- Transcript BART1_0-u52598.004
- Transcript IDBART1_0-u52598.004
- Gene IDBART1_0-u52598
Exon Structure of BART1_0-u52598.004
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr7H | 1 | 244648196 | 244647975 | + |
| chr7H | 2 | 244649011 | 244648303 | + |
| chr7H | 3 | 244649246 | 244649157 | + |
| chr7H | 4 | 244649454 | 244649354 | + |
| chr7H | 5 | 244649654 | 244649541 | + |
| chr7H | 6 | 244649826 | 244649745 | + |
| chr7H | 7 | 244649975 | 244649908 | + |
| chr7H | 8 | 244650141 | 244650076 | + |
| chr7H | 9 | 244650345 | 244650221 | + |
| chr7H | 10 | 244651074 | 244650741 | + |
| chr7H | 11 | 244651289 | 244651173 | + |
| chr7H | 12 | 244652881 | 244651667 | + |
| chr7H | 13 | 244653452 | 244653142 | + |
| chr7H | 14 | 244654255 | 244653589 | + |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials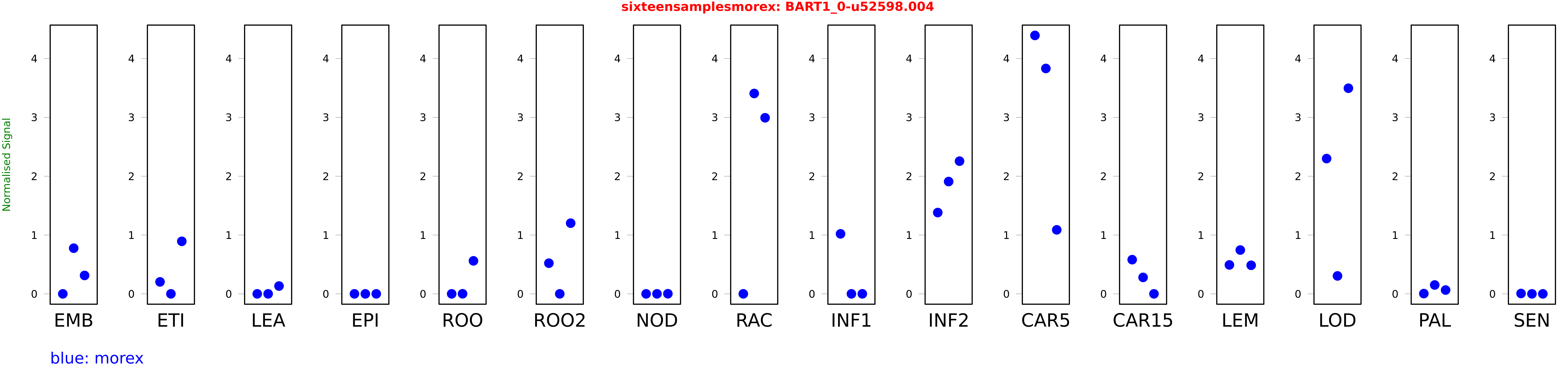
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0.0000251309 | 0.776292 | 0.31285 |
| morex | ETI | 0.203531 | 0.000176507 | 0.893079 |
| morex | LEA | 0.0000000124399 | 0 | 0.132781 |
| morex | EPI | 0 | 0.00000000580549 | 0.0000357086 |
| morex | ROO | 0.000013928 | 0.00130687 | 0.56111 |
| morex | ROO2 | 0.521382 | 0.000137113 | 1.20153 |
| morex | NOD | 0.000000000247394 | 0.0000664326 | 0.00198826 |
| morex | RAC | 0.000456449 | 3.40612 | 2.99284 |
| morex | INF1 | 1.02 | 0.000192176 | 0.000528483 |
| morex | INF2 | 1.38175 | 1.90897 | 2.25585 |
| morex | CAR5 | 4.39294 | 3.83139 | 1.08805 |
| morex | CAR15 | 0.581083 | 0.279571 | 0.0000307738 |
| morex | LEM | 0.491688 | 0.744817 | 0.485542 |
| morex | LOD | 2.29939 | 0.304863 | 3.49562 |
| morex | PAL | 0.0043883 | 0.150777 | 0.0647875 |
| morex | SEN | 0.00625592 | 0.0000000108893 | 0.00000000197304 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os08g43400.2 | +1 | 0.0 | 1494 | 749/1023 (73%) | protein|kinesin motor domain containing protein, expressed |
| TAIR PP10 | AT5G66310 @ ARAPORT AT5G66310.1 @ TAIR |
