Probe CUST_848_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_848_PI426222305 | JHI_St_60k_v1 | DMT400001331 | GGCAATAAATACCACGAGAAAAGGAAATACTGAGCTTTAGAGGCATGTATGTTTGAATAA |
All Microarray Probes Designed to Gene DMG400000501
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_829_PI426222305 | JHI_St_60k_v1 | DMT400001329 | AAGTGTTTGATAACCCCAGTAATGGATATTGAAAGTTGGGGCCGTCGAAATCCCATAAAG |
| CUST_848_PI426222305 | JHI_St_60k_v1 | DMT400001331 | GGCAATAAATACCACGAGAAAAGGAAATACTGAGCTTTAGAGGCATGTATGTTTGAATAA |
| CUST_1064_PI426222305 | JHI_St_60k_v1 | DMT400001330 | TTATGTTGGCAACTGCAAATTCAACCAAAATCCTAACAAGAGCTGCTTCTCTGCAGATAG |
Microarray Signals from CUST_848_PI426222305
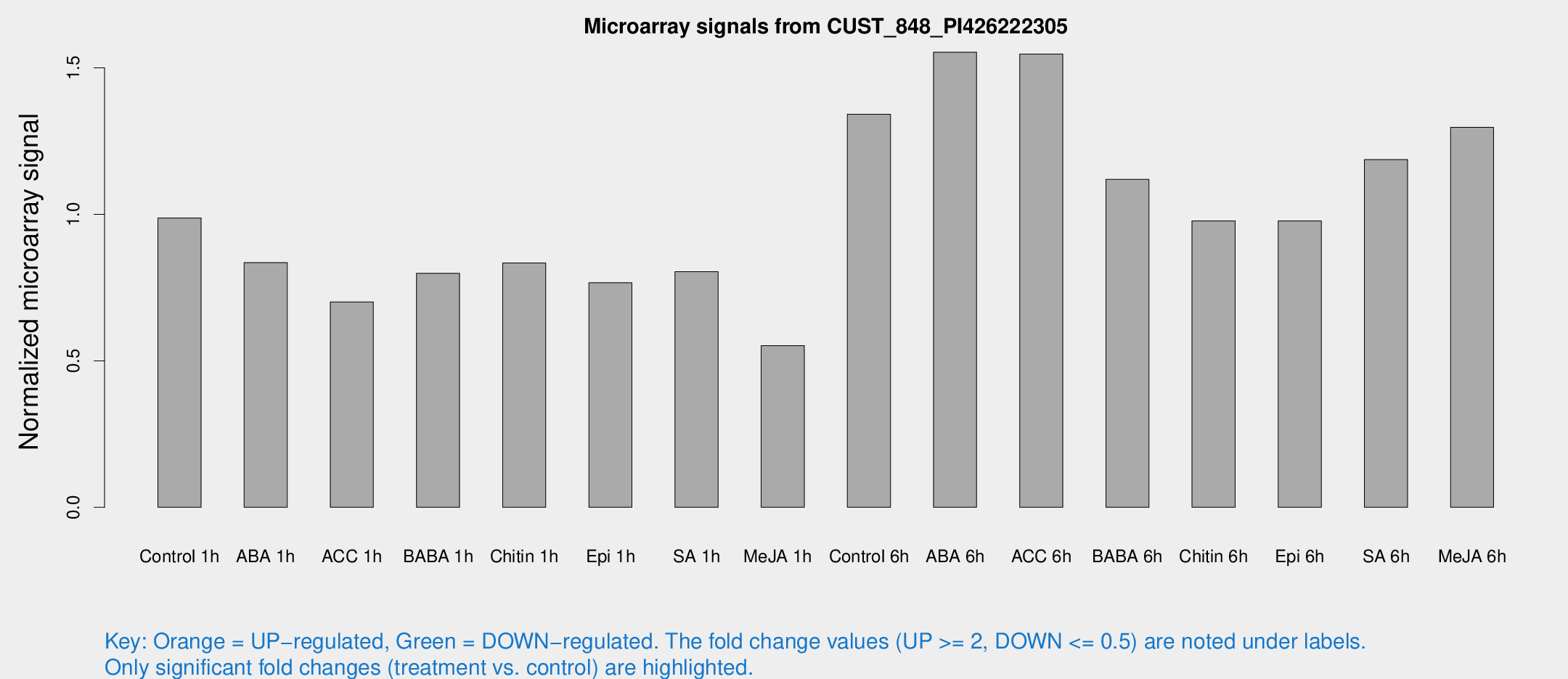
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 878.698 | 69.714 | 0.987797 | 0.057152 |
| ABA 1h | 654.496 | 38.0058 | 0.834989 | 0.0935594 |
| ACC 1h | 649.502 | 87.226 | 0.701017 | 0.0492023 |
| BABA 1h | 737.145 | 186.511 | 0.79902 | 0.158043 |
| Chitin 1h | 663.097 | 38.4359 | 0.833902 | 0.0626944 |
| Epi 1h | 594.242 | 65.1479 | 0.766619 | 0.0727435 |
| SA 1h | 753.507 | 133.045 | 0.804329 | 0.123534 |
| Me-JA 1h | 409.113 | 68.4695 | 0.551939 | 0.0440255 |
| Control 6h | 1244.74 | 247.611 | 1.34168 | 0.185579 |
| ABA 6h | 1494.89 | 230.299 | 1.55331 | 0.151712 |
| ACC 6h | 1572.62 | 91.0949 | 1.54683 | 0.195287 |
| BABA 6h | 1110.99 | 64.4505 | 1.11914 | 0.0647375 |
| Chitin 6h | 921.796 | 53.451 | 0.977685 | 0.0565964 |
| Epi 6h | 999.726 | 150.966 | 0.977619 | 0.108667 |
| SA 6h | 1081.46 | 203.308 | 1.18682 | 0.114273 |
| Me-JA 6h | 1160.1 | 138.262 | 1.29704 | 0.0859539 |
Source Transcript PGSC0003DMT400001331 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | Solyc02g087000.2 | +2 | 0.0 | 694 | 342/349 (98%) | genomic_reference:SL2.50ch02 gene_region:44137630-44143256 transcript_region:SL2.50ch02:44137630..44143256+ go_terms:GO:0009674 functional_description:Potassium transporter (AHRD V1 **** D2JYH2_GOSHI); contains Interpro domain(s) IPR018519 Potassium uptake protein, kup IPR003855 K+ potassium transporter |
| TAIR PP10 | AT5G14880.1 | +2 | 3e-169 | 498 | 247/354 (70%) | Potassium transporter family protein | chr5:4814244-4817667 FORWARD LENGTH=781 |
