- Transcript BART1_0-u39181.005
- Transcript IDBART1_0-u39181.005
- Gene IDBART1_0-u39181
Exon Structure of BART1_0-u39181.005
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr5H | 1 | 602222746 | 602220219 | - |
| chr5H | 2 | 602224004 | 602223248 | - |
| chr5H | 3 | 602224386 | 602224150 | - |
| chr5H | 4 | 602225112 | 602224892 | - |
| chr5H | 5 | 602228014 | 602226500 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials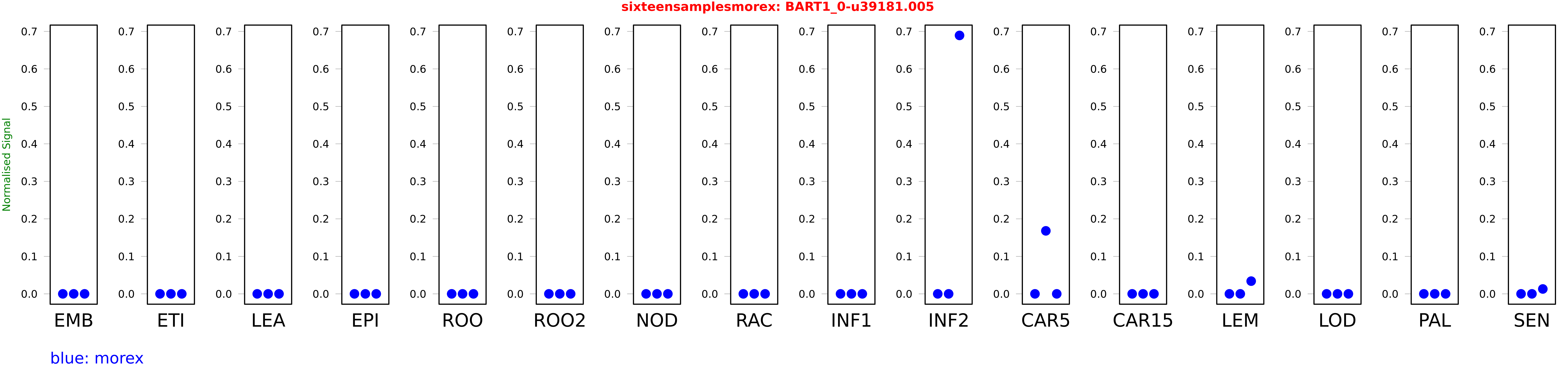
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0 | 0 | 0 |
| morex | ETI | 0 | 0 | 0 |
| morex | LEA | 0 | 0 | 0 |
| morex | EPI | 0 | 0 | 0 |
| morex | ROO | 0 | 0 | 0 |
| morex | ROO2 | 0 | 0 | 0 |
| morex | NOD | 0 | 0 | 0 |
| morex | RAC | 0 | 0 | 0 |
| morex | INF1 | 0 | 0 | 0 |
| morex | INF2 | 0 | 0 | 0.689221 |
| morex | CAR5 | 0 | 0.168029 | 0 |
| morex | CAR15 | 0 | 0 | 0 |
| morex | LEM | 0 | 0.00000343053 | 0.0339357 |
| morex | LOD | 0.00000250729 | 0 | 0.0000000445778 |
| morex | PAL | 0 | 0 | 0 |
| morex | SEN | 0 | 0 | 0.0130237 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os03g55130.2 | +3 | 0.0 | 566 | 384/669 (57%) | protein|expressed protein |
| TAIR PP10 | None | - | - | - | - | - |
| BRACH PP3 | Bradi1g08000.3.p | +3 | 0.0 | 1424 | 784/1010 (78%) | pacid=32799386 transcript=Bradi1g08000.3 locus=Bradi1g08000 ID=Bradi1g08000.3.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (5258 bp)
>BART1_0-u39181.005 5258 150831_barley_pseudomolecules CGCACGGAAGTAAAACTAGTAAAACAAGGACGTGACACAACAATAATGTAAACAACCACT TCCTCTGCCTAGGTTATAAGTCCCTATTGATTTTAGAGTCAAACTTTGACCATAGATTTA GCTAGCAAAATATAAGTTTGACGCCGTATAATTCATAGCATTGGATTCGTATTTGAAAGA AGCTTCAAATGACATTATTTTCAATACATACATTTATACAATCCAAGTTTGCCATAAAAT ACGAGAGGGGCTAATAAACCCGTACGGACGTAGCAGTAACTCATTGACATACACTATTCC ACAATCCAAAGCCACAAGTGCACACGAAATCCAATAAGTAATAGTAGTACTAGTAAAATA TACATCAACACATCAAACATGCACTATTCCAAAGCCATAAGTGCATGGAAAATACATTCA ACATGTCCAACCACCATTACCAGTACCAGTACATGCCGCTATAGTGCTCCTTGCAGTAGT