- Transcript BART1_0-u44239.022
- Transcript IDBART1_0-u44239.022
- Gene IDBART1_0-u44239
Exon Structure of BART1_0-u44239.022
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr6H | 1 | 180782519 | 180781937 | - |
| chr6H | 2 | 180782974 | 180782908 | - |
| chr6H | 3 | 180783205 | 180783085 | - |
| chr6H | 4 | 180783565 | 180783362 | - |
| chr6H | 5 | 180783770 | 180783666 | - |
| chr6H | 6 | 180786158 | 180785988 | - |
| chr6H | 7 | 180786706 | 180786233 | - |
| chr6H | 8 | 180787333 | 180786860 | - |
| chr6H | 9 | 180787544 | 180787451 | - |
| chr6H | 10 | 180787695 | 180787634 | - |
| chr6H | 11 | 180787865 | 180787778 | - |
| chr6H | 12 | 180788070 | 180788009 | - |
| chr6H | 13 | 180789168 | 180789050 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials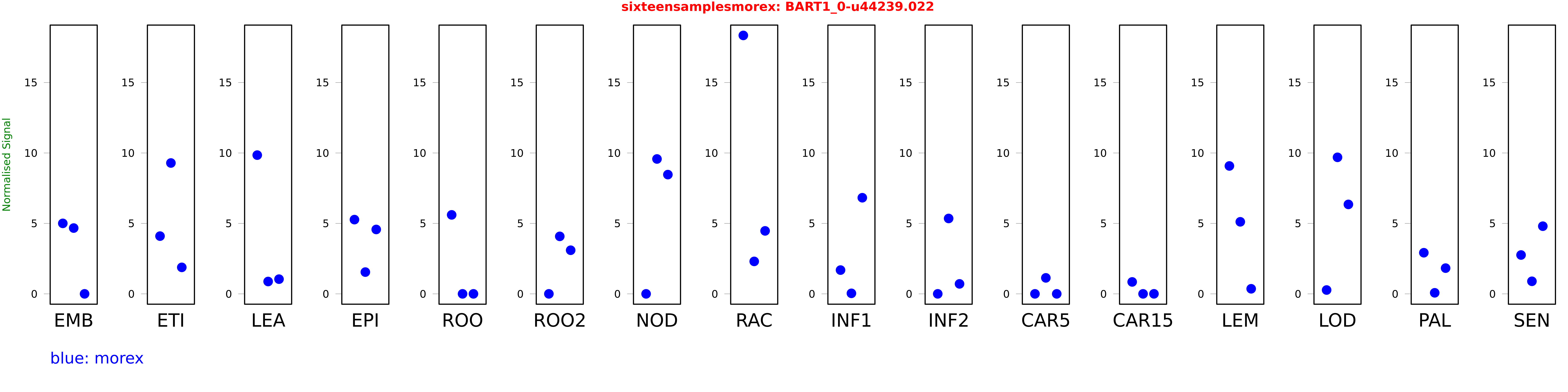
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 5.00295 | 4.67024 | 0.0000612926 |
| morex | ETI | 4.09833 | 9.28822 | 1.88033 |
| morex | LEA | 9.84713 | 0.876627 | 1.04357 |
| morex | EPI | 5.26946 | 1.54417 | 4.57117 |
| morex | ROO | 5.60541 | 0.0000463679 | 0 |
| morex | ROO2 | 0.00000980285 | 4.08059 | 3.09652 |
| morex | NOD | 0.00000702628 | 9.57362 | 8.46365 |
| morex | RAC | 18.3428 | 2.30376 | 4.46939 |
| morex | INF1 | 1.68733 | 0.0368829 | 6.82243 |
| morex | INF2 | 0.000335255 | 5.35123 | 0.707813 |
| morex | CAR5 | 0 | 1.13445 | 0 |
| morex | CAR15 | 0.847051 | 0.000000040255 | 0.00179219 |
| morex | LEM | 9.07801 | 5.11327 | 0.360164 |
| morex | LOD | 0.272417 | 9.6905 | 6.34914 |
| morex | PAL | 2.91854 | 0.0740118 | 1.82631 |
| morex | SEN | 2.75718 | 0.891679 | 4.80287 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os02g11820.2 | +3 | 0.0 | 762 | 507/749 (68%) | protein|GTPase-activating protein, putative, expressed |
| TAIR PP10 | AT4G13350 @ ARAPORT AT4G13350.2 @ TAIR |
