Probe CUST_2256_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2256_PI426222305 | JHI_St_60k_v1 | DMT400072463 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
All Microarray Probes Designed to Gene DMG400028192
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2317_PI426222305 | JHI_St_60k_v1 | DMT400072464 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
| CUST_2114_PI426222305 | JHI_St_60k_v1 | DMT400072461 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
| CUST_2256_PI426222305 | JHI_St_60k_v1 | DMT400072463 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
| CUST_2198_PI426222305 | JHI_St_60k_v1 | DMT400072462 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
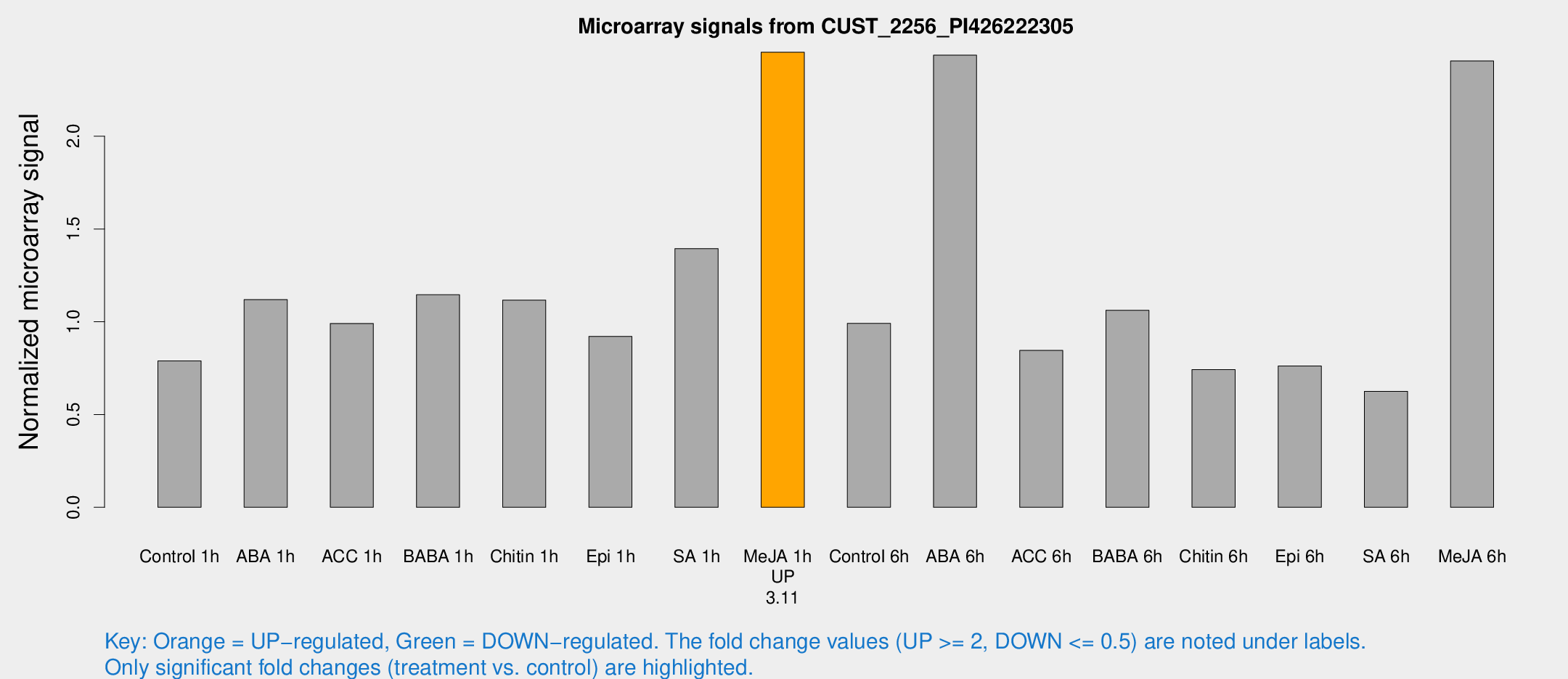
Microarray Signals from CUST_2256_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 163.064 | 28.4232 | 0.788843 | 0.143237 |
| ABA 1h | 210.924 | 54.7001 | 1.11931 | 0.205552 |
| ACC 1h | 215.63 | 48.9349 | 0.990779 | 0.182815 |
| BABA 1h | 237.093 | 56.5379 | 1.14554 | 0.206763 |
| Chitin 1h | 203.644 | 24.3807 | 1.11679 | 0.113378 |
| Epi 1h | 159.912 | 9.69118 | 0.92113 | 0.0557485 |
| SA 1h | 296.392 | 53.042 | 1.39419 | 0.18023 |
| Me-JA 1h | 410.251 | 61.0783 | 2.45265 | 0.20211 |
| Control 6h | 232.724 | 77.4114 | 0.991312 | 0.371523 |
| ABA 6h | 546.872 | 118.922 | 2.43685 | 0.471902 |
| ACC 6h | 199.496 | 34.4834 | 0.845424 | 0.0511426 |
| BABA 6h | 243.052 | 36.3833 | 1.06183 | 0.119251 |
| Chitin 6h | 158.501 | 9.75798 | 0.742148 | 0.0455812 |
| Epi 6h | 192.983 | 67.1579 | 0.761997 | 0.255143 |
| SA 6h | 134.926 | 35.4588 | 0.625134 | 0.126724 |
| Me-JA 6h | 508.988 | 112.563 | 2.40549 | 0.445667 |
Source Transcript PGSC0003DMT400072463 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc10g080930.1 | +2 | 5e-166 | 495 | 263/279 (94%) | evidence_code:10F0H1E1IEG genomic_reference:SL2.50ch10 gene_region:61435700-61439911 transcript_region:SL2.50ch10:61435700..61439911- go_terms:GO:0005634 functional_description:Transcription elongation factor A protein 2 (AHRD V1 *--- TCEA2_HUMAN); contains Interpro domain(s) IPR010990 Transcription elongation factor, TFIIS/elongin A/CRSP70, N-terminal |
| TAIR PP10 | AT5G05140.1 | +2 | 2e-81 | 222 | 139/286 (49%) | Transcription elongation factor (TFIIS) family protein | chr5:1520353-1522297 FORWARD LENGTH=436 |
