- Transcript BART1_0-u52907.019
- Transcript IDBART1_0-u52907.019
- Gene IDBART1_0-u52907
Exon Structure of BART1_0-u52907.019
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr7H | 1 | 305303613 | 305303325 | - |
| chr7H | 2 | 305303888 | 305303817 | - |
| chr7H | 3 | 305304132 | 305303990 | - |
| chr7H | 4 | 305304552 | 305304317 | - |
| chr7H | 5 | 305305059 | 305304860 | - |
| chr7H | 6 | 305305504 | 305305214 | - |
| chr7H | 7 | 305306742 | 305306483 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials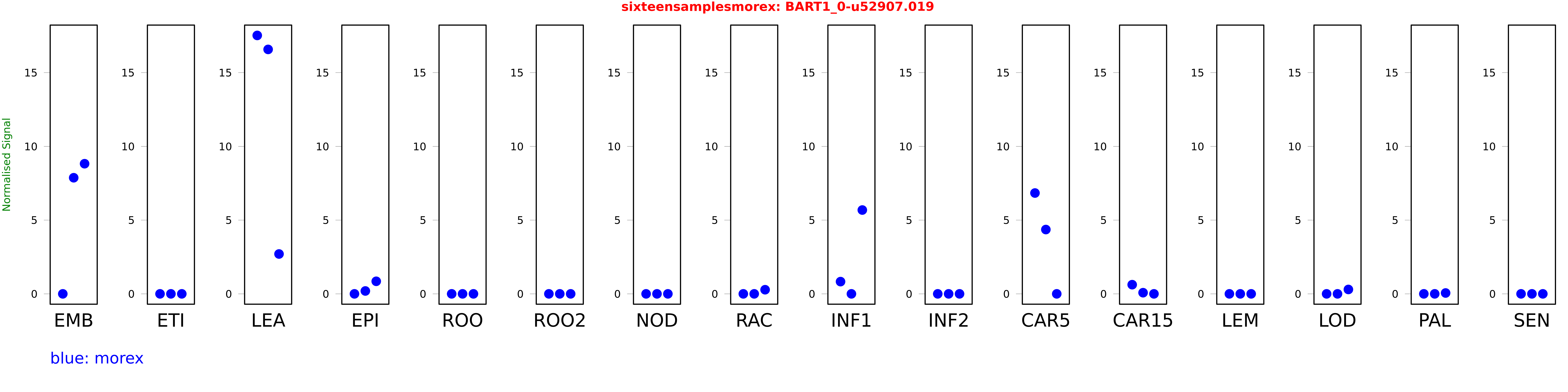
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0 | 7.87394 | 8.82514 |
| morex | ETI | 0 | 0 | 0 |
| morex | LEA | 17.5248 | 16.5763 | 2.70307 |
| morex | EPI | 0 | 0.195738 | 0.851844 |
| morex | ROO | 0 | 0 | 0.00000697418 |
| morex | ROO2 | 0 | 0 | 0 |
| morex | NOD | 0 | 0 | 0 |
| morex | RAC | 0 | 0 | 0.28034 |
| morex | INF1 | 0.830009 | 0 | 5.68429 |
| morex | INF2 | 0 | 0.0000133418 | 0.00000076901 |
| morex | CAR5 | 6.83764 | 4.36131 | 0.0000538032 |
| morex | CAR15 | 0.622065 | 0.078246 | 0 |
| morex | LEM | 0.00000000894794 | 0 | 0 |
| morex | LOD | 0 | 0 | 0.298334 |
| morex | PAL | 0 | 0 | 0.0583762 |
| morex | SEN | 0 | 0 | 0 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os08g29370.1 | +3 | 0.0 | 585 | 283/328 (86%) | protein|peptidyl-prolyl cis-trans isomerase, chloroplast precursor, putative, expressed |
| TAIR PP10 | AT3G01480 @ ARAPORT AT3G01480.1 @ TAIR |
+3 | 5e-175 | 499 | 241/329 (73%) | Symbols: CYP38, ATCYP38 | cyclophilin 38 | chr3:188569-190674 FORWARD LENGTH=437 |
| BRACH PP3 | Bradi3g34940.1.p | +3 | 0.0 | 622 | 301/328 (92%) | pacid=32818759 transcript=Bradi3g34940.1 locus=Bradi3g34940 ID=Bradi3g34940.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1491 bp)