AGTACTTGCTGAAAGGAGTCCAAGACAAGCGCGTTGATGCATTTGGTTTTTGTTGCCATG TAAAGGTCCGGACGGCGTGTACGTAGCAGCGCAGATGCAAAGCCCAGGAGGGAGATAAAA AAAGGCATGCATCACGAGGGGAGGGATAAAAAGATGGCCGGTTTTGCAAGGTTTGGAAAG CAGGTTTGGTGGTACTGTCCTCGAGTATTTTAACAGTTGCCGTTTCTCCCTCTCCCCCCC CACCGCACCAAGGGTGGACAGATTCTCTCCCATGTCCCAATCCCTAAACTCCCCCATTAT ATTACTCTAGAAACACACCAAACCGAGGAAAGAAATCAATCATCCTCGTGATAACTGTCT CGTAGGAAATCCGATCCTCCAGAATAATTGTATCATATCAGCACACTTGGGATCGGAAAG GAAGGAAAATGGAATGAACTAGGAGGACTGGTTCCACACCGGAATTGTGAGAGGGACAGC AGACGGTGGCGATGTGGTGGCAGGGCAGGGCAGGGCAGGGGAGGGGAGATAATAGTATTA TAATATTGGGAAATGTCTGAGTTGAGTTGGTGATTGATGAGAGAGAAAGACGGAGGGAGA GGGACAAGGGGTGGGGTGATGGGGAGGATCAGCATCAGGGGGGCGGAGAGAGAGAGAGAG AATATCAGATGTCAACAAATCTGCTGTCCTTGCGGCTCGTTTAGAAAGAGAGTAGAAGGT GGTTCCATTCCATCCCCTCTTCCCTTCCACCAGAGGAGAGAGAAAGAAAGAGAGAGAGAG GGAGGCCAGGCCAAACAACGCCAATCCAAAACCCCCAAAAGGAGCAAGGAAAGAGAAGAG AGAGTGGGGAGTGTGTGTGTGAGAGAGAGAACAAACCGAGATACTCCCACATCCGAGAGC AGACAGTGGTTCCTTCCTTCGTTCGTTCGTTCGTTCCGGCAGGAGCAAGAACCCACCGCT TTCTCCCCCTCTTCTTCCTGGCCTGGCCCCAGCAAATCCGATTCCAACCTCCGCCCGGAT ACATCGGAGCTCCGGGTTATTTCTCTTGAAGTGCCATTCAGATTGGGAGGTAGTATTGAT TCAGCTGAGCAAGTCGAGCTGTCTAGTTAGCTCAAAGCATTGAAGATTGGAATTTTACAC GTGCATTGCATTAACACTGTGATTTCCCTTTGGTGCCCTGTGATGTGATGATCAGTTCTA GTTGCCGTCCGGGTCAAGGCGAATTTTAGCATGATTGCTACATGCCCATTGTTTTGACTG TAACACTAATGGGAGCTAAAGTGGAAGCTGAGAGTTACAATACTGGATATTATGCCACAG GGGATCTGCACATGGACTCAAATGGTTGGCGCTCTCCATACTATGAGGATAAGCCATCAA ATGGGCAGTTGTACAATGGCTTCATGACAAAGCAGGCAGATGGTTATTCAGAGTACGACA ACGAAACGCTGAAGCGCACAATGCTTTCACATGAAGCTATGTTTAGACAGCAGGTTTATG AACTGCACCGTGTTTACAAAATACAGAGAGACCTGATGAAGCATTATGAAAGCAAAGACA CATATGCATACCCAGTACTGGCAGATGCATCACAGACAAATTCATCATCACAAGTTCCAC CAAATGGCTCAAAGATGATATGGCAGATGCCTCTAGGGGCCAAAACATACGGAGAAGCTA CTGTTGCAGTGCACACTGATACAAACCACTCCTTGAAGTTCCTTAGTGATGGCAGTGTTC TGCCCTCTCCAGATGGATTCCCCTCAAATGGTGTTGCTTTAAATACCAAGCAAGGCGTTA TTGACCTTGAGCTTCCAGCTGATCATTATATAGATGATGACAACACATCAGATGATAAGC CTATAGATTTCCTTGGAGTAGATTCGGGTACAAAACTGCCGAATGATGCCGGGGTCACAT TCAGTGGCGCAGAAGGCTTGGGGAGGTTCAGTGATAATAGCTCAACCTCTGGTTTGCTGG