Probe CUST_2198_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2198_PI426222305 | JHI_St_60k_v1 | DMT400072462 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
All Microarray Probes Designed to Gene DMG400028192
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2317_PI426222305 | JHI_St_60k_v1 | DMT400072464 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
| CUST_2114_PI426222305 | JHI_St_60k_v1 | DMT400072461 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
| CUST_2256_PI426222305 | JHI_St_60k_v1 | DMT400072463 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
| CUST_2198_PI426222305 | JHI_St_60k_v1 | DMT400072462 | ACACAGATATTTGGGGAGGTGATGAGGATCAAAGATGTCATTGATAACAACCAAGCTGAG |
Microarray Signals from CUST_2198_PI426222305
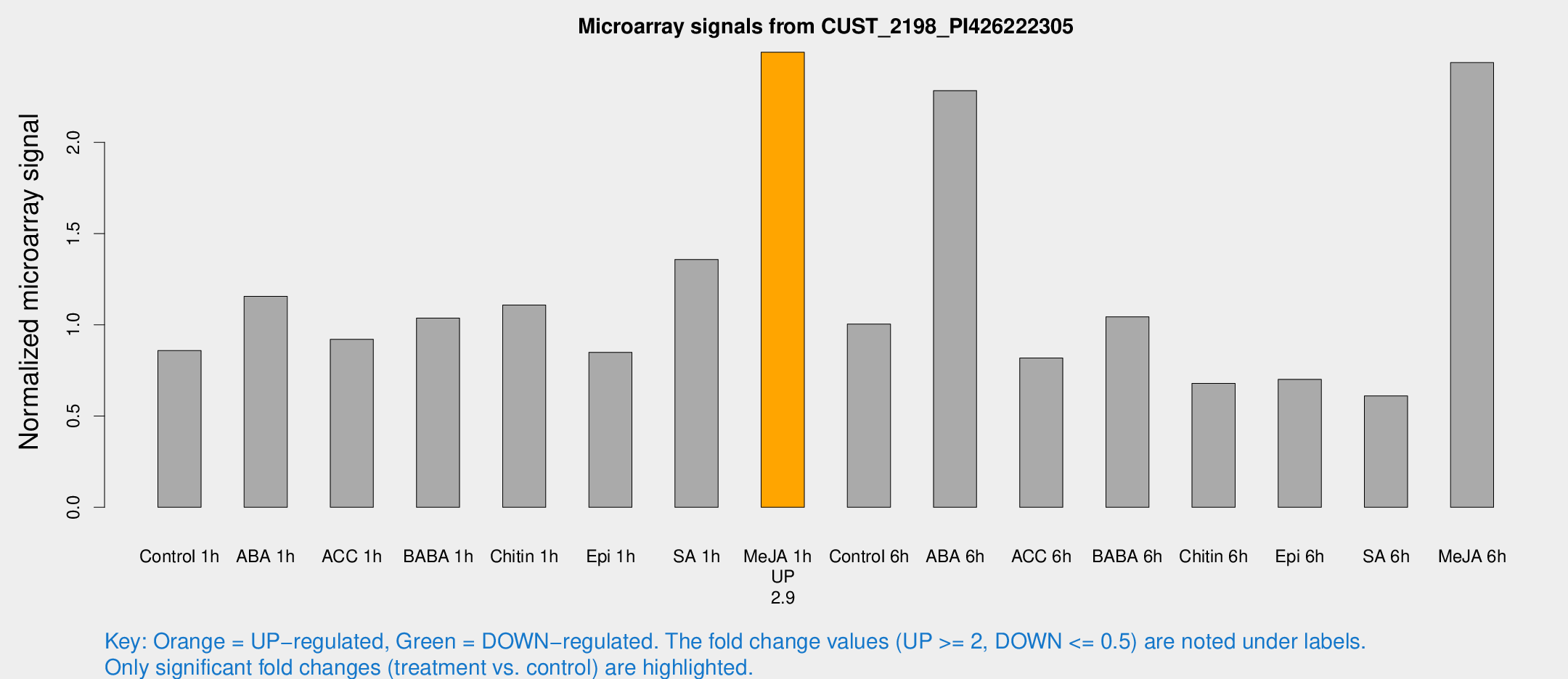
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 169.809 | 24.8484 | 0.858934 | 0.142083 |
| ABA 1h | 204.257 | 39.6727 | 1.15573 | 0.144061 |
| ACC 1h | 195.841 | 47.4502 | 0.920491 | 0.194874 |
| BABA 1h | 200.456 | 36.3683 | 1.03644 | 0.102949 |
| Chitin 1h | 196.915 | 29.8374 | 1.10828 | 0.153162 |
| Epi 1h | 143.626 | 15.3159 | 0.849215 | 0.086353 |
| SA 1h | 284.251 | 63.0905 | 1.35784 | 0.246078 |
| Me-JA 1h | 400.592 | 53.5695 | 2.49337 | 0.159265 |
| Control 6h | 233.774 | 90.4893 | 1.00425 | 0.41855 |
| ABA 6h | 516.042 | 145.512 | 2.28276 | 0.661221 |
| ACC 6h | 184.834 | 25.393 | 0.817895 | 0.0720618 |
| BABA 6h | 236.401 | 48.9304 | 1.04386 | 0.194337 |
| Chitin 6h | 140.909 | 14.2614 | 0.678907 | 0.0825742 |
| Epi 6h | 165.991 | 49.8292 | 0.701254 | 0.196619 |
| SA 6h | 126.829 | 33.8036 | 0.610457 | 0.121282 |
| Me-JA 6h | 505.639 | 123.687 | 2.43639 | 0.537864 |
Source Transcript PGSC0003DMT400072462 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc10g080930.1 | +2 | 0.0 | 473 | 252/259 (97%) | evidence_code:10F0H1E1IEG genomic_reference:SL2.50ch10 gene_region:61435700-61439911 transcript_region:SL2.50ch10:61435700..61439911- go_terms:GO:0005634 functional_description:Transcription elongation factor A protein 2 (AHRD V1 *--- TCEA2_HUMAN); contains Interpro domain(s) IPR010990 Transcription elongation factor, TFIIS/elongin A/CRSP70, N-terminal |
| TAIR PP10 | AT5G05140.1 | +2 | 1e-80 | 208 | 132/272 (49%) | Transcription elongation factor (TFIIS) family protein | chr5:1520353-1522297 FORWARD LENGTH=436 |
