- Transcript BART1_0-u57237.003
- Transcript IDBART1_0-u57237.003
- Gene IDBART1_0-u57237
Exon Structure of BART1_0-u57237.003
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr7H | 1 | 653252718 | 653252307 | - |
| chr7H | 2 | 653255206 | 653252838 | - |
| chr7H | 3 | 653255658 | 653255311 | - |
| chr7H | 4 | 653255899 | 653255838 | - |
| chr7H | 5 | 653256103 | 653255992 | - |
| chr7H | 6 | 653256324 | 653256220 | - |
| chr7H | 7 | 653259048 | 653257149 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials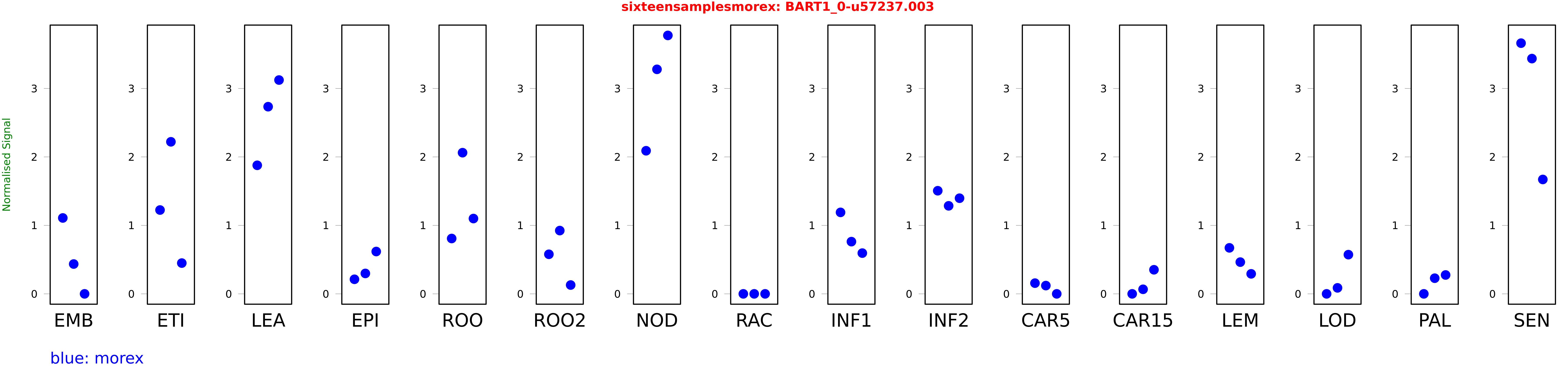
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 1.10839 | 0.435645 | 0 |
| morex | ETI | 1.22444 | 2.22067 | 0.449452 |
| morex | LEA | 1.87757 | 2.73339 | 3.12276 |
| morex | EPI | 0.212656 | 0.29826 | 0.618288 |
| morex | ROO | 0.808422 | 2.0618 | 1.09995 |
| morex | ROO2 | 0.577954 | 0.924161 | 0.128823 |
| morex | NOD | 2.08997 | 3.27924 | 3.7744 |
| morex | RAC | 0 | 0 | 0 |
| morex | INF1 | 1.18999 | 0.762725 | 0.595028 |
| morex | INF2 | 1.50602 | 1.28556 | 1.3969 |
| morex | CAR5 | 0.156457 | 0.120724 | 0 |
| morex | CAR15 | 0 | 0.0666183 | 0.351803 |
| morex | LEM | 0.671852 | 0.463562 | 0.292227 |
| morex | LOD | 0 | 0.0874998 | 0.572522 |
| morex | PAL | 0 | 0.228723 | 0.275746 |
| morex | SEN | 3.66151 | 3.43607 | 1.67076 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os06g51260.1 | +2 | 1e-119 | 386 | 292/518 (56%) | protein|MYB family transcription factor, putative, expressed |
| TAIR PP10 | AT5G37260 @ ARAPORT AT5G37260.1 @ TAIR |
+2 | 2e-34 | 134 | 101/238 (42%) | Symbols: RVE2, CIR1 | Homeodomain-like superfamily protein | chr5:14751344-14752972 REVERSE LENGTH=287 |
| BRACH PP3 | Bradi1g29680.1.p | +2 | 7e-161 | 498 | 316/508 (62%) | pacid=32806206 transcript=Bradi1g29680.1 locus=Bradi1g29680 ID=Bradi1g29680.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (5308 bp)