CTACTAATAAACCGGGAGGCTGCCATGTCGCTGACCTGAATGAGCCAATTACTGGGATGC ATATGGGTGGAACAAACGGGTCAGTACCCAGAGGTCTTCCGTATACCTTGGAGAACTCTT GGCGCCAATCTGCAGTGAGATCAGGCACAGCTAACTTTGGTTTCAATAAAGAATACTCCA AAGAGAAGCATACCGATGAAGGAACGAGCTCAAATTTCCTCGATGCAAGTGCCAAACTAA GACAGGAGGAGAAACCATTGATTGATAAAGGAAAACAAGTCAGCAGTCAATCTTTTTTTA CACCCAGATACAGCAATACTGATCTGCAAAAATCCTTCAAGGTTGCTGATGGGAGATCTG CAACCAATGAGTTTATTTACCATGGTCAAAATAGTTCAGTTGGATGGTTTTCAAAAGGTC CTATGGGAGCATCTGCCGTCAATAATTTTGCTAGACTTGACCATCCACATCATTCAAGTA TTGGTACACTTGTTGCTCCAATCTCCATTCCTCACATAGACCATCCTTCCGTCGCCTCTC CTATAGGTTCTTGCACAGTGGACCCAAGGAGCAGTGTTATCAACAATGCTTTCCAGCGAG TTCCAAGTTTCAATGGATCCTCAACTGTTAATTCATACAAGAGTCCTGGTGCTGTGACCC AAAGCATTGGACCTTCGAGTCACAGCATTGGACCTTCGATTCATAAGCTTAAAAGATTCG ATAATTTGGATGGTAGTTACTTTGGCTTTCCGCTTGATCCATTTTCCGCATCACGATCCA GACAACAGGTTGCAATGTCAAGTGAGTTGGAGCAAAGGAACTGCCTGATGTTTGAACATT CAGCTGCACGGCAGTCTCTTGGGTATACTCAGCCCACAAGCACCAAGGGCACAAAGAACT TTAATCTGAATGAAGCACTGTCAGATGGTCCAGAAGAGCTTGTAGAACAGGATGGAAGAT CTGTTGGCAGTTTGCAGCATAGTAAAGGTGAGGGATCAGTATTTGGGATCTCTTGGCTAA AAAATAGAGCTACTTGTGCTGATCCAACAGGTTTGGAAAAGCAAAGGAAGGTGTTTGGCC ATTCCAATGGGACTGCAACAGACACGAAAAACAACAACGACTTGACGGGAACAACACCGA TTGTTTGCAATTTGTCAGATTCCGCCTCGACGTCCCTAGGTTGCAGAATTAAGATAGATG ATGCTTCTGATGTCATCACTGATAGAACTTGGCTGGTATGTAATAAGACTGAGGAGAGTA CCACGAGACATTTACCACTTTCATGCCAGAAACCCTTTGACAGAGATGAACAAACTGCTG AAGGTGTTGTCAAGAAAAGTGGCGCCGTCATCAGAAATTTATTTGATCTGAATGATGACA TACCACATGAAGATAACTCAGAGTCATCAGTAGTGTCACATGAATGCGACGTGGCGTCGT TACAGAACAACCATGCTAAGCGCACATTTTTGATTGACTTGGAAGAAATGCCAGCTTGTG AAGATGGTGCTGCCTGGACTTCTCAACAGGAATGCACACCTTCTGGTACGCTTGATGCAT GCAAGGAAGCTGATATGTGTTCTCCATCCGCTATGGATGCAGCGCAGATCATTCTTGCGT TGTCAACGGATGTGCTGACCACCACAGGCACACCCGATGATATGCTGCAGTGGTTTGCTG AGCTCGCGATATCAAGCACGGATGCCCATGCCGATGAGCAAGCAGAAGCCCAGGGTTGCA TCAATAATTTCAGTGATGATGGTTTGGATTCATTTGAAGCCTTGACACTGAAGCTGGAAG AGTGCAATACTGATGAGCGTTGGACACCGGAACCACCACTGACAGACGATGAACAAGCTG TTTCAGCGGTGAACCTGCTGTCCAAGCCAAAAAGAGGACCGCAAAGGAAGCGACGGCAGA