Probe CUST_22047_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22047_PI426222305 | JHI_St_60k_v1 | DMT400041940 | CCTCAATTATGTGGAGTCAAGGAGTGTGTCACATAAGCCAAAAAGTGTGCAAAATTACAA |
All Microarray Probes Designed to Gene DMG400016268
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_22047_PI426222305 | JHI_St_60k_v1 | DMT400041940 | CCTCAATTATGTGGAGTCAAGGAGTGTGTCACATAAGCCAAAAAGTGTGCAAAATTACAA |
| CUST_22041_PI426222305 | JHI_St_60k_v1 | DMT400041941 | CAGTAGCATGTATCCAAACCCATAGCCGTGTATGTGTTTATTTGTTCAACTTTTACAAAT |
Microarray Signals from CUST_22047_PI426222305
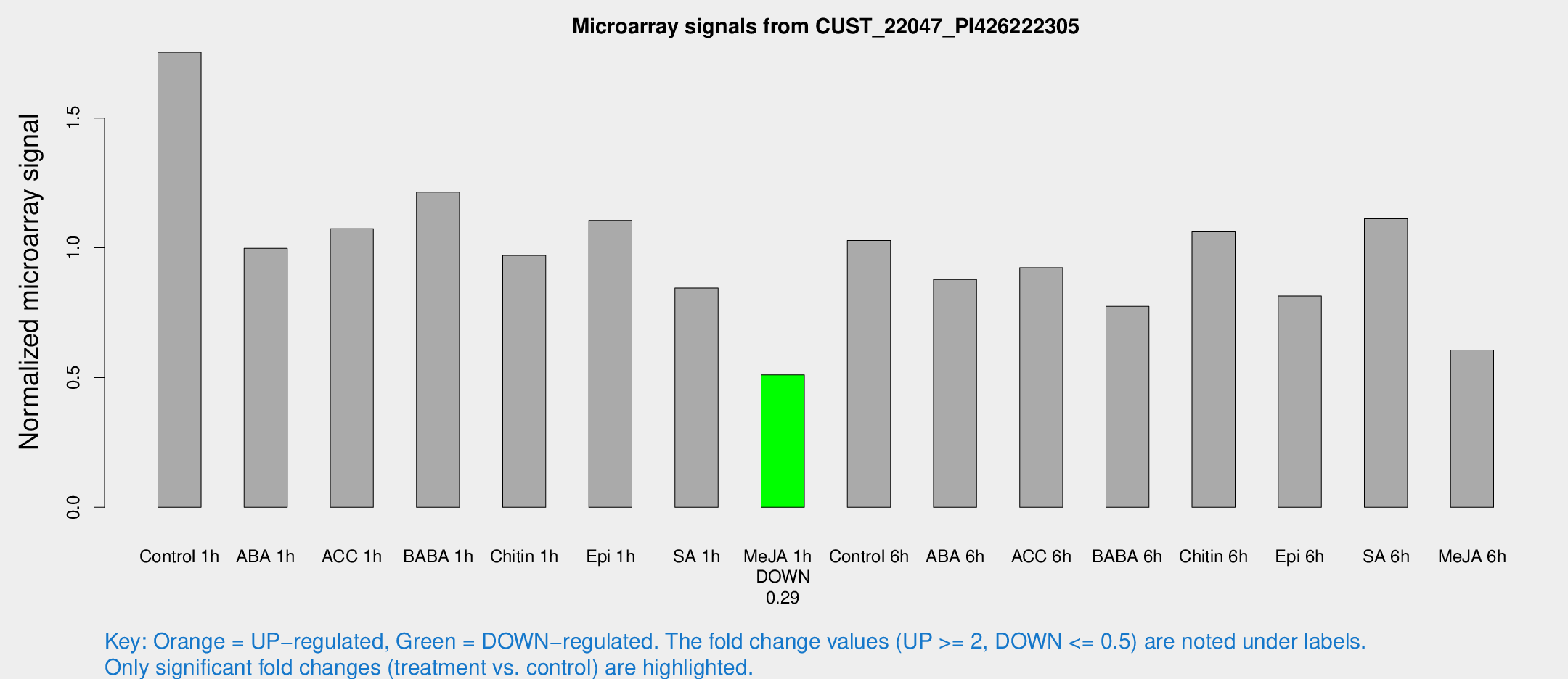
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 31.4728 | 3.79836 | 1.75365 | 0.212767 |
| ABA 1h | 18.4177 | 6.66761 | 0.998241 | 0.530616 |
| ACC 1h | 20.9479 | 4.73831 | 1.07375 | 0.227492 |
| BABA 1h | 25.4776 | 9.52569 | 1.21495 | 0.508204 |
| Chitin 1h | 20.2983 | 8.33849 | 0.970734 | 0.579553 |
| Epi 1h | 18.2147 | 4.05925 | 1.10602 | 0.332792 |
| SA 1h | 18.9128 | 7.2499 | 0.844959 | 0.425467 |
| Me-JA 1h | 7.63398 | 3.48869 | 0.510368 | 0.240801 |
| Control 6h | 23.7581 | 9.39094 | 1.02828 | 0.57833 |
| ABA 6h | 18.4497 | 5.49118 | 0.878158 | 0.247072 |
| ACC 6h | 20.0296 | 4.43106 | 0.923643 | 0.251649 |
| BABA 6h | 16.3756 | 4.01191 | 0.774393 | 0.215297 |
| Chitin 6h | 22.3087 | 6.23411 | 1.06158 | 0.389893 |
| Epi 6h | 18.5038 | 6.06057 | 0.814315 | 0.269741 |
| SA 6h | 25.8491 | 13.81 | 1.11225 | 0.952863 |
| Me-JA 6h | 11.6933 | 3.51606 | 0.606449 | 0.224812 |
Source Transcript PGSC0003DMT400041940 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G43860.1 | +3 | 5e-20 | 85 | 42/79 (53%) | chlorophyllase 2 | chr5:17630492-17632184 FORWARD LENGTH=318 |