>BART1_0-u57237.003 5308 150831_barley_pseudomolecules CTCGTGATATATGGTCAGTTAAAATAGCAGTTGTCCGTGCCTCTCCAATTGTGTTGATTT ATTATGTTACTCCTTTTGTGGGTGTAGAGTGAAGATTGTCCGTTAAATACTTACTAAGTC TCTCAAACATGAGGAAGCGACAAGAAAAAATGTGGAGAAGCTTCTCGTCACCATGCGATC AAAAATATGTAACATCACAACTCATATATATGTAGGAGTATCTGTCAATTAAATGGACAA TAAAATGTCTTAATTAATTTATGTGGCAGAAAAAGGAGGGCAGCCTAGATTTAGACAAGC CGAGATATGGAAAGGCCACTTGCAAGCCATGTGTCTTGGAGTTGCACTTATTCTAAAATT CACAACCGTCAGCACGATTTATCCAACCTCCCAATTCAACGATGTCGAGCAAGACGTATT GCTCTTAAATCTGTTTGGATGATTCCCCTTCTTTTCCAAAAATATACTCATGATATGTAT GGTTTATTTTGGGACCGGAGAAAATATGACATGCACATTACTAATTTTTGCAAAAATGAT TCCTTTTTTCTGGAGCGCATGGTTCCTATCCGAGACGTACGAAGTCTCTCCAATAAGATC ATGAAATTATCAAGCTAAATCCAGAAGAGATTCTTTTATTGGCCCCACACCAAGGGCTAA AAATGTAACATCGTCATTGATATCTACTCGACAATTATATTCATAGAAATGTGCTTCAAA AAATATGTGGACAAGACGAGAGGAACCTAGAGTTAGAGACACATTGAGAAAGGGAAAGGC CATTCATGGCATGATTACATAATTGGCCTTATCTAAAAGTTACGCGTGCTGTTATACTTT ATATATATCATATACCCTAGAAATATCTCCACGGGTTCGCTCCTGGCCTCTCATGTTCGA ATGGCTCTAGATACTCGTGGCTCTCTTCCGACTCGTTCCTCTCTCTTTGTTTCCACTTGT GCAAAACACCGAGACCCCCACGCGTTGTCACGTGGATTATTCTCGCGTACTACAATAACA ACCGTAATGCAACAACAATGTCCCATCAAATAAATTGTGTATGAGCCAACAAACCATAGG AAAACAATTCTCTGGTATTACAAACCGTATTACAAATCGCATCAGGATGATTACATATAA AATCGCACTCAAATCGTACCACATCATCGTTTCAAATTTACGTCGAAGAAAAACACAACT AGTGAAATAATGTAAACTTATTTTTGAGCTACTAGTAAGAAAATACACTTGTATATGGTG AGTAGATAAGGGAGAAAGCGCATCCAATGGGAAGCAAAGCCAGAAAAGGAAAGGGCGCGT CGGTCGCTGCATGTGGGCCCGGGGAGGCCGGGGCGTCCTCAGCCACGAGCTGGCCGGCGC TTCTGGAAATTTCATGGCTCCGACAGGTGGGACCCACACATGACGTGGCCACCCTCCCCG CCCCTCCTCGCGAAAATATATTTCCGTCGCTGCTCCACCCCACCTTCCTCCTCCTCCTCC GCCTCCGCTTCCTCCTCCTCCGTCTCCCCCGGCAGCACGTGCGCGCCCCGGCGGCCTCCT CTCTTAGCTGCGCCGCTCGCCGGAGCGTCTCTTCTCGTTTGGCTTCTTTTCTTTGGGTCA GTTTCCTTGGGCCACGGTGTCTCGCCGGCCGGCTCCGGTCGCCGAGAGGGAGGGTGTTTT TTCGGTGCTTCCTGGGCGATCCCGGCGAGAGACGCTTCTGTCCCAGCCTCCGCCGCAGGC CGCCCGCTTCTGTCCCCGCCTTTGCCGCCCGCCGTCTGCTTCTGTCCCCGCCTTCGCCGT CCGCCGTCTGCCGTCTGCTTCGTTCGATCTCGGTGACGGGGACGCCGAGATTCGGACCTC CGCCGGGGAACGGCTCATGGCTTCCATGGCGCTCGCGGAGGAAACGGATGCACTGGATTC ATCCAGATTGCCGAACGGCAAAGGGTTATCGGTGGATGTTGCAATGCAACCCAATGAGGA