AGCGAGACTTCCAGAAGGACATCTTACCTGGTCTCTCGTCACTTTCCCGTCCTAAAATCA TAGAGGACATTCAGCTAATTGAGGGGTTAGTACAGGCATCTGGTGGATCATGGGAGTCGA GTTTCACTAGGCGAGCGCGCTGTGGGCGAACAAGAGGTAGAAAGCCCCGCAAAAACCAGC CAGCTACCGTAGAGATAGAAGCTGAGGCCGAGGCCGAGGTGAGCCCGCCCTTGAAACCCG ACTCCATTGGCTTAGAAGCTGATGAAAGGGGCATGGTTAGCTGGGGACGAACAACAAGAC GGTGTCGCAAGCCGAGATGCCCCTCGGGCAGTAACATCGCGGCTTCGTCTTGAGCGACTG ATCTTGGTCAGAGAGAACGTCGGTCACGATCATGTACTTGGGTTCGGGTTCCAGATTTGC CATCGAGATTGCGGTGGACTGGTATAAAGGTAAAAAATCACAGCCATGAGTGTGCATACT CTGCTGGCCTGTTTGGCCGGCCTCCCCCCTGCAGAGATTTCATGCCTTTTCGGGTCGAAA CATGGTGTACATTTTTTTCTATAATAAATTCTTAAACTAGTCTATATGTGACAATTTCTG TAACAGCGTTGTATTAAGCTGGGCTCCAGGGCCTGAGGGGCCTGCTGTAATTCTTGAGAT AGTGGCGAGGTACCTTGACTGTCCGGTATTCAGAAATCAAAAAGGCTAATCGGTGTAGTT CTATATAGTTCTTTTAAATCTTGTTTTGTTTTGTGTTGTGCAAGTTGCTGCTTCATTTGG TTGCAATCTTTGCTTCGCAGCCTGTACAAGTTGGAACGCCGATTCGTGTGGTATACCACC GTTGAGGACTTGATCCAGACAGAACTGGTAGTTCATGG
Protein Sequence (1003 aa)
>BART1_0-u39181.005 1003 150831_barley_pseudomolecules MPIVLTVTLMGAKVEAESYNTGYYATGDLHMDSNGWRSPYYEDKPSNGQLYNGFMTKQAD GYSEYDNETLKRTMLSHEAMFRQQVYELHRVYKIQRDLMKHYESKDTYAYPVLADASQTN SSSQVPPNGSKMIWQMPLGAKTYGEATVAVHTDTNHSLKFLSDGSVLPSPDGFPSNGVAL NTKQGVIDLELPADHYIDDDNTSDDKPIDFLGVDSGTKLPNDAGVTFSGAEGLGRFSDNS STSGLLATNKPGGCHVADLNEPITGMHMGGTNGSVPRGLPYTLENSWRQSAVRSGTANFG FNKEYSKEKHTDEGTSSNFLDASAKLRQEEKPLIDKGKQVSSQSFFTPRYSNTDLQKSFK VADGRSATNEFIYHGQNSSVGWFSKGPMGASAVNNFARLDHPHHSSIGTLVAPISIPHID HPSVASPIGSCTVDPRSSVINNAFQRVPSFNGSSTVNSYKSPGAVTQSIGPSSHSIGPSI HKLKRFDNLDGSYFGFPLDPFSASRSRQQVAMSSELEQRNCLMFEHSAARQSLGYTQPTS TKGTKNFNLNEALSDGPEELVEQDGRSVGSLQHSKGEGSVFGISWLKNRATCADPTGLEK QRKVFGHSNGTATDTKNNNDLTGTTPIVCNLSDSASTSLGCRIKIDDASDVITDRTWLVC NKTEESTTRHLPLSCQKPFDRDEQTAEGVVKKSGAVIRNLFDLNDDIPHEDNSESSVVSH ECDVASLQNNHAKRTFLIDLEEMPACEDGAAWTSQQECTPSGTLDACKEADMCSPSAMDA AQIILALSTDVLTTTGTPDDMLQWFAELAISSTDAHADEQAEAQGCINNFSDDGLDSFEA LTLKLEECNTDERWTPEPPLTDDEQAVSAVNLLSKPKRGPQRKRRQKRDFQKDILPGLSS LSRPKIIEDIQLIEGLVQASGGSWESSFTRRARCGRTRGRKPRKNQPATVEIEAEAEAEV SPPLKPDSIGLEADERGMVSWGRTTRRCRKPRCPSGSNIAASS
