- Transcript BART1_0-u53703.028
- Transcript IDBART1_0-u53703.028
- Gene IDBART1_0-u53703
Exon Structure of BART1_0-u53703.028
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr7H | 1 | 385863972 | 385861670 | - |
| chr7H | 2 | 385865757 | 385864078 | - |
| chr7H | 3 | 385865991 | 385865930 | - |
| chr7H | 4 | 385866338 | 385866227 | - |
| chr7H | 5 | 385867084 | 385866409 | - |
| chr7H | 6 | 385867226 | 385867200 | - |
| chr7H | 7 | 385867543 | 385867389 | - |
| chr7H | 8 | 385872394 | 385872313 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials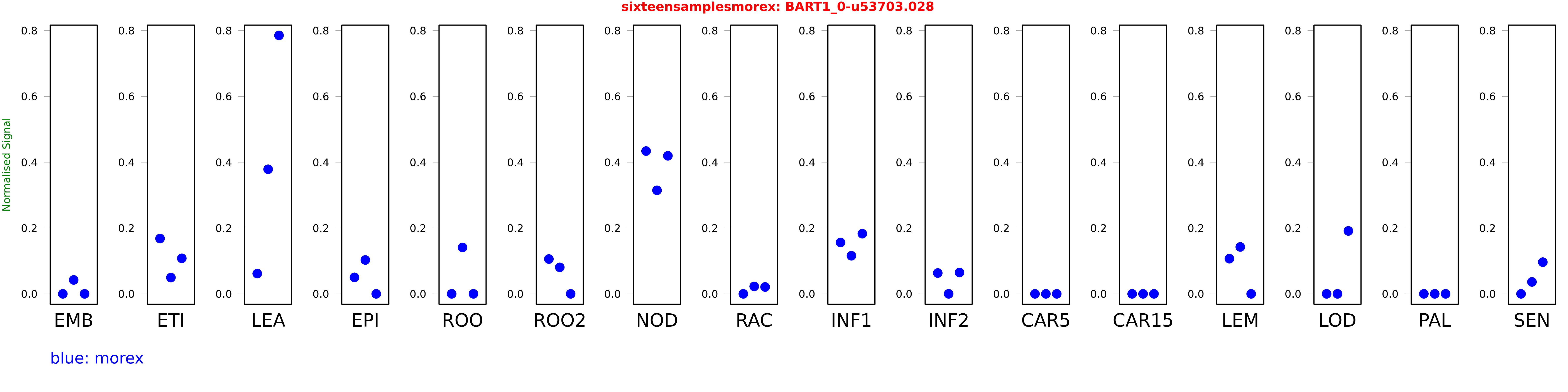
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0 | 0.0422963 | 0 |
| morex | ETI | 0.168199 | 0.0495291 | 0.108 |
| morex | LEA | 0.061637 | 0.378684 | 0.785497 |
| morex | EPI | 0.0502305 | 0.102931 | 0 |
| morex | ROO | 0 | 0.141141 | 0 |
| morex | ROO2 | 0.105829 | 0.0806095 | 0 |
| morex | NOD | 0.434037 | 0.314665 | 0.419613 |
| morex | RAC | 0 | 0.0226542 | 0.0209203 |
| morex | INF1 | 0.156238 | 0.115761 | 0.182822 |
| morex | INF2 | 0.0633392 | 0 | 0.0648996 |
| morex | CAR5 | 0 | 0 | 0 |
| morex | CAR15 | 0 | 0 | 0 |
| morex | LEM | 0.106843 | 0.142617 | 0 |
| morex | LOD | 0 | 0 | 0.19132 |
| morex | PAL | 0 | 0 | 0 |
| morex | SEN | 0 | 0.0362833 | 0.0962875 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os08g06110.2 | +2 | 0.0 | 549 | 369/539 (68%) | protein|MYB family transcription factor, putative, expressed |
| TAIR PP10 | AT1G01060 @ ARAPORT AT1G01060.3 @ TAIR |
+2 | 3e-32 | 135 | 60/72 (83%) | Symbols: LHY, LHY1 | Homeodomain-like superfamily protein | chr1:33992-37061 REVERSE LENGTH=645 |
| BRACH PP3 | Bradi3g16515.1.p | +2 | 0.0 | 597 | 393/532 (74%) | pacid=32817815 transcript=Bradi3g16515.1 locus=Bradi3g16515 ID=Bradi3g16515.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (5097 bp)
