Probe CUST_24905_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24905_PI426222305 | JHI_St_60k_v1 | DMT400045162 | CAGCAATCACTCAGATGTTGTGAAAGAGTGGAAGATGGGAAAGTAGAGTGCAAGTGTGCA |
All Microarray Probes Designed to Gene DMG400017517
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24905_PI426222305 | JHI_St_60k_v1 | DMT400045162 | CAGCAATCACTCAGATGTTGTGAAAGAGTGGAAGATGGGAAAGTAGAGTGCAAGTGTGCA |
| CUST_24793_PI426222305 | JHI_St_60k_v1 | DMT400045161 | GGCTTTAGTTTGTTTGTATTGAAGCCTTTGAGTATGTTTCTGTGTTTTCAGTTTGGCTTA |
| CUST_24786_PI426222305 | JHI_St_60k_v1 | DMT400045163 | TTTAGCGGCAATTATCACTCTGTGTATGTCCCTAAAGTCTTTAGCGATACTGGATTTAAT |
Microarray Signals from CUST_24905_PI426222305
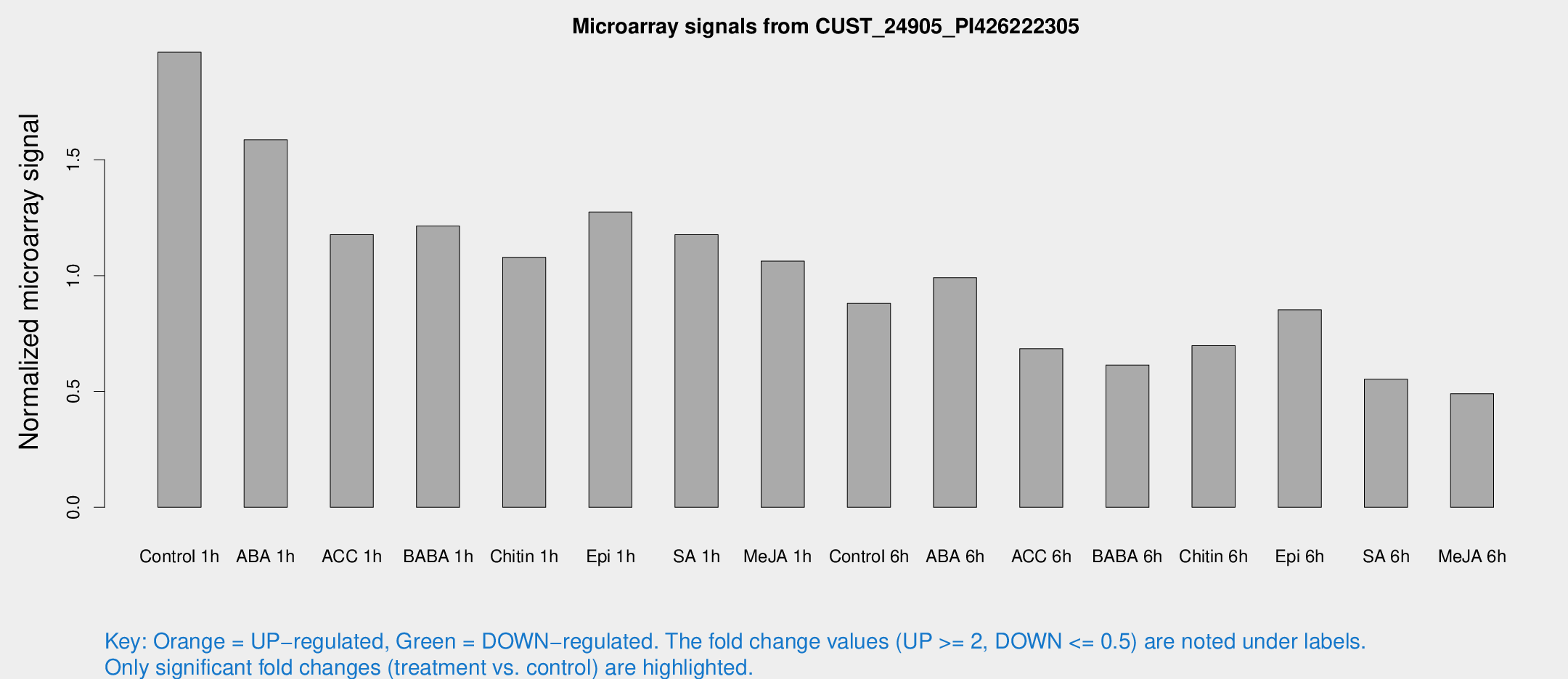
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9501.09 | 791.048 | 1.96402 | 0.113396 |
| ABA 1h | 6864.94 | 888.424 | 1.58593 | 0.156995 |
| ACC 1h | 6078.63 | 1234.94 | 1.17686 | 0.18174 |
| BABA 1h | 5811.06 | 1000.88 | 1.21399 | 0.106312 |
| Chitin 1h | 4726.57 | 576.936 | 1.07893 | 0.0813171 |
| Epi 1h | 5322.1 | 362.727 | 1.27419 | 0.0907425 |
| SA 1h | 5893.48 | 696.55 | 1.17657 | 0.076648 |
| Me-JA 1h | 4175.85 | 241.631 | 1.06261 | 0.0647223 |
| Control 6h | 4322.84 | 626.55 | 0.879961 | 0.0969336 |
| ABA 6h | 5223.63 | 867.55 | 0.991037 | 0.147829 |
| ACC 6h | 3832.15 | 474.947 | 0.684056 | 0.0557048 |
| BABA 6h | 3366.26 | 483.262 | 0.61411 | 0.0817736 |
| Chitin 6h | 3607.08 | 360.228 | 0.697866 | 0.0736896 |
| Epi 6h | 4625.42 | 267.499 | 0.852697 | 0.074196 |
| SA 6h | 2933.19 | 833.7 | 0.552817 | 0.137376 |
| Me-JA 6h | 2471.23 | 583.215 | 0.490285 | 0.0737542 |
Source Transcript PGSC0003DMT400045162 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
