Probe CUST_17891_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17891_PI426222305 | JHI_St_60k_v1 | DMT400018223 | AAGTGTTAGTGTTATTGACATTTTCCCTGGTTCACCATTTTCTCAAACAGCATATGCTTC |
All Microarray Probes Designed to Gene DMG400007074
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17863_PI426222305 | JHI_St_60k_v1 | DMT400018226 | GACAAATGGGCAAAACATAGAAAAATCATCAATCCCGCGTTCCATCTAGAGAAGTTAAAG |
| CUST_18070_PI426222305 | JHI_St_60k_v1 | DMT400018225 | CGTGCTAGCACGAGCCTAATATATGTTGCTCTCTTAATATCAAATTATGTGATATAGTTG |
| CUST_17984_PI426222305 | JHI_St_60k_v1 | DMT400018224 | GTCTTGCAAGTGTTTGGGAGTCGGAAACCAGATTTTGATGGCTTAAATCATCTAAAAGTT |
| CUST_17891_PI426222305 | JHI_St_60k_v1 | DMT400018223 | AAGTGTTAGTGTTATTGACATTTTCCCTGGTTCACCATTTTCTCAAACAGCATATGCTTC |
| CUST_17945_PI426222305 | JHI_St_60k_v1 | DMT400018219 | CTGATCATAATTTAGCACCAAGAATGGTGCCTTTCTTTCTTGAAACCATCAAGAAATATG |
| CUST_18057_PI426222305 | JHI_St_60k_v1 | DMT400018220 | AGTGCTTGGGAGTCGGAAACCAGATTTTGATGGATTAAATCATCTAAAAGTTGCAAAAAA |
| CUST_18102_PI426222305 | JHI_St_60k_v1 | DMT400018222 | CTGATCATAATTTAGCACCAAGAATGGTGCCTTTCTTTCTTGAAACCATCAAGAAATATG |
Microarray Signals from CUST_17891_PI426222305
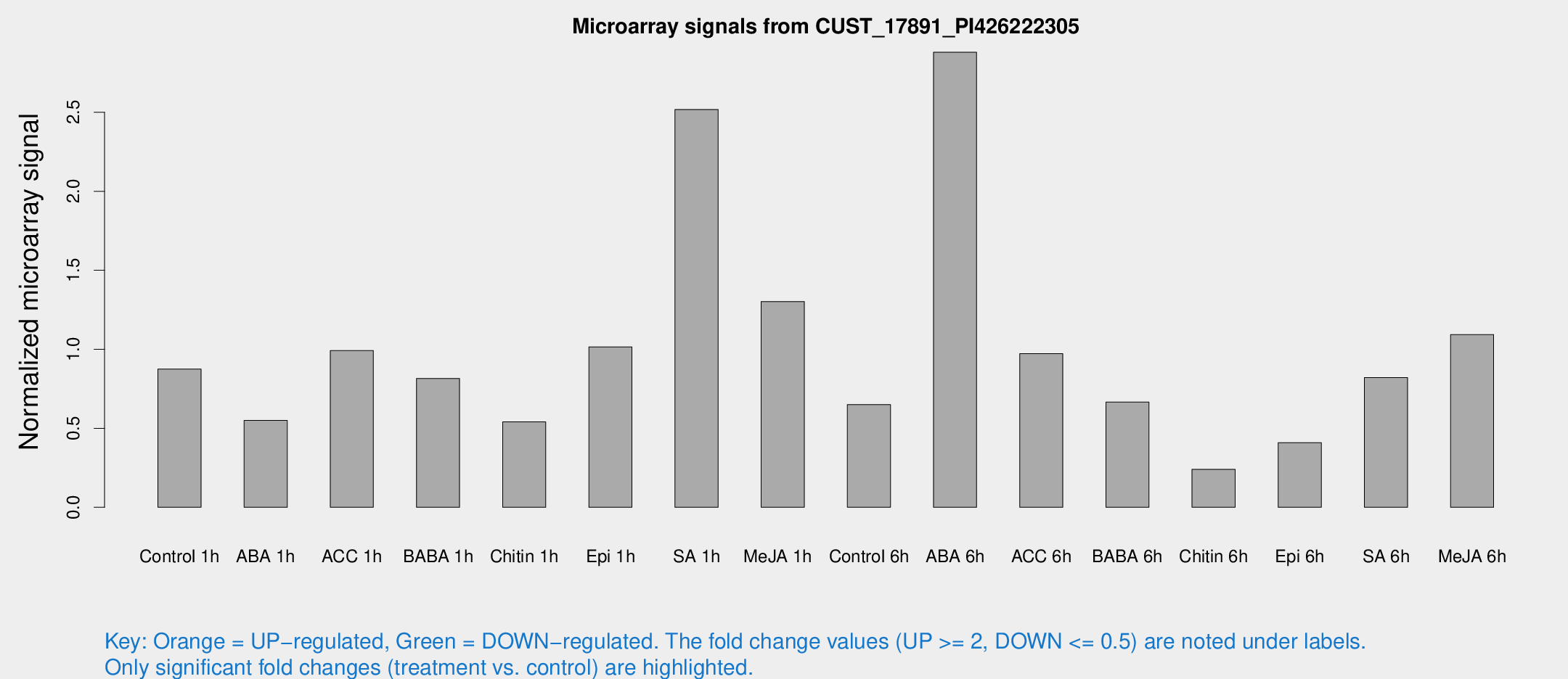
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 25.7938 | 5.48837 | 0.874743 | 0.243871 |
| ABA 1h | 17.449 | 8.28199 | 0.550467 | 0.307754 |
| ACC 1h | 30.5844 | 6.70771 | 0.991864 | 0.181009 |
| BABA 1h | 29.8521 | 12.9007 | 0.81542 | 0.503808 |
| Chitin 1h | 16.0748 | 6.53915 | 0.540778 | 0.256192 |
| Epi 1h | 24.8898 | 3.81806 | 1.01538 | 0.157145 |
| SA 1h | 79.4121 | 22.9645 | 2.51786 | 0.703942 |
| Me-JA 1h | 35.4601 | 12.4212 | 1.30243 | 0.506845 |
| Control 6h | 23.004 | 8.72392 | 0.650431 | 0.324454 |
| ABA 6h | 110.801 | 41.4486 | 2.87997 | 1.84524 |
| ACC 6h | 31.8607 | 4.96789 | 0.972358 | 0.193091 |
| BABA 6h | 24.9832 | 9.34503 | 0.665526 | 0.295966 |
| Chitin 6h | 7.23338 | 4.19765 | 0.240239 | 0.139362 |
| Epi 6h | 18.1023 | 10.5104 | 0.409514 | 0.27918 |
| SA 6h | 24.5741 | 5.69194 | 0.821531 | 0.307138 |
| Me-JA 6h | 31.5362 | 4.78206 | 1.09291 | 0.240272 |
Source Transcript PGSC0003DMT400018223 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14620.1 | +2 | 2e-26 | 111 | 53/82 (65%) | cytochrome P450, family 72, subfamily A, polypeptide 8 | chr3:4914978-4916853 FORWARD LENGTH=515 |