>BART1_0-u52907.019 1491 150831_barley_pseudomolecules GCATCTCCTCCACCAATCGCAACCCGCCCCCCGCTCCTATGGCTCCATATTTTGTCCGGC CTTATCCACATCCCGAAACCACCACCAGCTCCTCCGCTTCCATCGTTTTTCCTACCACCA ACGCCCGCTTCACCGGAACCGTACCCCCACCCAACCGCATGGCCGCGGCGCTCGCCTCCT CCCGCTGCTGCCGCCGCCCTTCGCTTCTTAACAGTGATCGTCGCCGCCGCTCCGTCGTCC GGTGCGCGCTCTCCGGCGAGAAAGGAAACTCATTTAGCTGGAACAAATGTGCAATTTCTA TAGCATTGTCTGTTGGATTAATTACTTGCCCACCAACATTTGGATGGTCGGCCCATGCTT TTCCGCTCGAACCTGTAATCCCTGATATTTCAGTTCTGATTTCTGGACCTCCCATTAAAG ATCCAGGGGCTTTGTTGAGATATGCTTTGCCCATAGATAATAAAGCTATTCGTGAAGTTC AAAAACCGTTAGAGGATATCACCGATAGTCTCAAAGTTTCTGGTGTTAGAGCGTTGGATT CAGTTGAAAGAAATGTGAGGCAGGCCTCGAGAGCGCTGACTAATGGGAGGAGTTTAATAC TTACTGGTCTTGCTGAATCAAAAAGAGAAAATGGAGAAAAAATATTGGATAAATTAGCTG TTGGACTTGAAGAGCTTCAACGAATTATTGAGGATAGAAACAGGAATGCGGTAGCTCCAA AGCAGAAAGAGCTTCTCAATTATGTTGGAATTGTAGAAGAAGATATGGTCGATGGCTTCC CCTATGAAGTACCAGAAGAATACAACAATATGCCTCTTCTTAAAGGAAGAGCTACTGTGG ATATGACGGTTAAGATTAAAGATAATCCAAACGTAGAAGATTGTGTATTTCGGATAGTTC TGGATGGATATAATGCTCCTGTGACTTCTGGGAACTTCGTAGATCTGGTGGAACGGAAGT TCTATGATGGCATGGAAATCCAAAGAGCTGATGGTTTTGTTGTTCAGACGGGAGATCCTG AGGGGCCAGCCGACGGTTTTATTGATCCCAGTACTGGCAAGAGTCGTACCATACCTCTTG AGATAATGGTTGATGGTGACAAGGCGCCTATATACGGTGAAACACTTGAGGAACTTGGCC TTTACAAGGCTCAAACAAAGCTTCCTTTTAATGCATTTGGAACGATGGCAATGGCTAGAG AATTGGATTATTCTTCGGAGGTTAAACTTCAATAACGGGTACGTCTTATGTTTCTGGATG GAGGGAGTAACATTTTCTTAGTTGTAACCAAGGTTTTTTCTTCTTCTTCTTCTCGATAAT GTTTTTTTGGATGGGCTAGAAGAATTGGTTTCTGGAAAATATCAGTATTCAGACAAAATT TGAAATCATTTTGTTTTGGGACAAAATATAAGTTTGCAGCCTGACCTACATATAGCTGCG CGTGTCCAAGCAGTTAGTACAATTCATCTGTTCCTCAAAGTAAGTCTCGGG
Protein Sequence (358 aa)
>BART1_0-u52907.019 358 150831_barley_pseudomolecules MAAALASSRCCRRPSLLNSDRRRRSVVRCALSGEKGNSFSWNKCAISIALSVGLITCPPT FGWSAHAFPLEPVIPDISVLISGPPIKDPGALLRYALPIDNKAIREVQKPLEDITDSLKV SGVRALDSVERNVRQASRALTNGRSLILTGLAESKRENGEKILDKLAVGLEELQRIIEDR NRNAVAPKQKELLNYVGIVEEDMVDGFPYEVPEEYNNMPLLKGRATVDMTVKIKDNPNVE DCVFRIVLDGYNAPVTSGNFVDLVERKFYDGMEIQRADGFVVQTGDPEGPADGFIDPSTG KSRTIPLEIMVDGDKAPIYGETLEELGLYKAQTKLPFNAFGTMAMARELDYSSEVKLQ