+3 | 8e-90 | 297 | 219/553 (40%) | Symbols: NIG | NSP (nuclear shuttle protein)-interacting GTPase | chr4:7770170-7773321 REVERSE LENGTH=602 |
| BRACH PP3 | Bradi3g07910.1.p | +3 | 0.0 | 933 | 594/719 (83%) | pacid=32814567 transcript=Bradi3g07910.1 locus=Bradi3g07910 ID=Bradi3g07910.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (2624 bp)
>BART1_0-u44239.022 2624 150831_barley_pseudomolecules AGCAGGGTTTGCGGCGGGCGATGGCGAGCCGGGTGAAGGAGGACGAGAAGAACGAGCGGG TAATACGCGGGCTGCTCAAGCTGCCCGCAAACAAGAGGTGCATCAACTGCAACAATTTGG GACCACAATATGTATGCACAAATTTCTGGACCTTCGTTTGTACCAATTGCAGTGGAGCAC ACCGAGAGTTCACGCACCGGGTGAAATCTGTTTCCATGGCTAAATTTACTGCACAAGAAG TCACTGCTCTACAAGGAGGAGGAAATGAGCGTGCTAGAGAAATATTTTTCAAGGAATGGG ATTCTCAGCGCAATCCATACCCTGACAGCAGTAATACGGACAAATTGAGGAATTTCATCA AATATGTTTACGTGGAACGGCGATATACAGGAGAAAGAAGCTCTGATAGACCACCGAGGG GAAAGGACGATAAGGATGAACATAGCGAAAATAGAAGATCTGATGGAAATCGGGGTGGTT CAAGAAGTCCACCATATAATGAAAGTTACTCTGACCGTCGGAGTTATAGTGGACGGAGTG ATGACAGAAATTCCAGACATTCCTATGGAGAACGTAGTCCTGGCTATGACCAGAATGATT ATAAGAAAAGCCCCCGCCACTTTGAAGTTGTCGACGATAGATCTGGGAAAACAACTCCAG TTCAAAGGTTCGAGGACCGCAGGTTTTCTGAACCACGAAAGCCAGAGACTGGGTCTCCAA ATTACCAGAGAGAAGCCAATGGATCTAGTCCTCCTGTTGTACGGCCAGTGAGGGAGATAT TGGGTGATGATGCTCCTCAACTTCGGATTGGTGAACCTCCAAAGCCGAATGTAGCCAGAC CGATTGACCCTCCAAAACCAAATGGAGCCAGGACCATTGAACCTCCACCTCAGGCACAGA GGACAGCTACTGCCAGCAGCATTGGTTCTTCCGAGGGGATTTCGGAACAGACGAAGGCAG TAACTCCAGTCAGTTTAATTGATTTTGGTGCAGATCCCGAACCTACTGCATCTGCCCTGC CATCACAGACAGCCCCTACACCCCAGCAGCAGTCAGTTAATGCACAGCCTATTAACACAA CAGCGCCACAACATGTTCTTGAACAAGGCAGAAGTGCTCCATCAGTGAGCGGCGGTGATT GGGCATCTTTTGATGCTTTTGGCCAGCAACAACAAACACCACAATCAGGCAGCACTGTAA ATCCGCTTGAGTCTGTGCTTGCACAACTATCATTCTCTGAAACACCCTCTGCCTCCAGTA CATCAGCCTTTTCAACTTCTGTTGATCAGCAAGCAAATGGAGGACAGTCATCCATGATAG ATCCATCACATTCTTCATTATTTGGTCCACCCGTTGGAATCTCAGGCAATCAGGCCTCAA CAGGAATGCCCGTCCAAGGATCTGCAGTTCATCAATCTGCTGTGGCTGCCCCTATGGGTG GTCTTCCATCTCTATTGCCTTCAAATTCTCAAGGAACAAGTGGTATTCAAGAAGCAACGT CTTCTCATGACAGCAAATCCAGTGGGCGGACAGCACTTCCAGTGGATTTCTTTACTTCAC