>BART1_0-u53703.028 5097 150831_barley_pseudomolecules CTCCCGGTTGGCTTCGTCCGGAAGCATCGGCCGGTGGCGGCAGAGAGAGGAGCTCCTGCG CCCGGCGCGGCGGTCGGTTGAGTAACTTTCAGGTGTAGGAAGTGTTAAGGGGATACATAA TCGGTTTTGAGCTGCGAGGCCCTGCATTTCTTCTGCCTGCTGTTTGATTCACGGCCCACT CTTTTCCTTCTATCCATTCAAGCAAAGTACTGGAATTTCTGAGCCCAATTTATTCAGGTC ATGGGATTTGTTTGTCCTTAGCAGAGTTGAAGATAAAGAGAGGAAGGCTAAACTCTGCTG ATCCGGTTCATGCGAGCAATGAGCCAACTGCAGTAGAGGTTTACAGTAGGTGAGTACAAT GGTAGATCAAGCTATTCTTGTAGCTGTTGAATTCTATATTTTTCCTAATTAATCATGCCT GATTTTAGGATGAAATGAGGTTTGCTGTTCTCTTAAGTACCTGCATATTTAAAATGTTGG AGAGGGCGTAATTTTATTTATTTAAGGGGTGAATATATATTTAAATGTTACACTATTTTG CATATTAACACCAATTGGGTCAAAGAAAATAGGTGAGTGAACTCCTTAGATTATTTCCTT TGATTTTGATCAAAGTTCATTTATGAGTAGACTGGTGAACTCCTTAGATTATTTCTTTGG TCTGTGACAACTAATTTAGCTGCTCTTTTGTGGCTGGGGTCTACAAATTTGAAATATTTA TTGTGTTTGTGTGATAGAGAGGTTTAGGGGGAGGTATGAGATACTACTCGCGTCAGGTCC AGCGTTTGGCTGGAGACACAGAAGGAGGAGTGCTGTTGTTTGTAGACGCCCTCAACTCCA AGTAACAAAGCGACCAGTTGTCAGCACTTCCTTTTGTTTTCGCATTTCCCTGGAATTGGA GATGGAGATAAATTCTTCGGGTGAGGAAACGGTGATAAAGGTGCGAAAGCCGTACACAAT AACAAAACAGCGGGAGCGGTGGACTGAGGCAGAGCACAAACGGTTCCTTGAAGCCCTCAA ACTGTATGGCAGAGCTTGGCAGCGCATAGAAGAGCATGTTGGGACAAAGACGGCCGTGCA AATCAGAAGCCATGCTCAGAAGTTCTTCACAAAGTTTGTTCTGCAGTGCTGAGCATGATG AAAGATTCGTTTCCTTTGAATATCATGAGAACCATGCTTTATCTAATTGTTGCCCTGCTG ATTGATTAAAAGATCAAATTAAAGCATGTTAAAATGGATGAAACATGCTAGAAAGTCTTT GCTGATTGTAAGCAATGAGTAATGTTGCAGAGAAGATGATGTTGAACAACTCTGATTCTC TCATTGATTCATTTTACATATCCATGCTGTTATTGAACATACATATGACTAAGAAAACAT TATGGTACTAGTTAAATTTACATAGAGAGTCAATATAATACATAATGTGAAATAGGATAT CTGAATGCACTCAAGTGAACTACAGGAAATTCCTTAGGAATTGTATGGTAAATATGCTAA CATTTGTTGCATTTCATTATAGCCCCCATCAACGTAGGTAAACACTTTATCTAACATGTT TCACTATCAACATTGTGTAAATGAGTCCATTTCATTTTCTAAATCATGATATTTCTTCCG CAGAAGTACACAATGGCATGAACAAAATTTATCTTCTTTCTCCCCTTTTTGTCCTTGCTC TCTTCTCTGCTTTCTGTTCTTTGCTGCTTGTCCTATTTTTTTAGGGGCTGCATACAGCCA CATGGTGGAGGCTACATTGCCAATTTGCCATCATGTTGTAACATGCAACAGACTGTTGCC TTGTTAGGATTGCAGCCTCATCCTCTCAGAGGTTTCTTTGCCAGACCCATGGAGGAGGGA GGAGGCTAAGGCTAGTCTTTTCTTCCCTTGTTTTGACCACGATGGTTTCTGGGTCCTCAA