- Transcript BART1_0-u37955.005
- Transcript IDBART1_0-u37955.005
- Gene IDBART1_0-u37955
Exon Structure of BART1_0-u37955.005
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr5H | 1 | 564069900 | 564069445 | - |
| chr5H | 2 | 564070540 | 564070357 | - |
| chr5H | 3 | 564070767 | 564070626 | - |
| chr5H | 4 | 564071554 | 564071423 | - |
| chr5H | 5 | 564072263 | 564071649 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials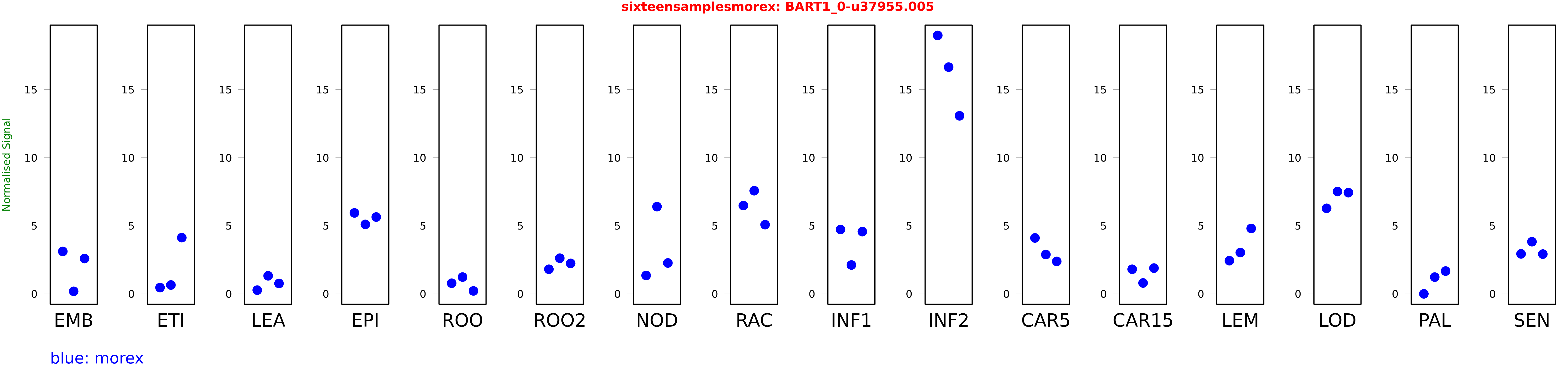
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 3.11458 | 0.187703 | 2.59101 |
| morex | ETI | 0.462015 | 0.650032 | 4.12638 |
| morex | LEA | 0.272444 | 1.32506 | 0.761701 |
| morex | EPI | 5.94381 | 5.10263 | 5.64149 |
| morex | ROO | 0.78074 | 1.2321 | 0.215642 |
| morex | ROO2 | 1.80664 | 2.61804 | 2.23939 |
| morex | NOD | 1.35032 | 6.40479 | 2.26784 |
| morex | RAC | 6.48225 | 7.57437 | 5.08283 |
| morex | INF1 | 4.72529 | 2.11885 | 4.56932 |
| morex | INF2 | 18.9788 | 16.6506 | 13.074 |
| morex | CAR5 | 4.10657 | 2.88761 | 2.38279 |
| morex | CAR15 | 1.80661 | 0.799048 | 1.89155 |
| morex | LEM | 2.43503 | 3.03359 | 4.80368 |
| morex | LOD | 6.28384 | 7.51503 | 7.43055 |
| morex | PAL | 0 | 1.23011 | 1.67392 |
| morex | SEN | 2.93561 | 3.83077 | 2.91858 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os09g36140.1 | +2 | 1e-47 | 168 | 88/138 (64%) | protein|RNA recognition motif 2 domain containing protein, expressed |
| TAIR PP10 | None | - | - | - | - | - |
| BRACH PP3 | Bradi4g36020.1.p | +2 | 5e-68 | 221 | 108/141 (77%) | pacid=32789987 transcript=Bradi4g36020.1 locus=Bradi4g36020 ID=Bradi4g36020.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1529 bp)