TTTATCCATCAGCAACTCCAGCAATGCCTGGCTGGCAGAGAGCTCCCCAGTCTGGAATGG GATTTGGCATGCAGTATCCTGCCGGAATGGGAATGCAGTCATATCCTCAGGCAGCATTTT CTCAACCTACATATCAACAGCCTACATATCAGCAACCTATGTATCAGCAACCTACATACC AACAACCTACGTATCAACAGCCTGTATATCAGCAAAATGCATATCCCCAACCTGCGAAAG CTTCAAACCCTTTTGACCTCGGAAATGAGGCAGCTCCAATTCAAGCTCACATGTCCCTAT CTGGACCACTGGGAGCATCAGCAGGCGCTGCTAACCCAACATTAGTTGGTAACTCTAGTT TTGGAGTTCCACCTCAGCAGCCTCAGCAGATGTATCAGCCACCTGTACATCAAAACCATT ACATGATGCAGCATGTTTCAAACAACATGCCTGAGCAGCTACCTAATGGCATGCTTCCAA GGCAACAAGGAGGGGCTGGTTCCCTCGGCGTCGGATATGATCAACAAGCTGCCCCTAGGT ACTCCCAGCCAAGCACCCCACCTTCATATGGTGCTGTCGGTGGAAATCCTTTTGGGTAAA AAAGGCTTATCAAATGTGGACGTACCATGTTACCTGTTTATGGTATCTTATGTATCTTCA GACCACTGGTGTGTAGTGCCCACTGCCCAGTGAGACCAGAATGCACAAAGCCCTGGAGAA AAATTTGTTTCATATTTACTGGTCATGTGTTCTGTCACTGTAGCTCCGTGTAACTTGGTC TGGGAGGTGGAAGCGACGTGGTGCATAAGCTTCAGGAGCAGTAGTTCGGTGCTTCTAGCC CCTATTTTCTTGTATATTTGTGTTTGGTGTTCTGTATGCGTTGCCTCCGATGTATATTCA CAGAGACCCTATGCCTTGTATGGTTTTTGATTGGGATCTTTCGAGCGGCCCTATCTGATA GTGTACTTAGCTCAGAATGTTACTTTTAGTCATGCTGGGTATTCAATTTTAATGTTTATT GCAATGTCGGACTGAAGGTTGTATAACTGTGGTTAATCAGTTGT
Protein Sequence (718 aa)
>BART1_0-u44239.022 718 150831_barley_pseudomolecules QGLRRAMASRVKEDEKNERVIRGLLKLPANKRCINCNNLGPQYVCTNFWTFVCTNCSGAH REFTHRVKSVSMAKFTAQEVTALQGGGNERAREIFFKEWDSQRNPYPDSSNTDKLRNFIK YVYVERRYTGERSSDRPPRGKDDKDEHSENRRSDGNRGGSRSPPYNESYSDRRSYSGRSD DRNSRHSYGERSPGYDQNDYKKSPRHFEVVDDRSGKTTPVQRFEDRRFSEPRKPETGSPN YQREANGSSPPVVRPVREILGDDAPQLRIGEPPKPNVARPIDPPKPNGARTIEPPPQAQR TATASSIGSSEGISEQTKAVTPVSLIDFGADPEPTASALPSQTAPTPQQQSVNAQPINTT APQHVLEQGRSAPSVSGGDWASFDAFGQQQQTPQSGSTVNPLESVLAQLSFSETPSASST SAFSTSVDQQANGGQSSMIDPSHSSLFGPPVGISGNQASTGMPVQGSAVHQSAVAAPMGG LPSLLPSNSQGTSGIQEATSSHDSKSSGRTALPVDFFTSLYPSATPAMPGWQRAPQSGMG FGMQYPAGMGMQSYPQAAFSQPTYQQPTYQQPMYQQPTYQQPTYQQPVYQQNAYPQPAKA SNPFDLGNEAAPIQAHMSLSGPLGASAGAANPTLVGNSSFGVPPQQPQQMYQPPVHQNHY MMQHVSNNMPEQLPNGMLPRQQGGAGSLGVGYDQQAAPRYSQPSTPPSYGAVGGNPFG