+1 | 0.0 | 717 | 458/1093 (42%) | Symbols: | ATP binding microtubule motor family protein | chr5:26485786-26490304 REVERSE LENGTH=1063 |
| BRACH PP3 | Bradi3g42190.1.p | +1 | 0.0 | 1578 | 792/1017 (78%) | pacid=32814334 transcript=Bradi3g42190.1 locus=Bradi3g42190 ID=Bradi3g42190.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (4221 bp)
>BART1_0-u52598.004 4221 150831_barley_pseudomolecules CCTTTACACCTTTTCCCTTCATTTTTTTTTCTACCAAAACCACTATAGTCCTCTCCCTTT CCCTTTCCCTTTCCCTTCCCCTCCTCCTCCTCCTATTCTTCTTCTCGGCACCAACAAGGA GGAGCAGAAGAGGAGAGGGGGCGACACAGTCTTGCCACTAGCTGCATTTCTCTCCTCTCG CATTGGTCGTCTTCTAGTTCCACCGTCCACCAGGGTGCGCAGAAAAACAAAGTCTTGATA CTCCACTGTCTCCCCCTCTATCCTCTCCTACCCCCCATGTGAGGTGCGTAAACAAAATTG AAGAGAGTGGCCGGGAGCGTGGCGGGGACCATGGTCCGTCCCTTCCTCTCGCGACAACAG CGCAAAACTCCAACCCAGCAAGCGGCGGCGGCCAAAGATAATCCGTAGCCGCGCAGGCAG CATTAGTACATGTGCGAGTAAGAGGGGGTGCGTCGGGCAGTGTCTAGTGGTACTAGTAGT TGCTGTTGTTACCTTGGGTCTGGGCAGGGTGGTCTCGCCGGATCGCCGTCCACGTCCGGA ACCCGGCGACAGACACGGCGCTCTCGGGCACAAGGACCTGCCACCAGCCGGCCGGCTTTG GATTCTTGCTTACAATGCTGCCTGTTCCCAGGGGAAGGGGGTGTTGGTCGGTGGTGACGG GTGCAGAGGAAGAGGGGCGATTTTGATACTGTTCTTGGGTGGAGGGGTGCAGGGCTGAGA AGGGCGATGGGGGCGGGGGAGGAGGATGCGGCGGCGGCATGGGAGAAGGTGGAGGCCAAG GAGGAGAGGATCATGGTGTCCGTGCGGGTGCGGCCCCTGAACGGCAGGGAGTCCGGCGAC AGCTCCGACTGGGAGTGCATTAGCCCCACCACCGTCATGTTCCGGAGTACCGTTCCTGAC CGCGCCATGTTCCCCACCGCCTACACCTATGACAGGGTGTTTGGCCCCAATTGCTCTACC AGACAGGTCTACGAGGAAGGAGCAAAAGAAGTTGCTCTATCAGTTGTCAGTGGAATCAAC TCAAGCATATTTGCGTATGGGCAAACAAGCAGCGGAAAGACTTACACAATGACTGGAATT ACTGAGTACAGTGTGATGGACATTTATGACTACATAGAGAAGCACCCTGAAAGAGAATTC ATGCTGAAATTCTCAGCAATAGAGATATATAATGAAGCTGTGAGGGATCTTCTGAGCCAT GATACGACACCTCTTAGACTGCTTGACGATCCTGAGAAAGGGACTACTGTTGAAAAGCTT ACCGAGGAAACATTGAGGGACAAGGACCATCTTAGGGATCTTCTTGGAATGTGTGAAGCT CAAAGAGAGATTGGAGAAACGGCTTTGAACGAAGCGAGTTCTAGGTCCCACCAAATACTC AGGCTGACAATCGAAAGTTCTGTTAAGCAGTATTTAGGAAGAAGCAAATCGAGCACCCTG GTGTCTTGTGTGAACTTTGTTGACCTAGCTGGAAGTGAGCGAGCATCTCAGACTGCTTCA