Probe CUST_24786_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24786_PI426222305 | JHI_St_60k_v1 | DMT400045163 | TTTAGCGGCAATTATCACTCTGTGTATGTCCCTAAAGTCTTTAGCGATACTGGATTTAAT |
All Microarray Probes Designed to Gene DMG400017517
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24905_PI426222305 | JHI_St_60k_v1 | DMT400045162 | CAGCAATCACTCAGATGTTGTGAAAGAGTGGAAGATGGGAAAGTAGAGTGCAAGTGTGCA |
| CUST_24793_PI426222305 | JHI_St_60k_v1 | DMT400045161 | GGCTTTAGTTTGTTTGTATTGAAGCCTTTGAGTATGTTTCTGTGTTTTCAGTTTGGCTTA |
| CUST_24786_PI426222305 | JHI_St_60k_v1 | DMT400045163 | TTTAGCGGCAATTATCACTCTGTGTATGTCCCTAAAGTCTTTAGCGATACTGGATTTAAT |
Microarray Signals from CUST_24786_PI426222305
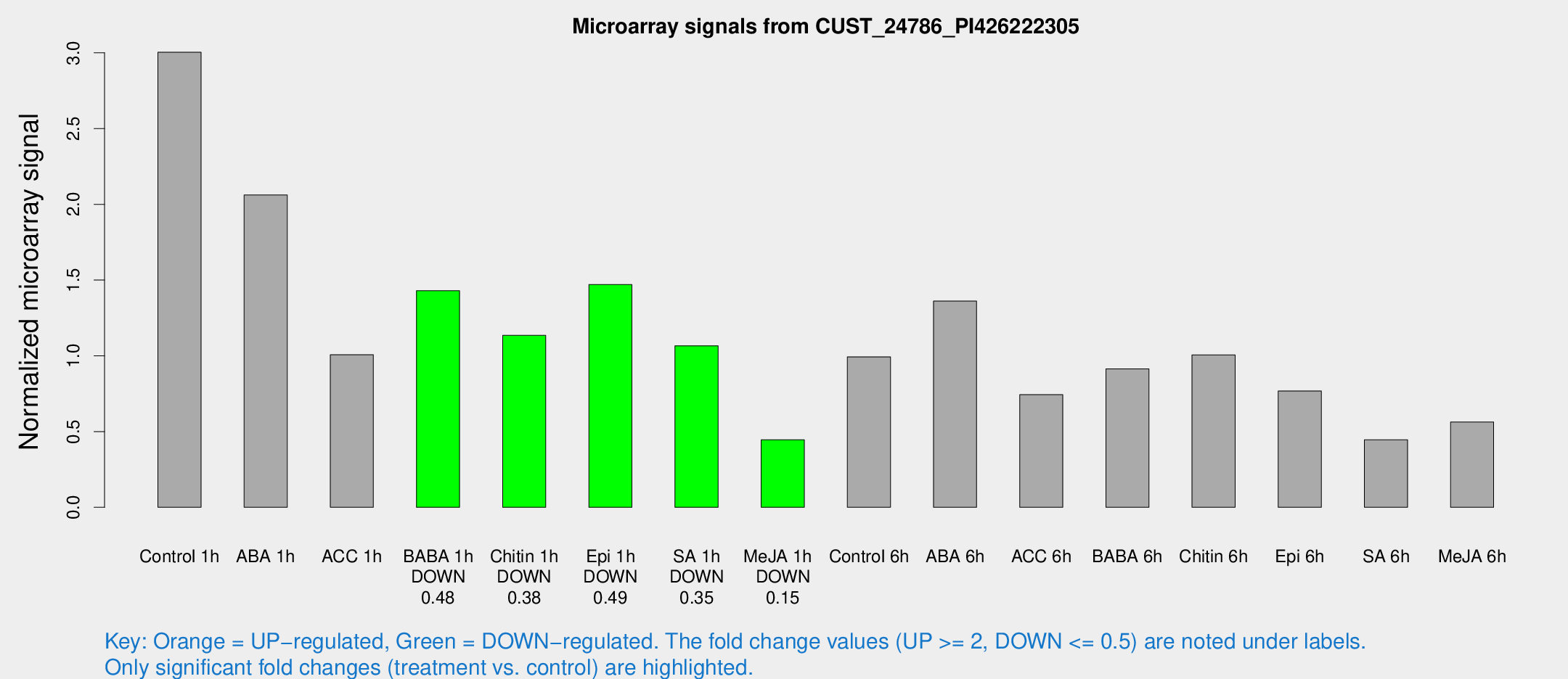
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 228.188 | 23.0925 | 3.00418 | 0.180458 |
| ABA 1h | 142.824 | 26.6776 | 2.06184 | 0.345873 |
| ACC 1h | 86.4211 | 24.2846 | 1.00696 | 0.285197 |
| BABA 1h | 110.696 | 25.3688 | 1.42932 | 0.241804 |
| Chitin 1h | 78.0385 | 10.0642 | 1.13486 | 0.138765 |
| Epi 1h | 97.5187 | 13.5726 | 1.47102 | 0.206409 |
| SA 1h | 82.3753 | 5.76088 | 1.06612 | 0.0744804 |
| Me-JA 1h | 28.2673 | 4.83685 | 0.446177 | 0.0790344 |
| Control 6h | 78.2982 | 17.0643 | 0.993457 | 0.169028 |
| ABA 6h | 122.604 | 35.6547 | 1.36149 | 0.489161 |
| ACC 6h | 65.301 | 8.36205 | 0.743584 | 0.100693 |
| BABA 6h | 80.9918 | 17.1347 | 0.913845 | 0.191273 |
| Chitin 6h | 81.8501 | 10.6148 | 1.00641 | 0.107675 |
| Epi 6h | 65.9931 | 7.87991 | 0.768113 | 0.111213 |
| SA 6h | 44.2523 | 18.5155 | 0.445483 | 0.263216 |
| Me-JA 6h | 51.2159 | 18.0019 | 0.563011 | 0.262204 |
Source Transcript PGSC0003DMT400045163 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
