Probe CUST_2149_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2149_PI426222305 | JHI_St_60k_v1 | DMT400072206 | GAGCCTATCTCCTTTCTTACCTTTTAGTAATTGGTTGTATGACGACTAATCTGTCATTTG |
All Microarray Probes Designed to Gene DMG400028097
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2149_PI426222305 | JHI_St_60k_v1 | DMT400072206 | GAGCCTATCTCCTTTCTTACCTTTTAGTAATTGGTTGTATGACGACTAATCTGTCATTTG |
| CUST_2237_PI426222305 | JHI_St_60k_v1 | DMT400072205 | ATAACAAAGAAAATTTAATGGAGAACCATGCTGAAGTAGGACGCCAGCTGGAGCCATGGG |
| CUST_2030_PI426222305 | JHI_St_60k_v1 | DMT400072204 | TTCTCGTTCTGATTTACAACCTGATCAATCGTTGTATTCTTCAAACGTAACAGGTGTTGG |
| CUST_2408_PI426222305 | JHI_St_60k_v1 | DMT400072207 | TGAAATCAACTCGAATTCGAGTCAGGATTTACGTGCCGTAGCGATTGAGCTTGACGTTAG |
Microarray Signals from CUST_2149_PI426222305
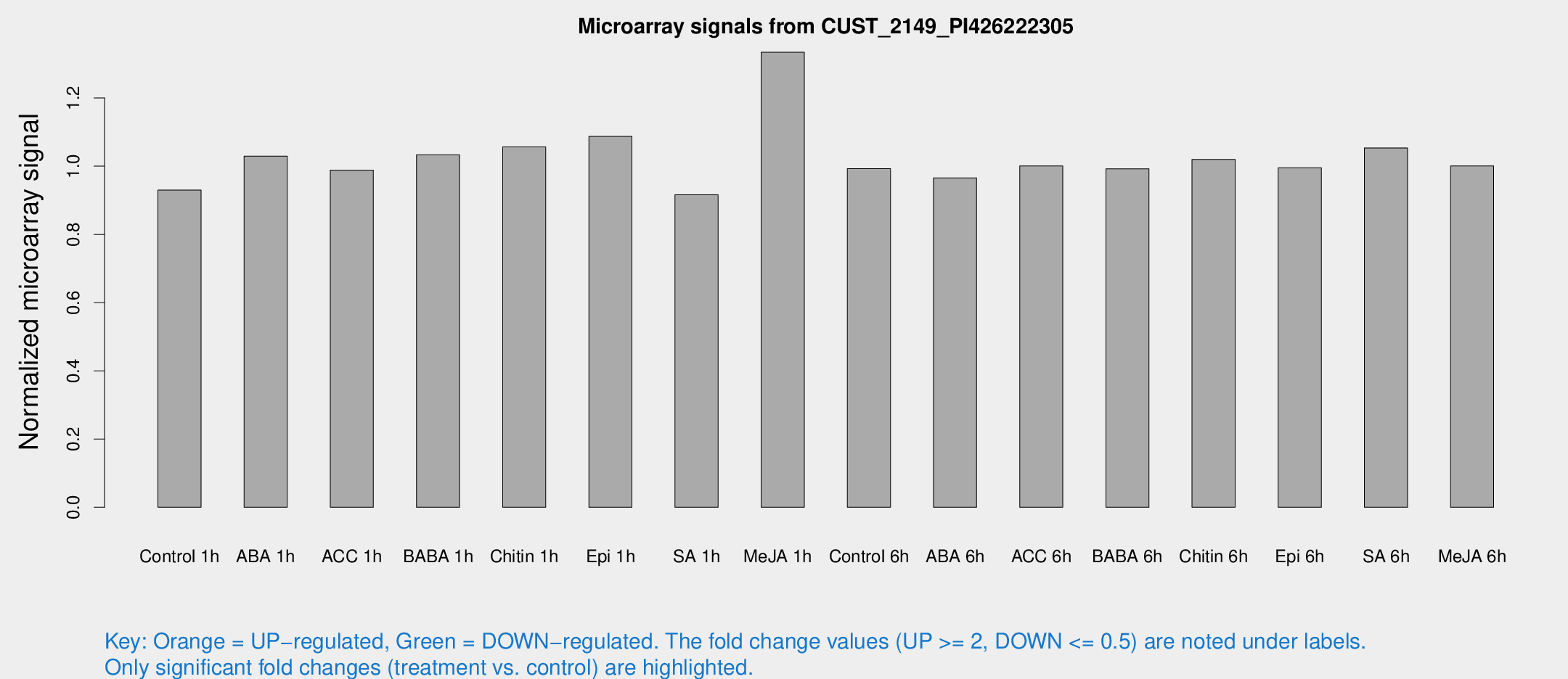
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.47234 | 3.16867 | 0.929865 | 0.538171 |
| ABA 1h | 5.34998 | 3.10274 | 1.02954 | 0.594531 |
| ACC 1h | 5.98108 | 3.47152 | 0.988693 | 0.573085 |
| BABA 1h | 5.86137 | 3.39617 | 1.03338 | 0.598513 |
| Chitin 1h | 5.59716 | 3.25193 | 1.05654 | 0.610723 |
| Epi 1h | 5.54514 | 3.21895 | 1.08726 | 0.630067 |
| SA 1h | 5.53553 | 3.20833 | 0.915928 | 0.530739 |
| Me-JA 1h | 6.45199 | 3.31777 | 1.33405 | 0.694314 |
| Control 6h | 5.85264 | 3.39308 | 0.992511 | 0.575264 |
| ABA 6h | 6.04432 | 3.50743 | 0.965618 | 0.559893 |
| ACC 6h | 6.92435 | 4.1189 | 1.00077 | 0.579913 |
| BABA 6h | 6.551 | 3.80333 | 0.992424 | 0.574694 |
| Chitin 6h | 6.38501 | 3.69751 | 1.01965 | 0.590417 |
| Epi 6h | 6.653 | 3.88359 | 0.995241 | 0.576308 |
| SA 6h | 6.1338 | 3.55513 | 1.05359 | 0.610263 |
| Me-JA 6h | 5.86727 | 3.4034 | 1.00079 | 0.579997 |
Source Transcript PGSC0003DMT400072206 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc10g080890.1 | +1 | 4e-112 | 330 | 164/170 (96%) | evidence_code:10F0H1E1IEG genomic_reference:SL2.50ch10 gene_region:61391472-61393927 transcript_region:SL2.50ch10:61391472..61393927+ go_terms:GO:0008152,GO:0055114 functional_description:3-oxoacyl-reductase (AHRD V1 **-- B6UEX0_MAIZE); contains Interpro domain(s) IPR002347 Glucose/ribitol dehydrogenase |
| TAIR PP10 | AT3G46170.1 | +1 | 1e-72 | 229 | 112/177 (63%) | NAD(P)-binding Rossmann-fold superfamily protein | chr3:16952723-16953589 REVERSE LENGTH=288 |
