Probe CUST_829_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_829_PI426222305 | JHI_St_60k_v1 | DMT400001329 | AAGTGTTTGATAACCCCAGTAATGGATATTGAAAGTTGGGGCCGTCGAAATCCCATAAAG |
All Microarray Probes Designed to Gene DMG400000501
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_829_PI426222305 | JHI_St_60k_v1 | DMT400001329 | AAGTGTTTGATAACCCCAGTAATGGATATTGAAAGTTGGGGCCGTCGAAATCCCATAAAG |
| CUST_848_PI426222305 | JHI_St_60k_v1 | DMT400001331 | GGCAATAAATACCACGAGAAAAGGAAATACTGAGCTTTAGAGGCATGTATGTTTGAATAA |
| CUST_1064_PI426222305 | JHI_St_60k_v1 | DMT400001330 | TTATGTTGGCAACTGCAAATTCAACCAAAATCCTAACAAGAGCTGCTTCTCTGCAGATAG |
Microarray Signals from CUST_829_PI426222305
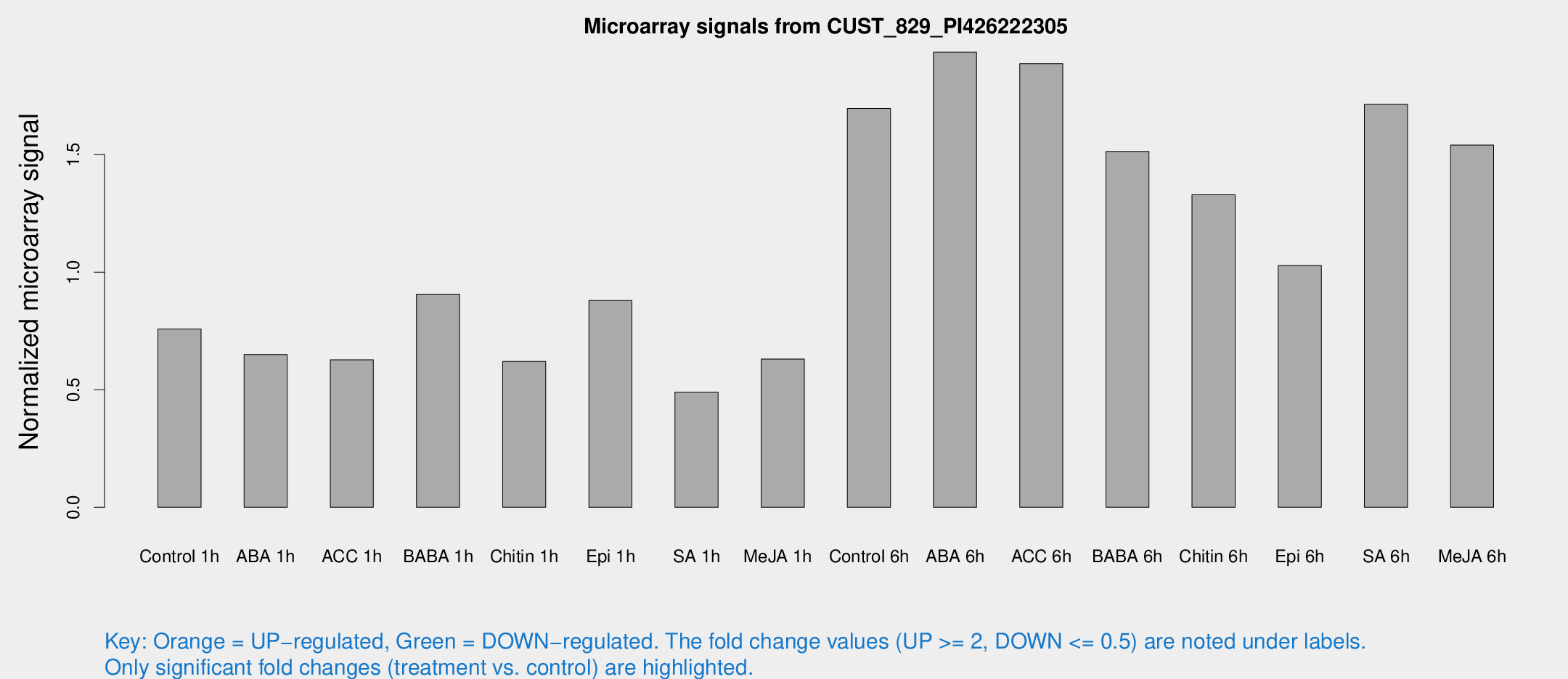
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 23.6803 | 3.83608 | 0.758201 | 0.127941 |
| ABA 1h | 18.3424 | 3.76866 | 0.649689 | 0.185533 |
| ACC 1h | 20.1749 | 3.39242 | 0.626936 | 0.112532 |
| BABA 1h | 27.2417 | 4.07114 | 0.90628 | 0.119872 |
| Chitin 1h | 17.3496 | 3.05403 | 0.62044 | 0.115157 |
| Epi 1h | 23.4115 | 3.1801 | 0.879174 | 0.122541 |
| SA 1h | 16.9928 | 5.47138 | 0.490154 | 0.131489 |
| Me-JA 1h | 15.7302 | 3.07061 | 0.630298 | 0.122974 |
| Control 6h | 54.0808 | 10.4942 | 1.69593 | 0.224302 |
| ABA 6h | 64.9089 | 11.2959 | 1.93504 | 0.276483 |
| ACC 6h | 69.1154 | 13.8039 | 1.88635 | 0.394586 |
| BABA 6h | 53.2082 | 8.57722 | 1.51307 | 0.217171 |
| Chitin 6h | 43.9344 | 5.68128 | 1.32896 | 0.19523 |
| Epi 6h | 36.5867 | 6.95626 | 1.02843 | 0.22533 |
| SA 6h | 61.7774 | 22.6175 | 1.7138 | 1.20062 |
| Me-JA 6h | 48.5533 | 8.7456 | 1.53985 | 0.190573 |
Source Transcript PGSC0003DMT400001329 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc02g087000.2 | +1 | 0.0 | 1476 | 761/774 (98%) | genomic_reference:SL2.50ch02 gene_region:44137630-44143256 transcript_region:SL2.50ch02:44137630..44143256+ go_terms:GO:0009674 functional_description:Potassium transporter (AHRD V1 **** D2JYH2_GOSHI); contains Interpro domain(s) IPR018519 Potassium uptake protein, kup IPR003855 K+ potassium transporter |
| TAIR PP10 | AT5G14880.1 | +1 | 0.0 | 1177 | 597/779 (77%) | Potassium transporter family protein | chr5:4814244-4817667 FORWARD LENGTH=781 |