GGGGATGGGTGAGCATCCTGTCAAGCCTAGGAAGCCATACACGATCACAAAGCAGCGGGA GAAGTGGACGGACGAAGAGCACGAGAAGTTCCTGGAGGCGCTGAAGCTGTATGGCCGGTC CTGGCGTCAGATACAAGAACACATTGGCACAAAGACCGCTGTCCAAATCCGAAGCCATGC CCAGAAGTTTTTCTCCAAGGTGGTGCGCGAACCCGGAGCTAAAATTGAGATCGAGATCCC TCCCCCTAGGCCAAAGAGGAAGCCGCTGCATCCATATCCTCGCAAGCGTGCGGACTCCTG CAACGGGGCAAACCCAGCAAATGGACCATCCAAGCTTGCTCAAATTTCATCATCTTCTGG TTCTGACCAAGAGAATGGTTCTCCCGTGTCAGTGTTGTCTGCAATGCAGTCGGATGCTTT TGGATCATCGATGTCAAACCCGTCATCTCGCAGTACTTCCCCAGAGTCGTCAGATGAGGA GAATAATGTTCCTCCAATGGTCAGCAGAGAGGAAGGTCAACAAACAGGGATAAATCAATC CCACAAGGAGGCTGATCAGGAAAACAAAGATACGGGTACCTCCGAAGAGGATTCTTCTGA TGAGGTGCAGGTAACAAGCGTGAAGCTGTTCGGGAAGACGGTCGTCATCCCAGACCCGAG GAAAAGGTGCTCCCCGGATCCAGGCTCAGGGCATGAAAACGGAGGGCAAACCTCACATTC CTCTAACAAAGGAACATCACAAGCCCCACTGGCTGTAGAGATTCCAACACATACAAAGGG AGAACAAATCTCACAATCCTCTAACAAGGCAACATCACAAGCCCCATTGGCTGTAGAGGT TCCGACGTATACAAAGGGAGAGCAAATCTCACAATTCTCTAACAAGGCAACTTCACAAGC CCCACTGGCTGTAGAAATTCCAGCATTTGCTGCGCCGCCAAGTGGGTGGGTTCTTCCGTA CAATTCGTTCCCTCTACATTTTGGGGAATCGGCGGAAGCCAGAATCACCCCTTTACACAT GTGGTGGCCATACTATGGGTTCCCTATCAGCCACCCTGGAGGGCTGAGTGTAGTGCCGCA CAACGAAGCTACTGATGAGAGCGAAGCTGCAAAGAGCCCTCTGGTTGAATCAAGCTCGAA GAGCCTTCCTGTTGAATCAAGCTCGGACTCCGATGACAATGCTGAGACAACAGCCAACAA GGAGTGGAAAGTGCTCGAGTCGCTCGGGACGGCACAGGTCCCTCGGTCAGCTTCAAGTTT TCAGCTAAAACCAAGCGAGAACTCGGCTTTTGTGAGAGTGAAGCCGATCATAGGCAGCGG AGAAGAGCCGGCGAGGGGGTTTGTGCCATACAAAAGATGTAGAGTTGAATGACGAAACCG GGGATCTTCTTCCCCGTGTCTAATTTAACTATGAACCCTAGGAGGAGCCTGTTCCACATA TTCCTTTGCTCCGGTGTATTCTTAGGTTTTACTCGCACACAGAGATTGGCGGTGGTCTGG CCCAGGAAATGCACGTTGTTTCCTGCTTGGGCCTTGTAATGAGCAGGGCACAGGTGGTTG TTGTAAGTGCTGTAGGATAAAAGCAGGAGGTTTTGTAGGGATGCCAATTCTCTGTCTGTC TCATGTTGTTGATGTAAAAGCAAAGAATTTTTGTGTCACCATTGTTGATTCTTAATGGAA ATCGACTTCTCTGAATGAATGAAGCTGGCATGTTTGGTTGTTGCGGGCTGTTGTGTTGGT AAGAGTCTGAAGATGCTGGTGCCTTTTGTCCCGAGCATGCAGGCAGAGCTTGGCCGTGTT CTTCAGACTGGAGTATGGAAGGAAACATTGTTCATCATTGGTAGCAATTGCATGCCTTTG AAGGTTTCCAGCCATGGCTGCTAAATTCAATTCCATCAAGTGGGACATTATGATATTTGT CTGAACATTGCAGTTGCCGAACATTCCAATGTTTGCCGCAACTGTTGGCTAGCAATGTCT