Probe CUST_2030_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2030_PI426222305 | JHI_St_60k_v1 | DMT400072204 | TTCTCGTTCTGATTTACAACCTGATCAATCGTTGTATTCTTCAAACGTAACAGGTGTTGG |
All Microarray Probes Designed to Gene DMG400028097
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_2149_PI426222305 | JHI_St_60k_v1 | DMT400072206 | GAGCCTATCTCCTTTCTTACCTTTTAGTAATTGGTTGTATGACGACTAATCTGTCATTTG |
| CUST_2237_PI426222305 | JHI_St_60k_v1 | DMT400072205 | ATAACAAAGAAAATTTAATGGAGAACCATGCTGAAGTAGGACGCCAGCTGGAGCCATGGG |
| CUST_2030_PI426222305 | JHI_St_60k_v1 | DMT400072204 | TTCTCGTTCTGATTTACAACCTGATCAATCGTTGTATTCTTCAAACGTAACAGGTGTTGG |
| CUST_2408_PI426222305 | JHI_St_60k_v1 | DMT400072207 | TGAAATCAACTCGAATTCGAGTCAGGATTTACGTGCCGTAGCGATTGAGCTTGACGTTAG |
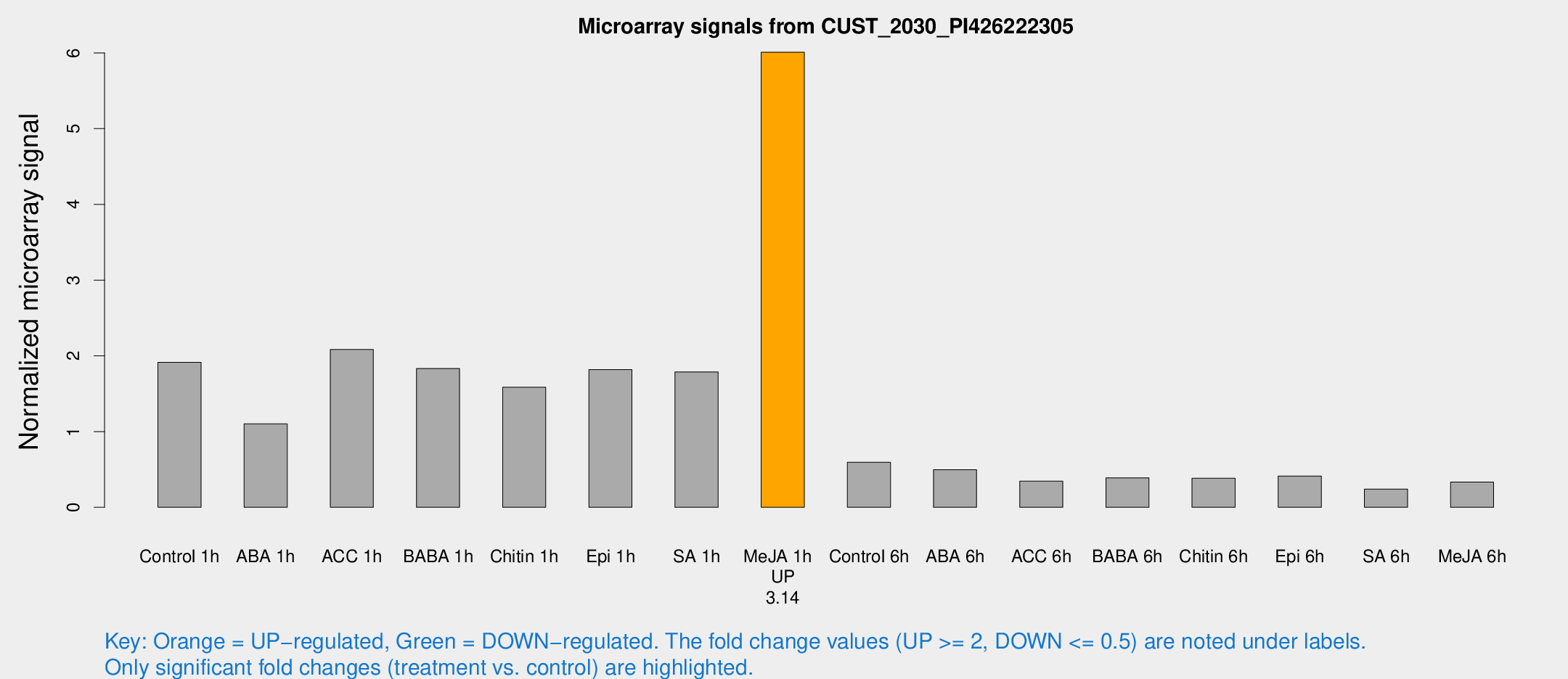
Microarray Signals from CUST_2030_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 55.9185 | 14.3387 | 1.91429 | 0.350634 |
| ABA 1h | 29.1661 | 8.59646 | 1.10196 | 0.252269 |
| ACC 1h | 66.0358 | 20.5494 | 2.08389 | 0.63717 |
| BABA 1h | 51.3744 | 11.6086 | 1.83257 | 0.27732 |
| Chitin 1h | 39.4335 | 4.01378 | 1.58605 | 0.2581 |
| Epi 1h | 43.3772 | 4.09293 | 1.81867 | 0.17249 |
| SA 1h | 58.8794 | 23.6395 | 1.78779 | 0.73653 |
| Me-JA 1h | 138.133 | 21.0294 | 6.0091 | 0.449183 |
| Control 6h | 19.6391 | 6.95666 | 0.594943 | 0.243232 |
| ABA 6h | 17.9439 | 7.8701 | 0.497321 | 0.256239 |
| ACC 6h | 13.7068 | 6.83904 | 0.34487 | 0.146914 |
| BABA 6h | 13.1135 | 4.02157 | 0.389614 | 0.145173 |
| Chitin 6h | 13.3062 | 5.75365 | 0.383518 | 0.187344 |
| Epi 6h | 13.7807 | 4.17709 | 0.4137 | 0.153151 |
| SA 6h | 6.53176 | 3.79197 | 0.240289 | 0.139449 |
| Me-JA 6h | 11.2652 | 5.41432 | 0.333032 | 0.154399 |
Source Transcript PGSC0003DMT400072204 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc10g080890.1 | +3 | 1e-88 | 278 | 178/201 (89%) | evidence_code:10F0H1E1IEG genomic_reference:SL2.50ch10 gene_region:61391472-61393927 transcript_region:SL2.50ch10:61391472..61393927+ go_terms:GO:0008152,GO:0055114 functional_description:3-oxoacyl-reductase (AHRD V1 **-- B6UEX0_MAIZE); contains Interpro domain(s) IPR002347 Glucose/ribitol dehydrogenase |
| TAIR PP10 | AT1G63380.1 | +3 | 2e-55 | 190 | 120/201 (60%) | NAD(P)-binding Rossmann-fold superfamily protein | chr1:23505582-23506504 FORWARD LENGTH=282 |
