Probe CUST_18070_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18070_PI426222305 | JHI_St_60k_v1 | DMT400018225 | CGTGCTAGCACGAGCCTAATATATGTTGCTCTCTTAATATCAAATTATGTGATATAGTTG |
All Microarray Probes Designed to Gene DMG400007074
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17863_PI426222305 | JHI_St_60k_v1 | DMT400018226 | GACAAATGGGCAAAACATAGAAAAATCATCAATCCCGCGTTCCATCTAGAGAAGTTAAAG |
| CUST_18070_PI426222305 | JHI_St_60k_v1 | DMT400018225 | CGTGCTAGCACGAGCCTAATATATGTTGCTCTCTTAATATCAAATTATGTGATATAGTTG |
| CUST_17984_PI426222305 | JHI_St_60k_v1 | DMT400018224 | GTCTTGCAAGTGTTTGGGAGTCGGAAACCAGATTTTGATGGCTTAAATCATCTAAAAGTT |
| CUST_17891_PI426222305 | JHI_St_60k_v1 | DMT400018223 | AAGTGTTAGTGTTATTGACATTTTCCCTGGTTCACCATTTTCTCAAACAGCATATGCTTC |
| CUST_17945_PI426222305 | JHI_St_60k_v1 | DMT400018219 | CTGATCATAATTTAGCACCAAGAATGGTGCCTTTCTTTCTTGAAACCATCAAGAAATATG |
| CUST_18057_PI426222305 | JHI_St_60k_v1 | DMT400018220 | AGTGCTTGGGAGTCGGAAACCAGATTTTGATGGATTAAATCATCTAAAAGTTGCAAAAAA |
| CUST_18102_PI426222305 | JHI_St_60k_v1 | DMT400018222 | CTGATCATAATTTAGCACCAAGAATGGTGCCTTTCTTTCTTGAAACCATCAAGAAATATG |
Microarray Signals from CUST_18070_PI426222305
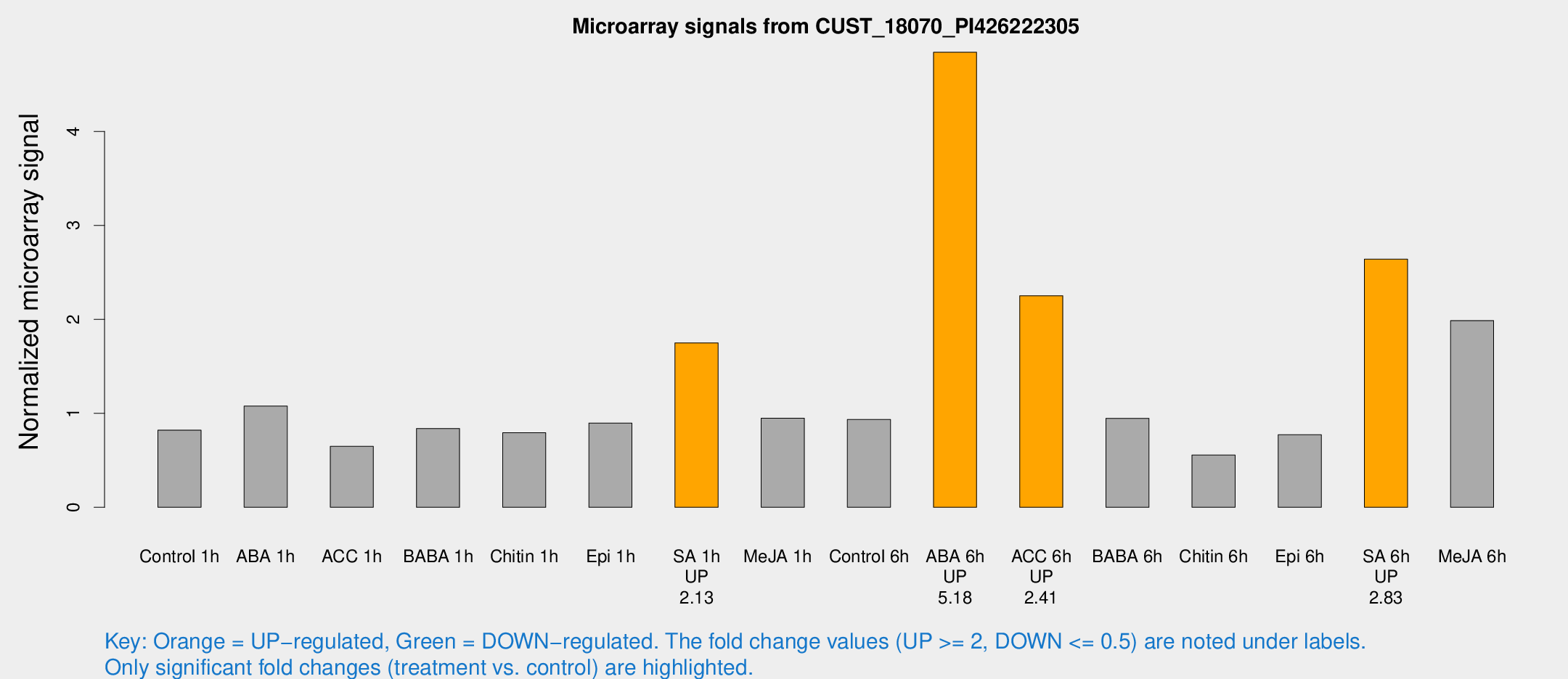
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 315.576 | 23.8347 | 0.821491 | 0.0483526 |
| ABA 1h | 364.168 | 21.3239 | 1.07684 | 0.07468 |
| ACC 1h | 284.176 | 85.8285 | 0.64811 | 0.191997 |
| BABA 1h | 337.471 | 93.0681 | 0.838252 | 0.183502 |
| Chitin 1h | 274.034 | 20.0149 | 0.793205 | 0.095845 |
| Epi 1h | 300 | 33.6152 | 0.896413 | 0.103599 |
| SA 1h | 690.352 | 50.2897 | 1.74937 | 0.101411 |
| Me-JA 1h | 305.556 | 54.6509 | 0.947643 | 0.0925881 |
| Control 6h | 370.861 | 65.6665 | 0.934159 | 0.0938418 |
| ABA 6h | 2242.93 | 703.476 | 4.84256 | 1.77754 |
| ACC 6h | 1005.89 | 142.117 | 2.2512 | 0.523808 |
| BABA 6h | 419.464 | 81.8443 | 0.947101 | 0.151343 |
| Chitin 6h | 226.612 | 13.7311 | 0.555491 | 0.0370737 |
| Epi 6h | 335.54 | 31.2317 | 0.771896 | 0.0499876 |
| SA 6h | 1047.37 | 215.804 | 2.64023 | 0.314691 |
| Me-JA 6h | 771.833 | 110.619 | 1.987 | 0.47514 |
Source Transcript PGSC0003DMT400018225 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14660.1 | +3 | 9e-57 | 198 | 87/141 (62%) | cytochrome P450, family 72, subfamily A, polypeptide 13 | chr3:4924960-4926911 FORWARD LENGTH=512 |
