Probe CUST_17863_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17863_PI426222305 | JHI_St_60k_v1 | DMT400018226 | GACAAATGGGCAAAACATAGAAAAATCATCAATCCCGCGTTCCATCTAGAGAAGTTAAAG |
All Microarray Probes Designed to Gene DMG400007074
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17863_PI426222305 | JHI_St_60k_v1 | DMT400018226 | GACAAATGGGCAAAACATAGAAAAATCATCAATCCCGCGTTCCATCTAGAGAAGTTAAAG |
| CUST_18070_PI426222305 | JHI_St_60k_v1 | DMT400018225 | CGTGCTAGCACGAGCCTAATATATGTTGCTCTCTTAATATCAAATTATGTGATATAGTTG |
| CUST_17984_PI426222305 | JHI_St_60k_v1 | DMT400018224 | GTCTTGCAAGTGTTTGGGAGTCGGAAACCAGATTTTGATGGCTTAAATCATCTAAAAGTT |
| CUST_17891_PI426222305 | JHI_St_60k_v1 | DMT400018223 | AAGTGTTAGTGTTATTGACATTTTCCCTGGTTCACCATTTTCTCAAACAGCATATGCTTC |
| CUST_17945_PI426222305 | JHI_St_60k_v1 | DMT400018219 | CTGATCATAATTTAGCACCAAGAATGGTGCCTTTCTTTCTTGAAACCATCAAGAAATATG |
| CUST_18057_PI426222305 | JHI_St_60k_v1 | DMT400018220 | AGTGCTTGGGAGTCGGAAACCAGATTTTGATGGATTAAATCATCTAAAAGTTGCAAAAAA |
| CUST_18102_PI426222305 | JHI_St_60k_v1 | DMT400018222 | CTGATCATAATTTAGCACCAAGAATGGTGCCTTTCTTTCTTGAAACCATCAAGAAATATG |
Microarray Signals from CUST_17863_PI426222305
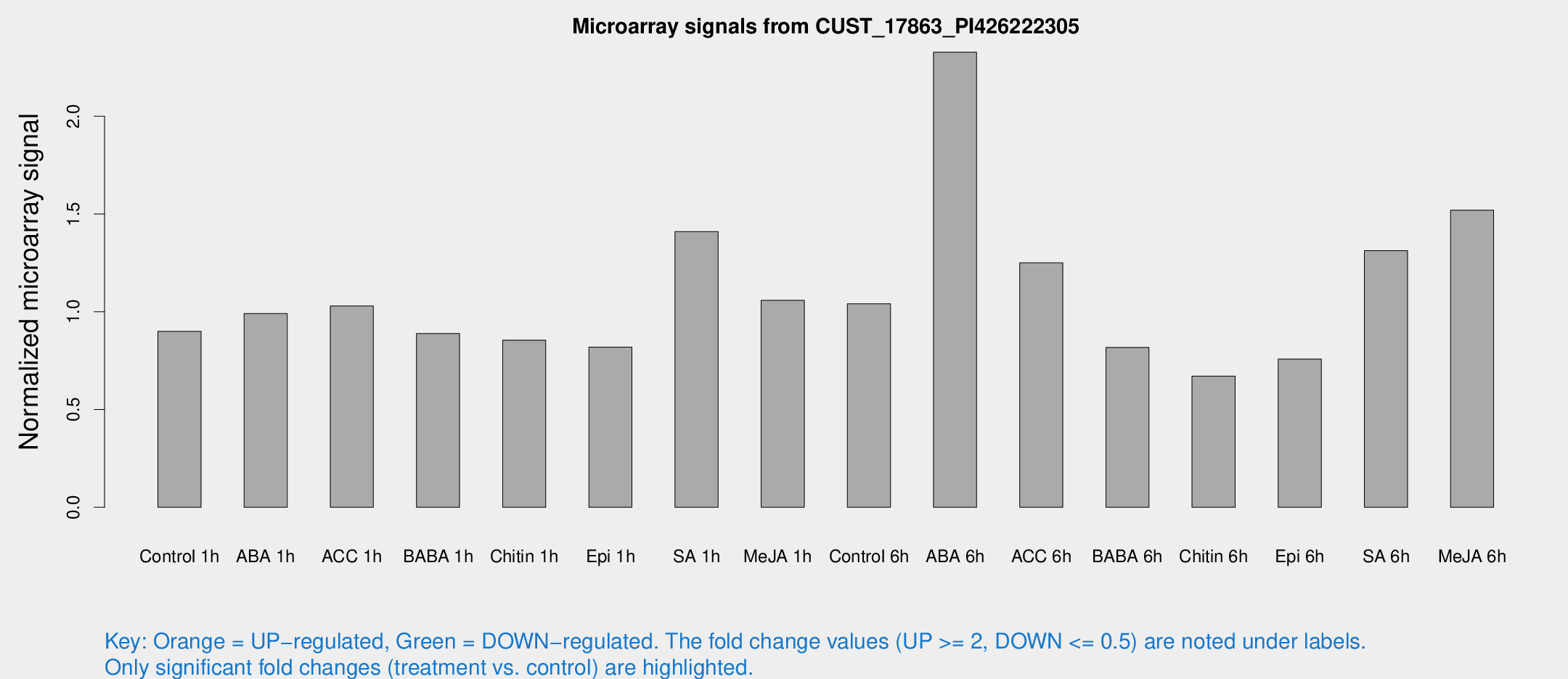
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 347.696 | 20.3337 | 0.899826 | 0.0707842 |
| ABA 1h | 340.203 | 22.7521 | 0.991109 | 0.0627827 |
| ACC 1h | 417.219 | 57.4083 | 1.02955 | 0.0755947 |
| BABA 1h | 355.763 | 92.209 | 0.888886 | 0.166503 |
| Chitin 1h | 298.702 | 22.2971 | 0.854831 | 0.050207 |
| Epi 1h | 275.968 | 22.3138 | 0.819689 | 0.0900452 |
| SA 1h | 605.76 | 149.385 | 1.41032 | 0.366175 |
| Me-JA 1h | 338.197 | 37.4835 | 1.05847 | 0.0620006 |
| Control 6h | 459.788 | 138.856 | 1.04113 | 0.334827 |
| ABA 6h | 1065.45 | 300.787 | 2.32747 | 0.761798 |
| ACC 6h | 558.58 | 51.099 | 1.25036 | 0.0761369 |
| BABA 6h | 355.202 | 22.8047 | 0.817846 | 0.0479735 |
| Chitin 6h | 278.408 | 26.0303 | 0.671314 | 0.0485003 |
| Epi 6h | 352.902 | 94.0121 | 0.757806 | 0.185409 |
| SA 6h | 527.56 | 115.317 | 1.31311 | 0.17807 |
| Me-JA 6h | 585.609 | 34.0164 | 1.51997 | 0.0881636 |
Source Transcript PGSC0003DMT400018226 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14690.1 | +2 | 0.0 | 539 | 254/419 (61%) | cytochrome P450, family 72, subfamily A, polypeptide 15 | chr3:4937410-4939310 FORWARD LENGTH=512 |