ATTTCATACTTTGCTCACCAGCCATTACCAATCTGACCTTGCTCAATATAGCAGGTTGGT TGTTGGGATGGACAGCTCTACCCTATTGTACATATACATATATATATAGTAAATGACGCA ACTAACGGTTCAGTACACCAACTTAGCTCATAGTTACTCGCCAATAAGTTTGTAAAAGCT CAGGGTGGAAACTGTCAGCCTTCAAGTTCAATCTAACCAATAGTTTAAAGGTTTTGTGCA TCGGTTGATGGAGAGGCTGGAGAGATGTCTCTTCCATAATCCAAAAACAATTTTACAGTG CTATTTTCTTACCTTTTATGTGGAATAGAGAAGAGCCAGTAGAGTGAAAAAGAAACCACC ATCCACTCTTTCAGTCCTCCACCCGCTTGAGGGTTTAGGCTGTAGAATGCTGTATTTGTC AGGGTTAGGATACCGGAACTGCCCCAAATTCCCATTGTAGGCGGTTCTGGTTTCTTGCAG AACCTGGCCAGTTAGTCTGGTTGCCCAAAAATGCCTTATTTTCTGGTGCTCTGAATCTTC ATTTTCATTTTGCAGATTTCAAATTGAAGATATAAATTACAGACCTATGCAACAATAGAT AGTTCAATTTTCCAATTCTAGTTTATCACCACTGTCTGTCTGTATCTAACTATTTGGGAA ATTGCAGTTGGAAAAGGAAGCTATCAACAATGGTACTTCTCCAGGACAAGCTCATGATAT AGACATACCTCCACCACGGCCTAAAAGAAAACCTAACTGTCCATATCCTCGAAAAGGTTG TCTCAGCTCTGAGACACCCACCAGAGAAGTTCCAAAATCAAGTGTTAGCTTGAGCAATAG CAATTCACAAATGGAAAGCAATGGAACTCTTCAGGTCACCAGCACTCAGAAACTTCAAAG GAAGGAGTTGTCTGGAAACGGCAGTTGCTCAGAAGTTATTAATATCTTTAGAGAAGCACC ATCTGCCTCATTTTCTTCCTCTAACAAGAGCTCTTCAAATCATGGTGTCTCTGGGGGAAT TGAACCGACTAAAACAGAAATCAAAGATATGGCAGCCATGGAAAGGAAATCTACTTCCGT TGATGTGGCGAAGGATGTAAAAGATATTAATGACCAGGAAATGGAAAGGAACAACAGAGT CCACATCAGTTCTAAATATGACCGTTCTCATGAAGATTGTTTGGATAGCTCAATGAAACA CATGCAGTTGAAGCCAAATACTGTGGAGACAACATACACGGGTCAACATGTTGCAAGTGC TCCACTCTACCAAATGAATAAGACTGGGGCAACTGGCACTCCAGACCCTGGAACTGAAGG AAGTCATCCTGATCAAACGAATGATCAAGTGGGAGGAGCTAATGGAAGTATGGACTGCAT CCATCCAACACTTCCCGTGGATCTAAAATTTGGCAGCAGCTCCACAGCGCAGCCCTTTCC CCACAACTATTCAGGCTTTGCACCAACGATGCAATGCCAGTGCAACCAAGATGCCTACAG GTCATCTGTTGATATGTCGTCCACCTTCTCCAACATGCTCGTTTCCACATTGTTATCAAA CCCCACAGTACATGCAGCTGCAAGGCTTGCAGCATCATACTGGCCAGCAGCAGACAGCAA CATCCCTGTCGATCCAAATCAAGGAATTTTTGCTCAGAATGCTCAAGGAAGACATATTGT TTCTCCTCCAAGCATGGCTTCTGTGGTAGCAGCTACAGTTGCTGCGGCTTCGGCATGGTG GGCAACACAAGGTCTTCTCCCTCTTTTTGCTCCCCCCATGGCTTTTCCATTTGTCCCAGT TCCTACCGCTTCCTTTCCCACAGCGGATGTCCAGCGAGCTACAGAGAACTGCCCAGTGGA