Probe CUST_12612_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12612_PI426222305 | JHI_St_60k_v1 | DMT400063775 | GCCCCATTGTACATTTTCTGTCATTACACTTCGTGGTATATAAACATATAGCTGTACATT |
All Microarray Probes Designed to Gene DMG400024785
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12612_PI426222305 | JHI_St_60k_v1 | DMT400063775 | GCCCCATTGTACATTTTCTGTCATTACACTTCGTGGTATATAAACATATAGCTGTACATT |
| CUST_12318_PI426222305 | JHI_St_60k_v1 | DMT400063777 | GCCCCATTGTACATTTTCTGTCATTACACTTCGTGGTATATAAACATATAGCTGTACATT |
| CUST_12536_PI426222305 | JHI_St_60k_v1 | DMT400063776 | GCCCCATTGTACATTTTCTGTCATTACACTTCGTGGTATATAAACATATAGCTGTACATT |
Microarray Signals from CUST_12612_PI426222305
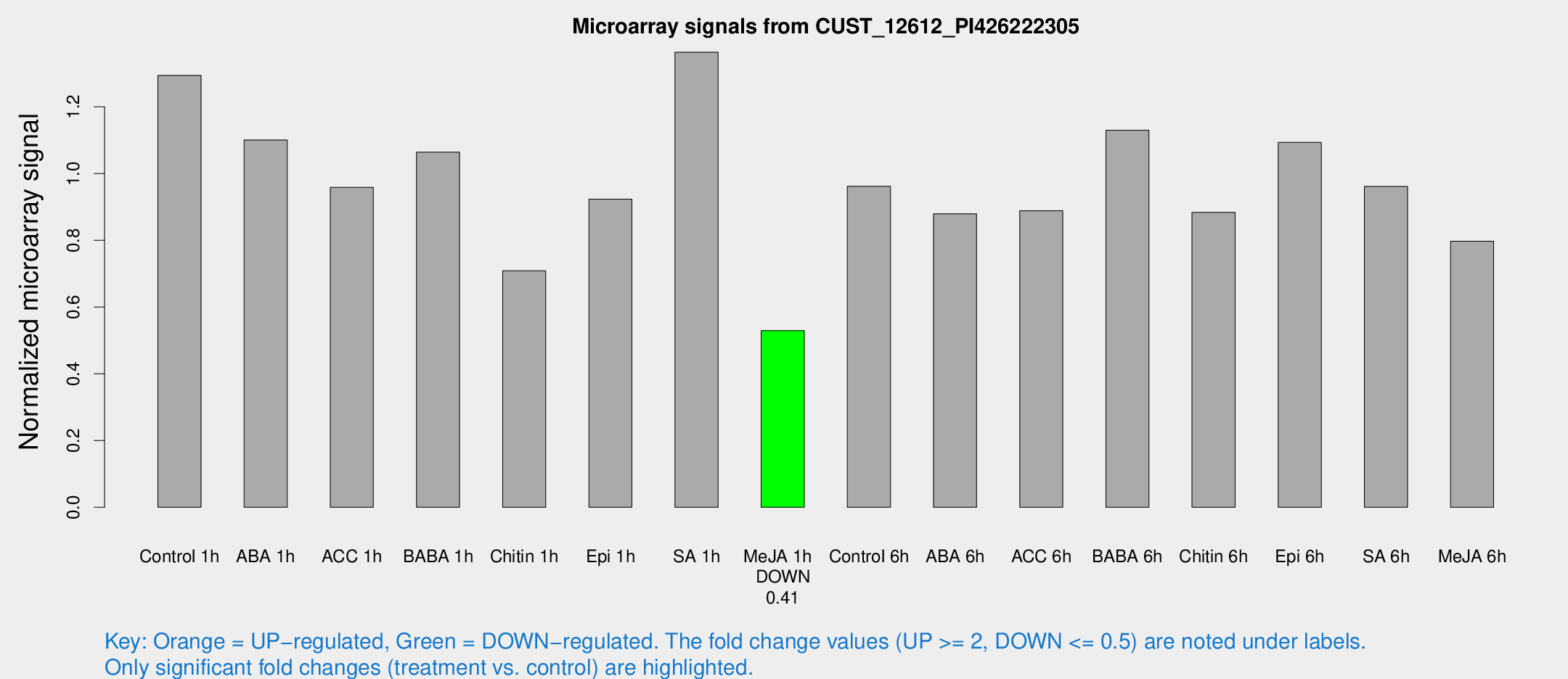
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 399.799 | 73.7928 | 1.29392 | 0.150305 |
| ABA 1h | 291.542 | 17.0586 | 1.10022 | 0.0998862 |
| ACC 1h | 354.105 | 120.223 | 0.958761 | 0.428915 |
| BABA 1h | 312.709 | 43.0464 | 1.06413 | 0.101425 |
| Chitin 1h | 193.72 | 24.7425 | 0.708404 | 0.0422374 |
| Epi 1h | 241.629 | 23.1474 | 0.923344 | 0.0741342 |
| SA 1h | 426.69 | 54.8878 | 1.36334 | 0.143864 |
| Me-JA 1h | 130.715 | 12.9914 | 0.52939 | 0.0327884 |
| Control 6h | 301.62 | 57.8945 | 0.961595 | 0.119671 |
| ABA 6h | 287.811 | 49.4373 | 0.87946 | 0.0979775 |
| ACC 6h | 305.927 | 18.0592 | 0.888269 | 0.119674 |
| BABA 6h | 380.075 | 23.8875 | 1.1296 | 0.0705159 |
| Chitin 6h | 282.276 | 16.6564 | 0.883499 | 0.0520294 |
| Epi 6h | 374.494 | 46.1957 | 1.09346 | 0.154266 |
| SA 6h | 291.651 | 42.4274 | 0.961306 | 0.0566231 |
| Me-JA 6h | 255.027 | 65.6601 | 0.797411 | 0.149745 |
Source Transcript PGSC0003DMT400063775 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