GACCATTCAATTCTGTCAAGCAAACAGCAGAGGCTGCTTAGTGACTGGGAATTGTACGGG ACATAAATCGTGAGTTACTGGGATAAACATTAAGGCTTGTTTTTTGTACTTGTATATATC CAGAGGATGCAAATATCTCATGTGAAGAACCGAAATGGAAATGGATAGCTAAAGGATCCA TTGCGACAAAACGGGTAAATTTGTATCAGGAAGCAGAGAGTTGTTTTTAATGAGCTTGTC AACAGAAGCTTGATCCAGCTTGCTGATGTTAATGTTGACTATTATGATTTTAACAACATT GTCAGGTTCAATCAGATGAAGCGAATTTTGCCACAGTTTAAAACCGTGCCTGCAATCCTT TGCCAAACAAGATCCGTCGACTATCCCTGCAATCCTGTGGTTTGAACATAAATTGACAAT CCATGCCATCACAGGAAGCAAGTTACATATTCGCTCATGGAATGCCTTTGCACGTGTTTG ACATATGTGCGGGTCATTCGTCACCCGAAACACAAACATGTAAGAAATATAAGAAGTTTT CTATCAACTGATGTATTTGAGAATTCAGAGTGAAGGAATCACATAGCTTCCTGAAGATAT TAGAAAGCTACAGAACTTAGAGACTCTTGATTTACGGGACAGTTCAGTTGGAAAATTGCC ATCAACCATAGCCCTCCGATGGCTATAAAATTTGGCGTGGCAAAGAAACGGGTATAAAAA TTTGCAGCTGGTGCGACGCCACCACTCCGATGGCTCAACCTTTCCTGGAGTGCAGCTCAA GTTTGGTTGAGATATAGTGATGGGTCTGACATAGGCATCAAACATCTTGCAAGCCTTATG TACATCAAACTTTTTGTGATGATGCAACTCGGGAGGTTGTCTTGAAGATGAGAGGTTTAA GAGTTACGAGAGGGCAATGCGTAGGGAAGGAAGCAGGATATTTGGAAAACACAAGATTGG TATGCTTCAGCAACCAATTGAACTGAACCGGCTGTAGTTGCCTCGAGGTGTCAGGTTCAA ATTTCTGAATTTCAGCTGCACCTTCGTCGACAGCTTCGGTTTGTTCGTAGCATGCTAGCC TAACTGTTTGACATTGGTGTATATATAATTGTGGCAAACTGGCAACCAATAGTTAGTGCC CAGCTCACACCCCAAATTTCACTTCTGAAACTGTGGTAACTACTGCTCAGGTTATGGCAA ACGAACCAGCTTTTCTTCAGCTCTGCAATTGTTTCACTCTCCTCCAAATGAACCTCTGCC CATTGCTCGGCTTGTCAAGACAAATGTACCTTTGGCGGAACAAAGAGGATACTGCTAGTT AAATCAGGAAGCAAGATGCTACCGAAAG
Protein Sequence (491 aa)
>BART1_0-u57237.003 491 150831_barley_pseudomolecules MASMALAEETDALDSSRLPNGKGLSVDVAMQPNEEGMGEHPVKPRKPYTITKQREKWTDE EHEKFLEALKLYGRSWRQIQEHIGTKTAVQIRSHAQKFFSKVVREPGAKIEIEIPPPRPK RKPLHPYPRKRADSCNGANPANGPSKLAQISSSSGSDQENGSPVSVLSAMQSDAFGSSMS NPSSRSTSPESSDEENNVPPMVSREEGQQTGINQSHKEADQENKDTGTSEEDSSDEVQVT SVKLFGKTVVIPDPRKRCSPDPGSGHENGGQTSHSSNKGTSQAPLAVEIPTHTKGEQISQ SSNKATSQAPLAVEVPTYTKGEQISQFSNKATSQAPLAVEIPAFAAPPSGWVLPYNSFPL HFGESAEARITPLHMWWPYYGFPISHPGGLSVVPHNEATDESEAAKSPLVESSSKSLPVE SSSDSDDNAETTANKEWKVLESLGTAQVPRSASSFQLKPSENSAFVRVKPIIGSGEEPAR GFVPYKRCRVE