GCTGGTATGAGGTTAAAAGAAGGTAGTCATATTAATCGAAGTCTGCTAACACTGGGAAAG GTTGTTCGCCAACTCAGCAAGGGAAGAAATGGTCATATCCCTTATAGAGATTCAAAGCTG ACACGCATATTACAGTCCTCTTTGGGAGGCAATGCAAGAACAGCCATTGTCTGCACAATG AGCCCTGCACACACTCATATTGAGCAATCCAGGAACACACTTCTGTTTGCGACTTGCGCA AAGGAGGTAGTTACAAATGCACAGGTCAATGTAGTGATGTCTGACAAAGCACTACTGAAG CATCTACAGAGAGAGCTCGCAAGATTAGAAAATGAACTGAAATTGCCGGAGTCAGCCTCC TGCACTAGTCATGCTGAAGCTTTAAGAGAGAAGGATGCACAGATTAAAAAGCTGGAAAAA CAGCTTAGGGAATTGATAGAAGAAAAGGACACCGTTCAGTCTCAACTTAACTGTTTACTG AGAAGTGATGGTGACGACCATGACAATGATCGTGCTGCAAAGCGATGGGATGTCCATAGT CGGTCCTCAGAGTCTCTTGCACGAAATGCATCGGAGGAAGCACTTTCGGTTTCAGACACT TATGGAGGTTCTTATCAAGATCAAGGTCACGCAGTGTTTGATGGATCATATGTCTTCAGT GCTGATCATGATGGCTTGTCATCCCCCAATAAACCAATGGATCTTCCTCAGCAAACAAGG GTTCGGCAGCCAGTTTCACCTTGGCACCCTACTAGCAACTATAGCTCTGATGGTGCGGAG TCATATAACATAAAGGAGGTAGCTTTCAGAACTGCATCTGAGGTATCCGAAGAACATTGT AGGGAAGTGCAATGTATAGAGATACATGAGCATAGAAGAAGCACAAGCCAGGAATTTAAC GTATTACTCCATGAGGGCACTAAGCTCCACATACCTGAAGTAGAGGACATCTCTAGAGAC GCAGTCCCGCAACCTGATGAAGTGCTAGAAGCAGGGAGCATAACAGAAAAAATGGAAGAT CATATGAAGATATACGCTAGCAAAGAGGAGCAGCAGGCCGAGATCATAGCAAATGCAGTA GAAGATCCTATTGAAGTGCATCAATGTGAATCTGATGACTTTGCACACAACTTTGTCAAA CTGTACCCATGTGATTCTAACATTTCTCTGGACGTTGGCAAACCTTACCCTCATGAATGT CTTACTGTAAAGAGGTGTATAACGAGCTCCAAGGATAGTGCATTGGCTAGAAGTCAAAGC TGCAGGGCTAGCTTCATGATCATTCCAAATAGTTGGTTTGATGATTCGGAAAACACCAGG CAGACACCACCTGATGAAATCTTCAGGTATCCTCCCAGAAGGCCTGATAAAGTCAGGAGA AGTCTGTACCAAGGAAATGATGATTGCCAAAACAATAACACTTCAGTAGACCTCTCTGCT GATTCTGGAGAAGTAGTTTGTGATGAAGTAGTTAAGGACACAGGTACCAGTGATGCAGTC AAGGACATGAGCACCAGTGCTGAAGTAGTCAAGGACATGAGTACCAATGATGAAGTAGTT AAGGCCATGAGCACCAGTGATGAAGTAGTCAAGGACATGAGCACCAGTGATGAAGTAGGC AAGGAGTCGAGTACCAGTGAAGCAGAACAAGAAGTTTGTATGGGTGACATCAGTTGTGTC ACAGAGCTGGAACAGAAGACAGCAAAACACCATGAGGACCAACCCGAGGAACATGAGGCA GAGCAGCAGACTGTTAGAGATGAGTGTACAGCAGTTAAAACTGTCAAAGATGTTGGCATA GACGCAGTCCCGAGTACCGCCGAGTCTCCTTCGTGCTGGCCTATTGATTTTGCAAATAGG CAGCGGGAGATCATCGAGTTGTGGCACGACTGCAACGTGTCCCTAGTACACAGGACCTAC