Probe CUST_22041_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22041_PI426222305 | JHI_St_60k_v1 | DMT400041941 | CAGTAGCATGTATCCAAACCCATAGCCGTGTATGTGTTTATTTGTTCAACTTTTACAAAT |
All Microarray Probes Designed to Gene DMG400016268
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22047_PI426222305 | JHI_St_60k_v1 | DMT400041940 | CCTCAATTATGTGGAGTCAAGGAGTGTGTCACATAAGCCAAAAAGTGTGCAAAATTACAA |
| CUST_22041_PI426222305 | JHI_St_60k_v1 | DMT400041941 | CAGTAGCATGTATCCAAACCCATAGCCGTGTATGTGTTTATTTGTTCAACTTTTACAAAT |
Microarray Signals from CUST_22041_PI426222305
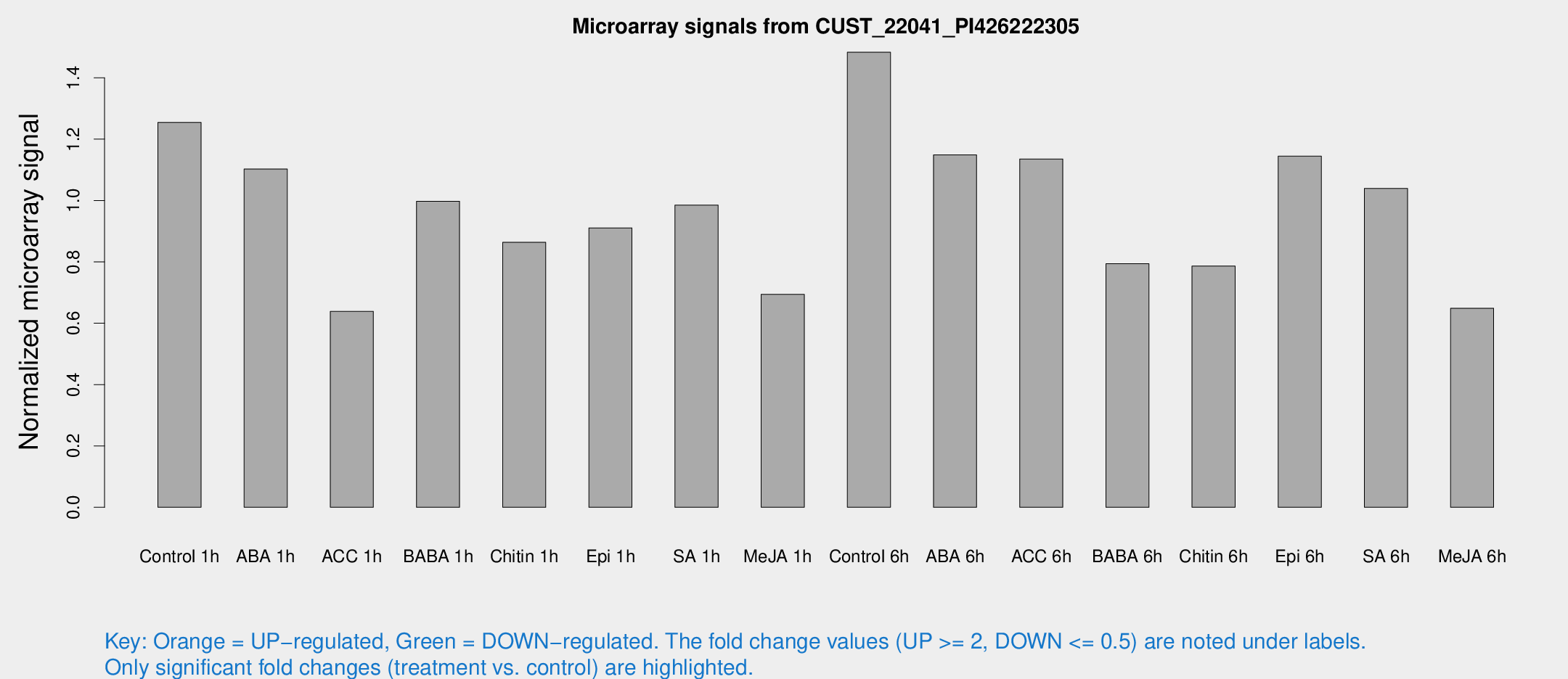
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3575.32 | 233.761 | 1.25467 | 0.0724476 |
| ABA 1h | 2783.31 | 204.528 | 1.10258 | 0.0724548 |
| ACC 1h | 2244.05 | 788.147 | 0.638565 | 0.289078 |
| BABA 1h | 2872.1 | 598.425 | 0.997625 | 0.133195 |
| Chitin 1h | 2243.05 | 297.278 | 0.863812 | 0.0949091 |
| Epi 1h | 2303.83 | 374.845 | 0.910831 | 0.154635 |
| SA 1h | 2928.73 | 390.28 | 0.98501 | 0.0998673 |
| Me-JA 1h | 1621.09 | 146.817 | 0.694098 | 0.0400998 |
| Control 6h | 4324.61 | 655.524 | 1.48341 | 0.129975 |
| ABA 6h | 3481.61 | 240.126 | 1.14855 | 0.0663216 |
| ACC 6h | 3711.72 | 235.257 | 1.13551 | 0.0927576 |
| BABA 6h | 2563.4 | 338.16 | 0.793873 | 0.101732 |
| Chitin 6h | 2382.41 | 137.959 | 0.786535 | 0.0454272 |
| Epi 6h | 3674 | 212.777 | 1.14466 | 0.0660981 |
| SA 6h | 3155.04 | 773.417 | 1.03945 | 0.188341 |
| Me-JA 6h | 1921.66 | 435.32 | 0.64884 | 0.0954136 |
Source Transcript PGSC0003DMT400041941 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G43860.1 | +3 | 3e-130 | 381 | 183/310 (59%) | chlorophyllase 2 | chr5:17630492-17632184 FORWARD LENGTH=318 |