CAACGCACCAAAGGAATGCCAAGTAGCTCAGGGGCAAGGTCAACCTGAAGCTATGATAGT TGTAGCATCTTCTGGGTCCGGCGAGAGTGGAAAAGGAGAGGTGTGTCCTCACACTGAGTT AAATATATCTCTGGCTGATAAAGCTGAGACAACACCTGCCACAGGAGCTGAAACAAGTGA TGCTTTGGGCAACAAGAAGAAGCAGGATCGCTCTTCATGTGGTTCCAACACACCATCAAG TAGTGATGTAGAGGCAGAACATGTTCCTGAGAACCAAGATCAAGCTAACGACAAGACACA GCAAGCATGTTGCAGTAATTCTTCAGCTGGTGACATGAACCATCGCAGGTTTAGGAACAT TTCAAGCACGAATGATTCATGGAAGGAAGTTTCCGAGGAGGTTGTAGTCTACCAGCATTG CCGAATTTCATTTCTATGTTACTCTTGAGCATCTATCACTGCTTTTCTAATTCACATTTC TGCTGTCAGGGTCGTATGGCTTTCGATAAACTGTTCAGTAGAGGAAAGCTTCCCCAAAGC TTTTCTCCTCCACAAGCAGAAGGATTGAAGGTGGTTCCCAGAGGGGAGCAAGATGAAGCT ACTACGGTGACGGTCGACCTCAACAAGAGTGCTGCAGTTATGGACCATGAACTTGACACA TTGGTTGGGCCAAGAGCTTCCTTTCCCATTGAATTGTCACACCTGAATATGAAATCCCGC CGGACAGGCTTCAAACCTTACAAGAGGTGCTCGGTGGAAGCAAAGGAGAATAGGGTGCCG GCTGCTGACGAGGTTGGTACCAAAAGGATTCGCCTTGACAGCGAACCCTCCACGTGATTT ACTTCCCACGACATGATTACCAGCCTGCACAAGTAGTGTATTTTCAAGAAATTGCGGTAT TTACATCTAAGCTACTATAGGACTTGCCAGTCCTTGCAATGCAGTGTTTCACAGAACTCA ATTTGTTATGTGTTGTATGTTAATGCTTGCCGAGCAGCCCTTTTAGACTGTCAATTAAGC ATTAAGCACCTATGCATGCTACTTTATTGGTAAAAAAGTAATTCTGCTTATTTTTCTGTG ATGAAGGATAATACTTGTAATTTTGCTTCATTTTACTGGATCATGTAGATTTAATTCCCG AAGGGAGAGCATGATCGCCTTGTGTTATTTTTATTTTCTTGAGAAAAATATTGTGCTATG TATATGCACCCTAGATTTGAAGTTTCTTGGATGGTCCAGCTTTCTCAAACAGTGTACTTC TTTTAATTTAAAATAAAATATATGGCTTGTATCTTTAATATCGAAGGTACTCCCTCT
Protein Sequence (485 aa)
>BART1_0-u53703.028 485 150831_barley_pseudomolecules MESNGTLQVTSTQKLQRKELSGNGSCSEVINIFREAPSASFSSSNKSSSNHGVSGGIEPT KTEIKDMAAMERKSTSVDVAKDVKDINDQEMERNNRVHISSKYDRSHEDCLDSSMKHMQL KPNTVETTYTGQHVASAPLYQMNKTGATGTPDPGTEGSHPDQTNDQVGGANGSMDCIHPT LPVDLKFGSSSTAQPFPHNYSGFAPTMQCQCNQDAYRSSVDMSSTFSNMLVSTLLSNPTV HAAARLAASYWPAADSNIPVDPNQGIFAQNAQGRHIVSPPSMASVVAATVAAASAWWATQ GLLPLFAPPMAFPFVPVPTASFPTADVQRATENCPVDNAPKECQVAQGQGQPEAMIVVAS SGSGESGKGEVCPHTELNISLADKAETTPATGAETSDALGNKKKQDRSSCGSNTPSSSDV EAEHVPENQDQANDKTQQACCSNSSAGDMNHRRFRNISSTNDSWKEVSEEVVVYQHCRIS FLCYS
