Probe CUST_18057_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18057_PI426222305 | JHI_St_60k_v1 | DMT400018220 | AGTGCTTGGGAGTCGGAAACCAGATTTTGATGGATTAAATCATCTAAAAGTTGCAAAAAA |
All Microarray Probes Designed to Gene DMG400007074
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17863_PI426222305 | JHI_St_60k_v1 | DMT400018226 | GACAAATGGGCAAAACATAGAAAAATCATCAATCCCGCGTTCCATCTAGAGAAGTTAAAG |
| CUST_18070_PI426222305 | JHI_St_60k_v1 | DMT400018225 | CGTGCTAGCACGAGCCTAATATATGTTGCTCTCTTAATATCAAATTATGTGATATAGTTG |
| CUST_17984_PI426222305 | JHI_St_60k_v1 | DMT400018224 | GTCTTGCAAGTGTTTGGGAGTCGGAAACCAGATTTTGATGGCTTAAATCATCTAAAAGTT |
| CUST_17891_PI426222305 | JHI_St_60k_v1 | DMT400018223 | AAGTGTTAGTGTTATTGACATTTTCCCTGGTTCACCATTTTCTCAAACAGCATATGCTTC |
| CUST_17945_PI426222305 | JHI_St_60k_v1 | DMT400018219 | CTGATCATAATTTAGCACCAAGAATGGTGCCTTTCTTTCTTGAAACCATCAAGAAATATG |
| CUST_18057_PI426222305 | JHI_St_60k_v1 | DMT400018220 | AGTGCTTGGGAGTCGGAAACCAGATTTTGATGGATTAAATCATCTAAAAGTTGCAAAAAA |
| CUST_18102_PI426222305 | JHI_St_60k_v1 | DMT400018222 | CTGATCATAATTTAGCACCAAGAATGGTGCCTTTCTTTCTTGAAACCATCAAGAAATATG |
Microarray Signals from CUST_18057_PI426222305
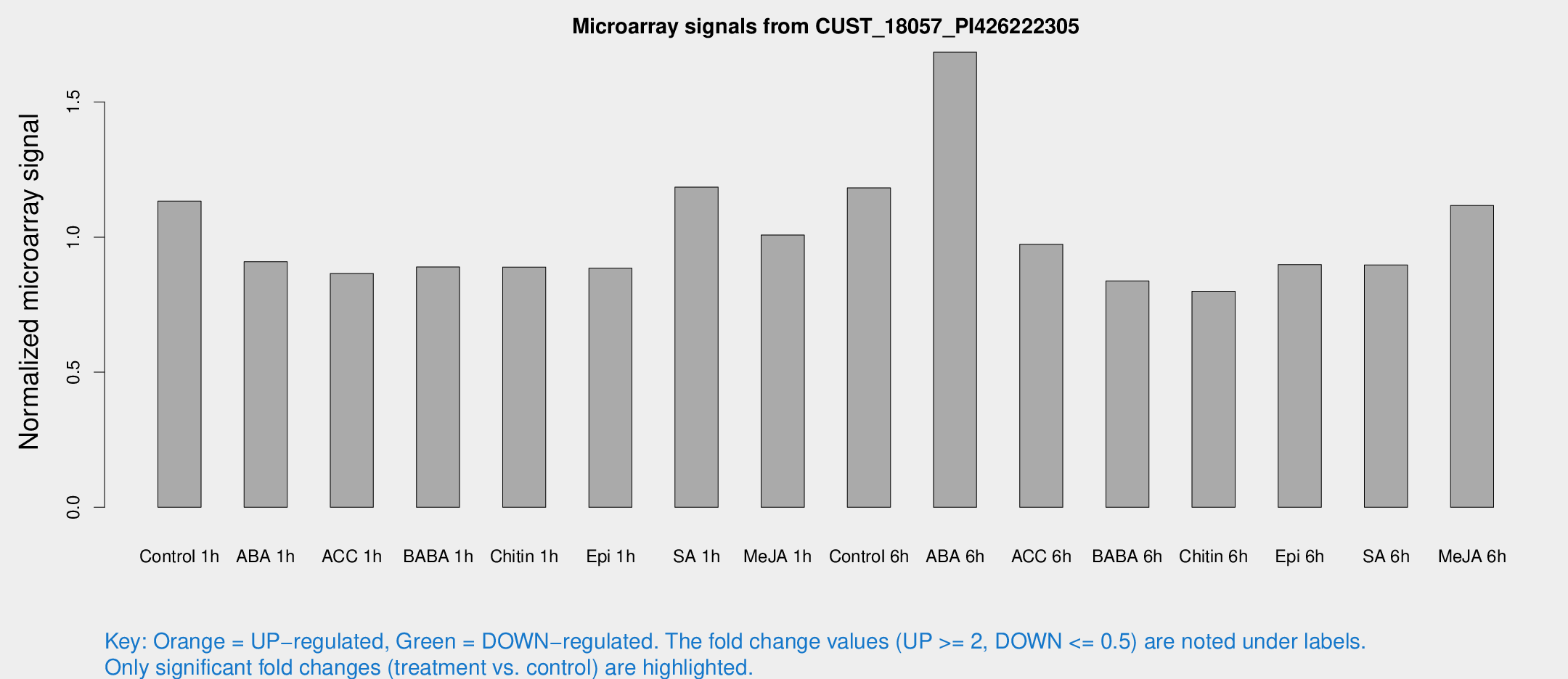
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2216.96 | 204.016 | 1.13336 | 0.0654564 |
| ABA 1h | 1565.51 | 91.5698 | 0.909021 | 0.0605361 |
| ACC 1h | 1830.69 | 401.963 | 0.865439 | 0.159148 |
| BABA 1h | 1805.32 | 450.598 | 0.889387 | 0.181406 |
| Chitin 1h | 1580.13 | 220.129 | 0.889336 | 0.0570035 |
| Epi 1h | 1492.6 | 92.4902 | 0.885316 | 0.0511518 |
| SA 1h | 2441.6 | 410.132 | 1.18534 | 0.163719 |
| Me-JA 1h | 1664.92 | 326.606 | 1.00807 | 0.125878 |
| Control 6h | 2487.81 | 631.119 | 1.18234 | 0.254154 |
| ABA 6h | 3609.59 | 689.807 | 1.6845 | 0.243035 |
| ACC 6h | 2195.27 | 242.763 | 0.973872 | 0.0660488 |
| BABA 6h | 1838.79 | 190.39 | 0.837773 | 0.0718889 |
| Chitin 6h | 1656.53 | 96.0118 | 0.79968 | 0.0462083 |
| Epi 6h | 2029.79 | 375.119 | 0.898672 | 0.153156 |
| SA 6h | 2111.25 | 780.125 | 0.896994 | 0.379698 |
| Me-JA 6h | 2168.8 | 152.376 | 1.11698 | 0.0645151 |
Source Transcript PGSC0003DMT400018220 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G17060.1 | +2 | 2e-128 | 240 | 107/226 (47%) | cytochrome p450 72c1 | chr1:5832282-5835255 REVERSE LENGTH=476 |
