Probe CUST_1064_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_1064_PI426222305 | JHI_St_60k_v1 | DMT400001330 | TTATGTTGGCAACTGCAAATTCAACCAAAATCCTAACAAGAGCTGCTTCTCTGCAGATAG |
All Microarray Probes Designed to Gene DMG400000501
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_829_PI426222305 | JHI_St_60k_v1 | DMT400001329 | AAGTGTTTGATAACCCCAGTAATGGATATTGAAAGTTGGGGCCGTCGAAATCCCATAAAG |
| CUST_848_PI426222305 | JHI_St_60k_v1 | DMT400001331 | GGCAATAAATACCACGAGAAAAGGAAATACTGAGCTTTAGAGGCATGTATGTTTGAATAA |
| CUST_1064_PI426222305 | JHI_St_60k_v1 | DMT400001330 | TTATGTTGGCAACTGCAAATTCAACCAAAATCCTAACAAGAGCTGCTTCTCTGCAGATAG |
Microarray Signals from CUST_1064_PI426222305
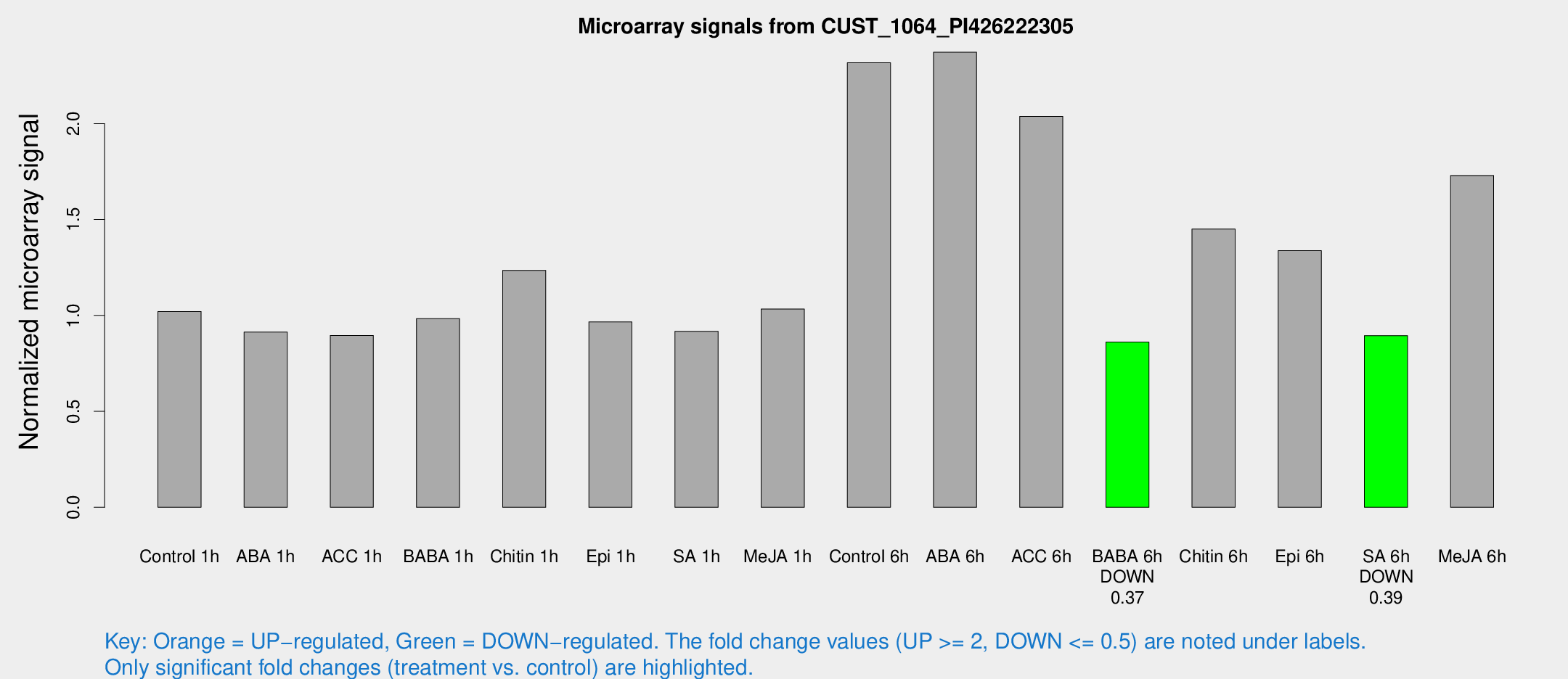
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.35037 | 2.93673 | 1.02073 | 0.475833 |
| ABA 1h | 4.86568 | 2.81957 | 0.913113 | 0.512 |
| ACC 1h | 5.76653 | 3.36841 | 0.895819 | 0.519192 |
| BABA 1h | 5.95223 | 3.09728 | 0.983689 | 0.520797 |
| Chitin 1h | 7.55361 | 2.95127 | 1.23526 | 0.564876 |
| Epi 1h | 5.06466 | 2.90111 | 0.966259 | 0.53724 |
| SA 1h | 5.93982 | 2.96555 | 0.916982 | 0.472183 |
| Me-JA 1h | 5.17754 | 3.00795 | 1.03319 | 0.588475 |
| Control 6h | 14.7499 | 3.11889 | 2.31745 | 0.519861 |
| ABA 6h | 20.3286 | 8.0277 | 2.37201 | 1.5893 |
| ACC 6h | 17.9036 | 6.44277 | 2.03753 | 1.07487 |
| BABA 6h | 6.00866 | 3.39211 | 0.861431 | 0.485652 |
| Chitin 6h | 10.1392 | 3.39651 | 1.45049 | 0.551497 |
| Epi 6h | 10.2312 | 3.56087 | 1.33817 | 0.54782 |
| SA 6h | 5.5104 | 3.19622 | 0.894966 | 0.518733 |
| Me-JA 6h | 12.8334 | 5.27256 | 1.72941 | 0.765516 |
Source Transcript PGSC0003DMT400001330 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc02g087000.2 | +2 | 0.0 | 923 | 469/477 (98%) | genomic_reference:SL2.50ch02 gene_region:44137630-44143256 transcript_region:SL2.50ch02:44137630..44143256+ go_terms:GO:0009674 functional_description:Potassium transporter (AHRD V1 **** D2JYH2_GOSHI); contains Interpro domain(s) IPR018519 Potassium uptake protein, kup IPR003855 K+ potassium transporter |
| TAIR PP10 | AT5G14880.1 | +2 | 0.0 | 710 | 360/482 (75%) | Potassium transporter family protein | chr5:4814244-4817667 FORWARD LENGTH=781 |
