Probe CUST_44026_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44026_PI426222305 | JHI_St_60k_v1 | DMT400016508 | GACAACTTAACTGGACTTCCCAAAATTATGGATTGCATGGCAAATGAAGTTGACATATTT |
All Microarray Probes Designed to Gene DMG400006446
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44003_PI426222305 | JHI_St_60k_v1 | DMT400016509 | GGATTTATGTACACTGGCTTTCTCTGTTTATTGCCTTCTCTTTTGTATTTACAAGTGCGT |
| CUST_44026_PI426222305 | JHI_St_60k_v1 | DMT400016508 | GACAACTTAACTGGACTTCCCAAAATTATGGATTGCATGGCAAATGAAGTTGACATATTT |
| CUST_44039_PI426222305 | JHI_St_60k_v1 | DMT400016507 | GACAACTTAACTGGACTTCCCAAAATTATGGATTGCATGGCAAATGAAGTTGACATATTT |
Microarray Signals from CUST_44026_PI426222305
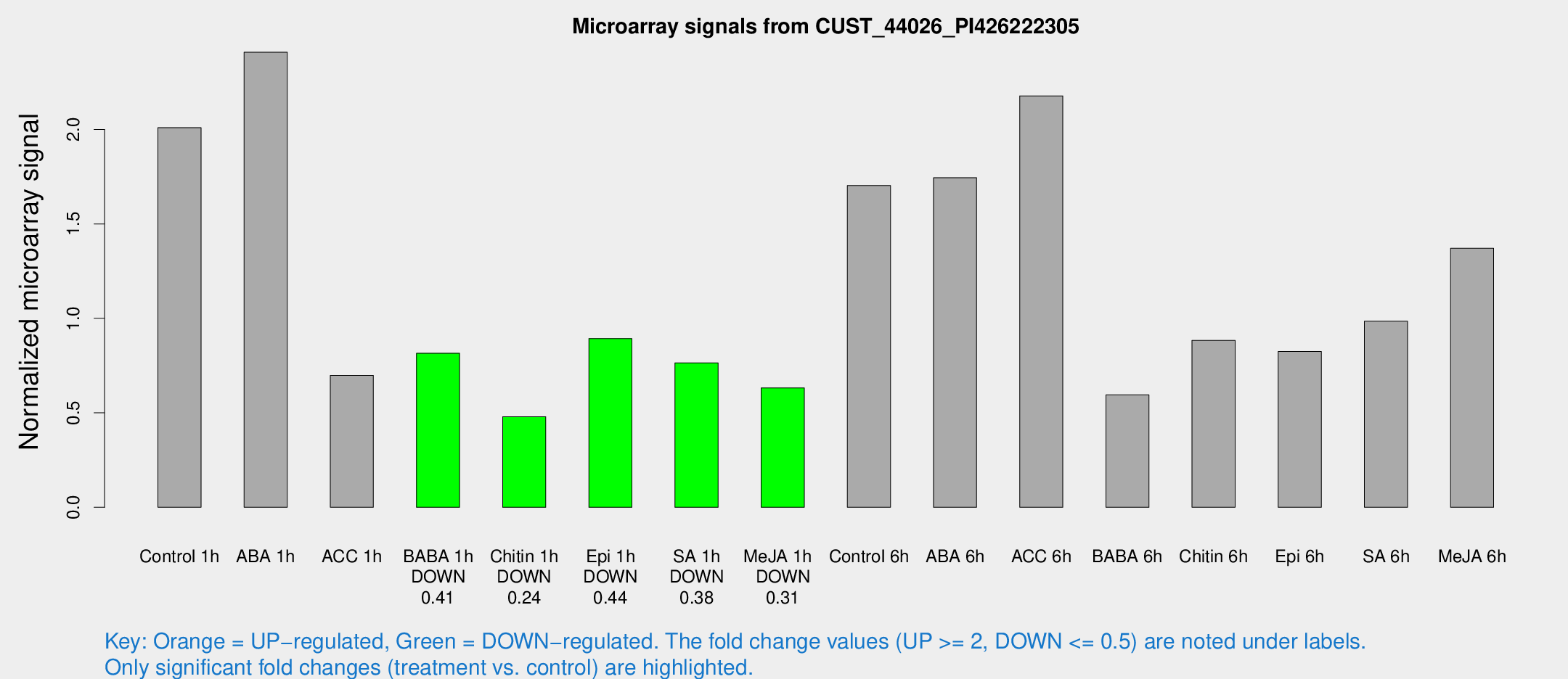
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 85.121 | 5.82325 | 2.00956 | 0.189366 |
| ABA 1h | 95.4244 | 24.3794 | 2.40851 | 0.640563 |
| ACC 1h | 44.2932 | 20.6931 | 0.698097 | 0.616006 |
| BABA 1h | 33.3107 | 3.7843 | 0.815451 | 0.140034 |
| Chitin 1h | 18.207 | 3.38154 | 0.479573 | 0.0890917 |
| Epi 1h | 34.1006 | 7.49657 | 0.893076 | 0.19798 |
| SA 1h | 34.0242 | 5.39443 | 0.764437 | 0.178667 |
| Me-JA 1h | 22.1863 | 3.44221 | 0.6316 | 0.14775 |
| Control 6h | 75.3729 | 16.5324 | 1.70293 | 0.557297 |
| ABA 6h | 85.4136 | 24.6902 | 1.74456 | 0.629202 |
| ACC 6h | 111.017 | 22.7436 | 2.17751 | 0.502548 |
| BABA 6h | 29.4125 | 6.21768 | 0.595081 | 0.140085 |
| Chitin 6h | 49.4195 | 23.458 | 0.883037 | 0.449964 |
| Epi 6h | 42.0319 | 11.5399 | 0.824899 | 0.16582 |
| SA 6h | 42.2835 | 6.56995 | 0.984865 | 0.10438 |
| Me-JA 6h | 59.7661 | 10.533 | 1.37059 | 0.325736 |
Source Transcript PGSC0003DMT400016508 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G04200.1 | +2 | 3e-102 | 334 | 161/242 (67%) | FUNCTIONS IN: molecular_function unknown; INVOLVED IN: N-terminal protein myristoylation; LOCATED IN: cellular_component unknown; EXPRESSED IN: 22 plant structures; EXPRESSED DURING: 13 growth stages; CONTAINS InterPro DOMAIN/s: Dymeclin (InterPro:IPR019142); Has 395 Blast hits to 389 proteins in 117 species: Archae - 0; Bacteria - 0; Metazoa - 262; Fungi - 21; Plants - 68; Viruses - 0; Other Eukaryotes - 44 (source: NCBI BLink). | chr1:1109676-1113875 FORWARD LENGTH=732 |
