Probe CUST_44039_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44039_PI426222305 | JHI_St_60k_v1 | DMT400016507 | GACAACTTAACTGGACTTCCCAAAATTATGGATTGCATGGCAAATGAAGTTGACATATTT |
All Microarray Probes Designed to Gene DMG400006446
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44003_PI426222305 | JHI_St_60k_v1 | DMT400016509 | GGATTTATGTACACTGGCTTTCTCTGTTTATTGCCTTCTCTTTTGTATTTACAAGTGCGT |
| CUST_44026_PI426222305 | JHI_St_60k_v1 | DMT400016508 | GACAACTTAACTGGACTTCCCAAAATTATGGATTGCATGGCAAATGAAGTTGACATATTT |
| CUST_44039_PI426222305 | JHI_St_60k_v1 | DMT400016507 | GACAACTTAACTGGACTTCCCAAAATTATGGATTGCATGGCAAATGAAGTTGACATATTT |
Microarray Signals from CUST_44039_PI426222305
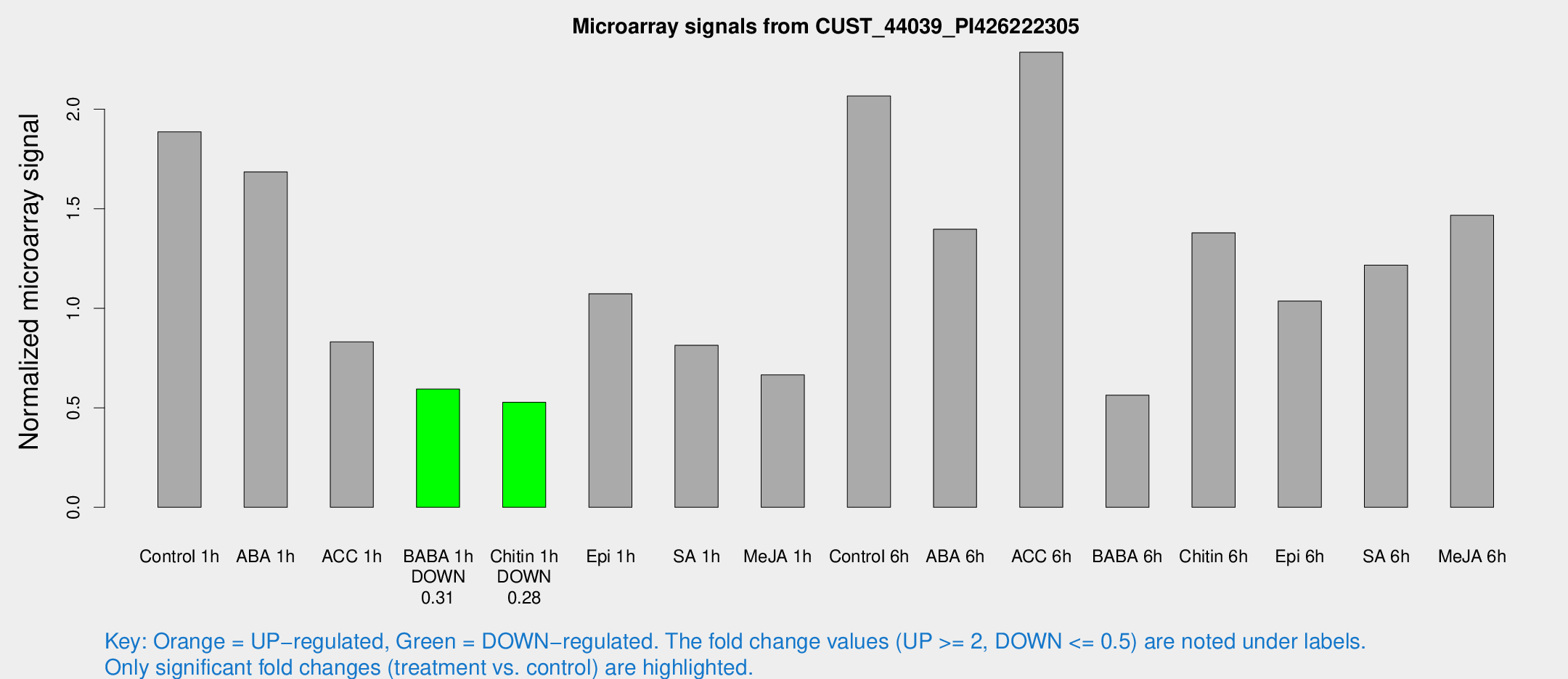
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 93.3072 | 6.2633 | 1.88622 | 0.12641 |
| ABA 1h | 84.8382 | 33.1625 | 1.68532 | 0.746107 |
| ACC 1h | 46.8652 | 13.9747 | 0.830837 | 0.239249 |
| BABA 1h | 28.348 | 3.75894 | 0.594021 | 0.0789924 |
| Chitin 1h | 23.7663 | 3.45569 | 0.527552 | 0.0784457 |
| Epi 1h | 48.3851 | 11.798 | 1.07299 | 0.262133 |
| SA 1h | 42.7319 | 8.14074 | 0.814246 | 0.208795 |
| Me-JA 1h | 28.5143 | 6.35552 | 0.665237 | 0.244564 |
| Control 6h | 118.443 | 47.1587 | 2.0664 | 1.14038 |
| ABA 6h | 77.3969 | 18.8502 | 1.39721 | 0.35541 |
| ACC 6h | 133.874 | 23.419 | 2.28637 | 0.340377 |
| BABA 6h | 31.5965 | 4.10632 | 0.563509 | 0.0792698 |
| Chitin 6h | 75.5949 | 15.8274 | 1.37923 | 0.260927 |
| Epi 6h | 60.4364 | 13.7583 | 1.03596 | 0.169603 |
| SA 6h | 62.1994 | 12.203 | 1.21682 | 0.127736 |
| Me-JA 6h | 74.7627 | 13.3597 | 1.46705 | 0.311009 |
Source Transcript PGSC0003DMT400016507 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G04200.1 | +1 | 1e-55 | 203 | 93/144 (65%) | FUNCTIONS IN: molecular_function unknown; INVOLVED IN: N-terminal protein myristoylation; LOCATED IN: cellular_component unknown; EXPRESSED IN: 22 plant structures; EXPRESSED DURING: 13 growth stages; CONTAINS InterPro DOMAIN/s: Dymeclin (InterPro:IPR019142); Has 395 Blast hits to 389 proteins in 117 species: Archae - 0; Bacteria - 0; Metazoa - 262; Fungi - 21; Plants - 68; Viruses - 0; Other Eukaryotes - 44 (source: NCBI BLink). | chr1:1109676-1113875 FORWARD LENGTH=732 |