>BART1_0-u37955.005 1529 150831_barley_pseudomolecules CTGCGGTGGTGCCACCTGGTGCGTGATACAGGGGGCAGGTGGTGCTCCTCCAAGTCCTCC TCGGGTCGGTCTTGCCCGGGTCCGGTCCTCCTTCCTCCTACTTTTCCCCAGCTCAAAAGA AAAATCAAGTTAGCCATGGCCGCCGCCTACACCTCGCGCCTGGGCCTGCCCCCCGCCGCG CACCCGTACGACGACGAATCCGTGGCCTGCGCCCTAGGGCATCTCTACCCCACCAAGGAT CCGTTCGCCCTGCCGGCCAGCTGCCTCCTCCCGCCGCCCGAGCTGCCGCTGGCCTTCGGG TTGCCCGCCGGCGGTCTCGCCCCCGTCTGCTGCTGCGCCGACGCCGCGGGGGCGCCCGCG TTCCCGCCCTTCCCATGGGCGAACGTGCCCTCGCCGCCAGTGCCGCCGCGTTGCGGCATC ACGGAGATCGACGAGTTCCGGGAGGTGGAGTCGGAGGATAACCTCTCCCCTTGTTCCGTC CTCACGCCCTGGAGGAGGCCGACGCCCGCCTCGGCGCTGCCTTCGCCTCCGCCCCTCGTG GTCGGGGGGAAACGGGCCTTCGACCCCACCAGCGAGAAGACTTCTCTGATGATCTGCAAT ATCCCCAACGGATTCGAAGCGGAGGTTCATGGCGATTCTGGACCAGCATTGCGTCCATGA GAACGACAATCCTGAATGGCGTGTCGTTGGGGGTGGCAAGTTCGTGAGATCGGAGTATGA CTTCCTCTACATCCCAATAGACTTCAGGACGAAGTACAACAAGGGCTATGCGTTCGTCAA CATGACGACGGCGACTGCAGCGCGGCGGCTCCATGCTTTTCTCCACGGGCACCGCTGGGC GCTGGCGGGCTCTCGGAAGGTATGCGAGGTTGTGCACGCGGACATCCAGGGGGTGGATGC TTTGTCAGCGCACTTCTCCAGCTCCAAGTTCCCATGCGGTAACAAGGACTTCCTGCCAGT GCGCTTCGGCCCGCCGCGGGACGGCCTTCGGCCGACGGTGGAGCGTGTAATTGGCCGCAC GGTGGTCCACCGTCCCGCTGATCAATCTGCTCACCCCACGCCGCGTGCTGCACAAAGAGG AGGCAAATGGAAACCTGGTGCAGTGGGGAACAAAACTGTTTGAGCTGGTGGCTACTGGCT AGTGATGTCTTTGCTTCGTCGCGAGGATGGAGGCCTACTCACCTTAGGGTAGGTGCTTAG TGTGCTTGGTTACCCAAGATGATCAAGTTAGATGGTTCTTTTGGGGCTGCAGGAGTATCT TCTAGTGTCTGTATGACTGTATCCTACCTTGGAGATGTTGGCTTCATTTGTCTTCCCTAG GTAGCAGCAATGCTTGGTGGTTCTGCCATCCGATTTCCTTTGAGTTGTCACTAACTGTAC TCATCTGCCCTGAACTTTCTTAGATTGGTTTGTGCATCTGTTCGCATGTTAACCGTATGT TATCCGCCCGTGTATGTTTTTGAAGATTTGGCATCCAGTTGCGATGTGTGATGTTGATGG CGCTGAAAGAAAAGGTTAATGGTGGGGAG
Protein Sequence (219 aa)
>BART1_0-u37955.005 219 150831_barley_pseudomolecules LRWCHLVRDTGGRWCSSKSSSGRSCPGPVLLPPTFPQLKRKIKLAMAAAYTSRLGLPPAA HPYDDESVACALGHLYPTKDPFALPASCLLPPPELPLAFGLPAGGLAPVCCCADAAGAPA FPPFPWANVPSPPVPPRCGITEIDEFREVESEDNLSPCSVLTPWRRPTPASALPSPPPLV VGGKRAFDPTSEKTSLMICNIPNGFEAEVHGDSGPALRP
