Probe CUST_24793_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24793_PI426222305 | JHI_St_60k_v1 | DMT400045161 | GGCTTTAGTTTGTTTGTATTGAAGCCTTTGAGTATGTTTCTGTGTTTTCAGTTTGGCTTA |
All Microarray Probes Designed to Gene DMG400017517
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_24905_PI426222305 | JHI_St_60k_v1 | DMT400045162 | CAGCAATCACTCAGATGTTGTGAAAGAGTGGAAGATGGGAAAGTAGAGTGCAAGTGTGCA |
| CUST_24793_PI426222305 | JHI_St_60k_v1 | DMT400045161 | GGCTTTAGTTTGTTTGTATTGAAGCCTTTGAGTATGTTTCTGTGTTTTCAGTTTGGCTTA |
| CUST_24786_PI426222305 | JHI_St_60k_v1 | DMT400045163 | TTTAGCGGCAATTATCACTCTGTGTATGTCCCTAAAGTCTTTAGCGATACTGGATTTAAT |
Microarray Signals from CUST_24793_PI426222305
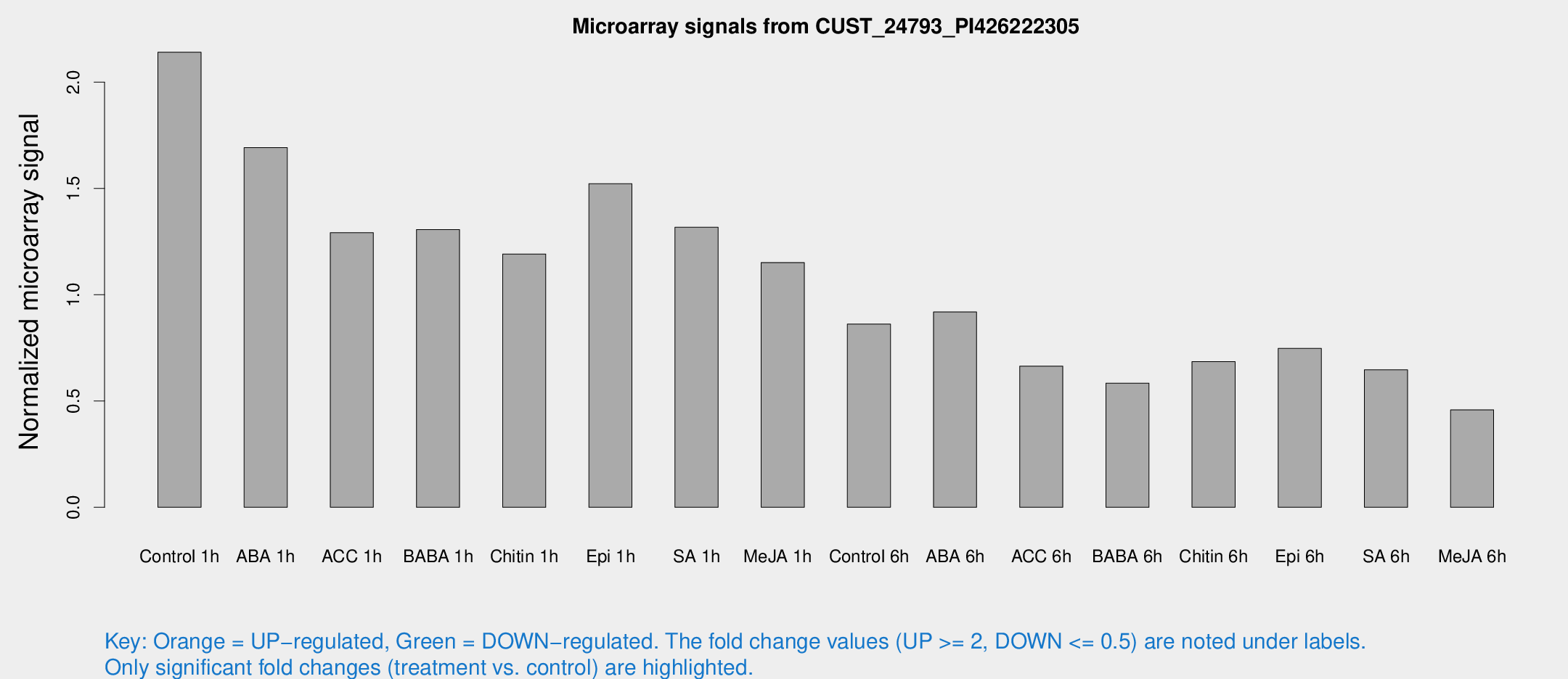
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 49405 | 3058.1 | 2.14107 | 0.123615 |
| ABA 1h | 34886.8 | 3944.57 | 1.69206 | 0.12743 |
| ACC 1h | 32565.2 | 7678.8 | 1.29167 | 0.259398 |
| BABA 1h | 30552.8 | 6538.45 | 1.30687 | 0.183641 |
| Chitin 1h | 24867.4 | 2499.92 | 1.19175 | 0.0719024 |
| Epi 1h | 30415.4 | 1999.3 | 1.52243 | 0.103237 |
| SA 1h | 31746.2 | 4272.07 | 1.31761 | 0.131439 |
| Me-JA 1h | 21696 | 1344.81 | 1.15169 | 0.0664931 |
| Control 6h | 20149 | 2397.84 | 0.86232 | 0.0647122 |
| ABA 6h | 22936.5 | 3229.38 | 0.919027 | 0.0876069 |
| ACC 6h | 17714.8 | 1811.21 | 0.664293 | 0.0741214 |
| BABA 6h | 15185.3 | 1589.72 | 0.58412 | 0.0627322 |
| Chitin 6h | 16948.4 | 1739.92 | 0.685075 | 0.0700533 |
| Epi 6h | 19376 | 1119.1 | 0.747414 | 0.0519919 |
| SA 6h | 15852.7 | 3822.35 | 0.646674 | 0.112983 |
| Me-JA 6h | 11056.6 | 2623.58 | 0.458476 | 0.0697542 |
Source Transcript PGSC0003DMT400045161 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