TTCTTCCTCCTCTTCAAGGGAGATGCTGCAGATAGCGTCTACATGGAAGTGGAGCACAGG AGACTGTCCTTCATCCTCAGCTCGTTCAGCACCAACTCCGCAGGAGGCGAGCTTAACTCC GCAATCGCGTCCAGCTTGAAGAATCTGAAACGTGAGAGGGACATGTTCTACAAGCAGATG CTCAAGAAGCTCGCCAACGGGGACAAGGAGGGCATCTACACCAGATGGGGGATCGATCTG GGCTCCAAACAGCGGCGGCTTCAGCTATCTCGCCTCGTATGGACGCGGGCTGACATGGAG CATGTCCGGGAGAGCGCCTCGCTGGTGGCGAGGCTGATCGACCTGGTTGAGCCAGGGCAG GCTCTCAAGGAGATGTTCAGCCTCAACTTCACACTGGCTCCCAGAACCGAACGGCGGTCT TTCAGCCTGCTACTAGGCGCGTGATCTTTGACGTACGAAGGTAGAAGGGTTTGATTTACC TAGTAAGTACAAGGAGGTTGACTTAACAGGAGAAGAGGACTATGCTGCGTATCATCGATT GGTTTCACGCATGTGTCTCTGCCTTGTTTTTGCTCGTTGTGTGCGTTTGAAAGAATTGTG TGCGCGTGCGTGTGTGATATTTCTGTGTGCAGTCTCACAAAAGTGAATCAATCAGACAGT ACAGCTCTCCCTACCTTGCGTCGTATGTTCGTTTGTAAGAGTTTGTTCGTCAACCAAAAT TTTGGAAGCCATGCGTGGGAGCATCGCACCATCCGGAATTTCCTATTTGCCCCACTTTGC ATGGGCCGGCCTAGTTTTGCC
Protein Sequence (1045 aa)
>BART1_0-u52598.004 1045 150831_barley_pseudomolecules MGAGEEDAAAAWEKVEAKEERIMVSVRVRPLNGRESGDSSDWECISPTTVMFRSTVPDRA MFPTAYTYDRVFGPNCSTRQVYEEGAKEVALSVVSGINSSIFAYGQTSSGKTYTMTGITE YSVMDIYDYIEKHPEREFMLKFSAIEIYNEAVRDLLSHDTTPLRLLDDPEKGTTVEKLTE ETLRDKDHLRDLLGMCEAQREIGETALNEASSRSHQILRLTIESSVKQYLGRSKSSTLVS CVNFVDLAGSERASQTASAGMRLKEGSHINRSLLTLGKVVRQLSKGRNGHIPYRDSKLTR ILQSSLGGNARTAIVCTMSPAHTHIEQSRNTLLFATCAKEVVTNAQVNVVMSDKALLKHL QRELARLENELKLPESASCTSHAEALREKDAQIKKLEKQLRELIEEKDTVQSQLNCLLRS DGDDHDNDRAAKRWDVHSRSSESLARNASEEALSVSDTYGGSYQDQGHAVFDGSYVFSAD HDGLSSPNKPMDLPQQTRVRQPVSPWHPTSNYSSDGAESYNIKEVAFRTASEVSEEHCRE VQCIEIHEHRRSTSQEFNVLLHEGTKLHIPEVEDISRDAVPQPDEVLEAGSITEKMEDHM KIYASKEEQQAEIIANAVEDPIEVHQCESDDFAHNFVKLYPCDSNISLDVGKPYPHECLT VKRCITSSKDSALARSQSCRASFMIIPNSWFDDSENTRQTPPDEIFRYPPRRPDKVRRSL YQGNDDCQNNNTSVDLSADSGEVVCDEVVKDTGTSDAVKDMSTSAEVVKDMSTNDEVVKA MSTSDEVVKDMSTSDEVGKESSTSEAEQEVCMGDISCVTELEQKTAKHHEDQPEEHEAEQ QTVRDECTAVKTVKDVGIDAVPSTAESPSCWPIDFANRQREIIELWHDCNVSLVHRTYFF LLFKGDAADSVYMEVEHRRLSFILSSFSTNSAGGELNSAIASSLKNLKRERDMFYKQMLK KLANGDKEGIYTRWGIDLGSKQRRLQLSRLVWTRADMEHVRESASLVARLIDLVEPGQAL KEMFSLNFTLAPRTERRSFSLLLGA
