Probe CUST_44003_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44003_PI426222305 | JHI_St_60k_v1 | DMT400016509 | GGATTTATGTACACTGGCTTTCTCTGTTTATTGCCTTCTCTTTTGTATTTACAAGTGCGT |
All Microarray Probes Designed to Gene DMG400006446
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44003_PI426222305 | JHI_St_60k_v1 | DMT400016509 | GGATTTATGTACACTGGCTTTCTCTGTTTATTGCCTTCTCTTTTGTATTTACAAGTGCGT |
| CUST_44026_PI426222305 | JHI_St_60k_v1 | DMT400016508 | GACAACTTAACTGGACTTCCCAAAATTATGGATTGCATGGCAAATGAAGTTGACATATTT |
| CUST_44039_PI426222305 | JHI_St_60k_v1 | DMT400016507 | GACAACTTAACTGGACTTCCCAAAATTATGGATTGCATGGCAAATGAAGTTGACATATTT |
Microarray Signals from CUST_44003_PI426222305
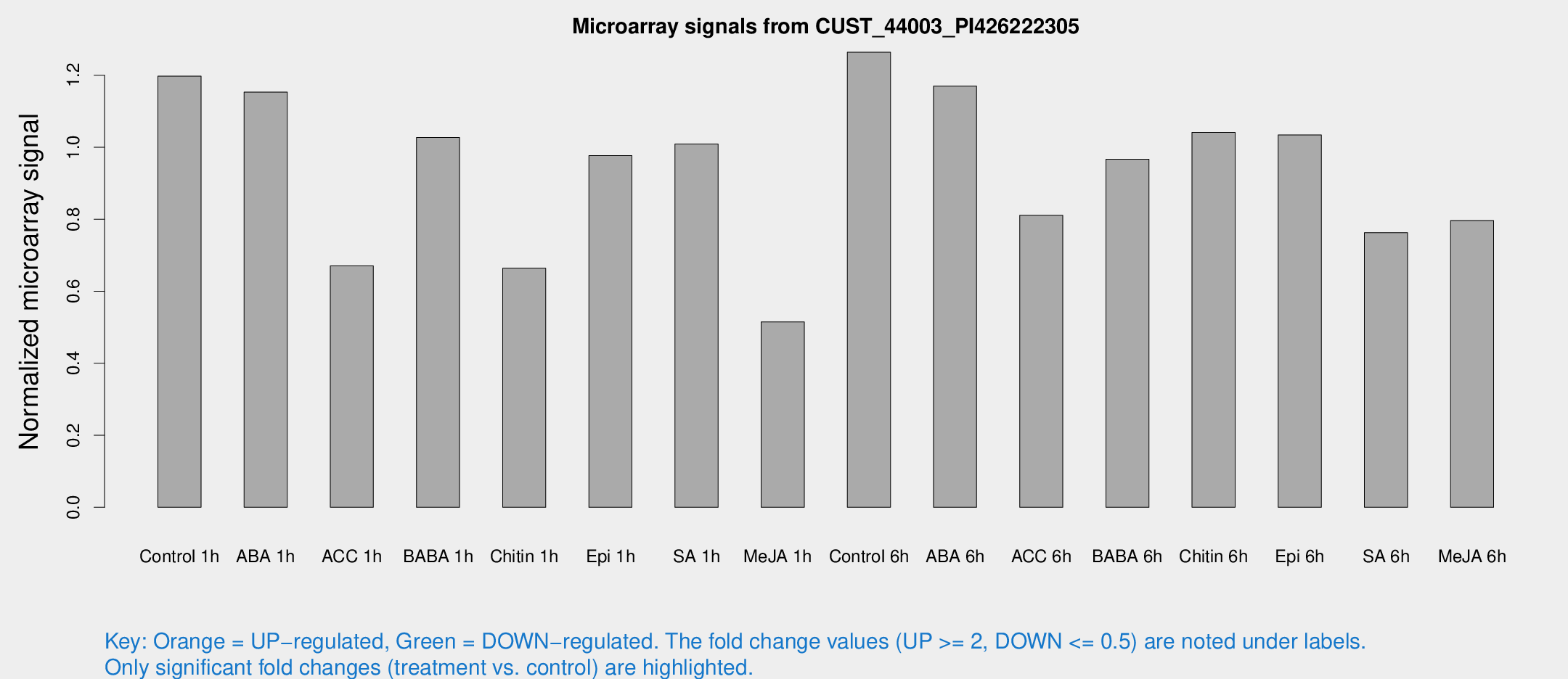
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 46.1201 | 4.69868 | 1.19758 | 0.119501 |
| ABA 1h | 39.9661 | 6.76878 | 1.15318 | 0.229512 |
| ACC 1h | 33.6786 | 13.035 | 0.67056 | 0.390828 |
| BABA 1h | 37.7745 | 4.51747 | 1.02714 | 0.122698 |
| Chitin 1h | 23.1887 | 3.98795 | 0.664084 | 0.11541 |
| Epi 1h | 32.8858 | 4.44943 | 0.976638 | 0.133517 |
| SA 1h | 39.88 | 4.23461 | 1.00895 | 0.107844 |
| Me-JA 1h | 18.2381 | 5.5573 | 0.514803 | 0.240009 |
| Control 6h | 52.0333 | 13.5804 | 1.26395 | 0.243317 |
| ABA 6h | 47.7689 | 4.7334 | 1.16996 | 0.117809 |
| ACC 6h | 40.1644 | 14.5534 | 0.810881 | 0.192597 |
| BABA 6h | 43.1336 | 8.29663 | 0.967006 | 0.180713 |
| Chitin 6h | 42.3694 | 5.10198 | 1.04145 | 0.126774 |
| Epi 6h | 47.538 | 11.4355 | 1.03402 | 0.231346 |
| SA 6h | 29.2432 | 4.50297 | 0.762526 | 0.154614 |
| Me-JA 6h | 34.1351 | 10.7833 | 0.796409 | 0.236338 |
Source Transcript PGSC0003DMT400016509 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G04200.1 | +1 | 0.0 | 872 | 471/706 (67%) | FUNCTIONS IN: molecular_function unknown; INVOLVED IN: N-terminal protein myristoylation; LOCATED IN: cellular_component unknown; EXPRESSED IN: 22 plant structures; EXPRESSED DURING: 13 growth stages; CONTAINS InterPro DOMAIN/s: Dymeclin (InterPro:IPR019142); Has 395 Blast hits to 389 proteins in 117 species: Archae - 0; Bacteria - 0; Metazoa - 262; Fungi - 21; Plants - 68; Viruses - 0; Other Eukaryotes - 44 (source: NCBI BLink). | chr1:1109676-1113875 FORWARD LENGTH=732 |
