- Transcript BART1_0-u35721.006
- Transcript IDBART1_0-u35721.006
- Gene IDBART1_0-u35721
Exon Structure of BART1_0-u35721.006
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr5H | 1 | 405760204 | 405760074 | + |
| chr5H | 2 | 405760512 | 405760450 | + |
| chr5H | 3 | 405761523 | 405761364 | + |
| chr5H | 4 | 405762633 | 405762486 | + |
| chr5H | 5 | 405762807 | 405762730 | + |
| chr5H | 6 | 405763442 | 405763358 | + |
| chr5H | 7 | 405763590 | 405763535 | + |
| chr5H | 8 | 405766075 | 405765455 | + |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials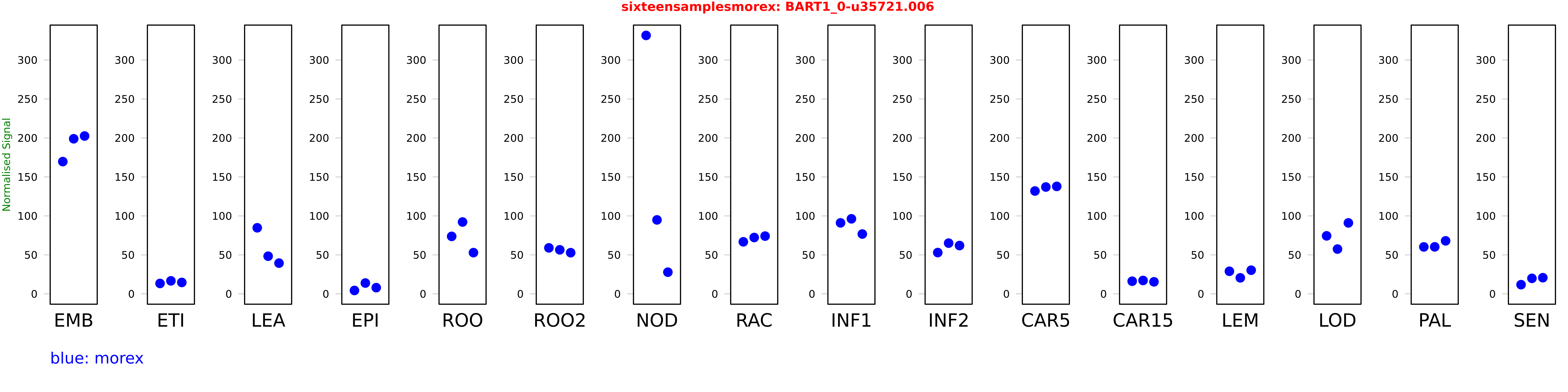
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 169.677 | 198.973 | 202.508 |
| morex | ETI | 13.3608 | 16.6811 | 14.637 |
| morex | LEA | 84.7651 | 48.3351 | 39.4258 |
| morex | EPI | 4.33869 | 13.8804 | 7.94598 |
| morex | ROO | 73.7533 | 92.1893 | 52.8816 |
| morex | ROO2 | 59.004 | 56.4931 | 52.813 |
| morex | NOD | 331.553 | 94.8816 | 27.8066 |
| morex | RAC | 66.7671 | 72.2905 | 74.1004 |
| morex | INF1 | 91.0835 | 96.2645 | 76.755 |
| morex | INF2 | 52.9479 | 65.0785 | 61.9602 |
| morex | CAR5 | 131.952 | 137.076 | 137.872 |
| morex | CAR15 | 16.2236 | 17.1713 | 15.4344 |
| morex | LEM | 28.9719 | 20.5523 | 30.3528 |
| morex | LOD | 74.4151 | 57.4808 | 91.0247 |
| morex | PAL | 60.2203 | 60.3004 | 68.0103 |
| morex | SEN | 11.7802 | 19.8913 | 20.7291 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os02g21250.2 | +3 | 2e-99 | 295 | 170/182 (93%) | protein|clathrin adaptor complex small chain domain containing protein, expressed |
| TAIR PP10 | AT3G09800 @ ARAPORT AT3G09800.1 @ TAIR |
+3 | 2e-72 | 226 | 127/178 (71%) | Symbols: | SNARE-like superfamily protein | chr3:3006731-3007916 REVERSE LENGTH=179 |
| BRACH PP3 | Bradi4g08770.4.p | +3 | 6e-104 | 307 | 177/182 (97%) | pacid=32783612 transcript=Bradi4g08770.4 locus=Bradi4g08770 ID=Bradi4g08770.4.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1342 bp)
>BART1_0-u35721.006 1342 150831_barley_pseudomolecules CCAGCTCGTCCATTTCCGCTCGCCTAATCCCTCCCGTCTTTCTCTACACTCGTCTGCTGC ATCTCAGATCTCGCGGGCACCAACAAACCAAAGCAAAGCTCTCCCGCTGTCCCGATCACT CTCCCTTCCAGGAAGCTTGGACCGGAGGAAGGTGCTGAGAGGGCGGAAAGGAGGCCATGG GGGAATTCTCCAAGGAATCTTGCCCGTCTGTGAAGAACATTTTGCTACTGGATTCTGAGG GAAAGCGCGTTGCTGTGAAGTACTTCTCGGATGATTGGCCGAACAACGCATCTAGGTTGA CCTTTGAAAAGTCCATTTTTACTAAAACTCTGAAGACAAATGCACGCTCAGAAGCTGAGA TAACATTGCTTGATGGTAATATTGTTGTTTACAAGTTTGTACAAGACCTTCACTTCTTCG TCACTGCCGGAGATGATGAGAATGAGCTAATCATAGCAAATGTGCTACAGGGTTTTGCTG ATTCGGTTGGTCTTCTACTCAGGGGTGACGTTGAGAAGAGGACTGCACTTGAGAACTTGG ACTTGATACTTCTCTGCATTGATGAGATTATTGATGGCGGCATAATTCTTGAAACAGATG CGAATACCATTGCGGGGAAGGTTGCAACCAATGCTGCTGACGGTTCTGTTCCCTTCTCTG AGCAGACAATATCCCAAGCTCTGGCCACAGCCAGGGAACACTTTGCAAGATCTCTTCTGA AGTTTACCCTACCAAACGAATTTCAAATGCCACAAAACTTTAATATGATTTTTTCTGGAC TATATGAAGACCTGCAAGCATCAGAAGTGGACGAGAGGCCGAACGAGGTAGGCACAAGCC AACAGGGCGCGGCCGCGCCCTAATGACTTGTGCCCCCCTAGATGCCTCCCGCCGTCCCTC TCTGACCTATAAATTCACAAATATTCCGATGCACTCCCGAAACACTTTTTCCGCCGCCGC AAGTCTCTGTTCTGACGGAAGGCCATCTAAACCTCCATTCCGGCGATCTGTCGGAGGGGA GATCAGTCACGGAGGGCCTCTACATCAACCTTGTTGCCCTCACCGTGATGTGTGAGTAGT TTACCACATACCTACGGGTCCGTAGGCAGTACCTAGATGACAATCTCTCTCTCTTATGAT CTTCAATACCATGTTCATCGCATCTTCGCTTTGATCTATCCGATGTAATCACTATGTATG GTGTGTTTGTTGGGGTCCGATGAATTGTGGGTTTATGTTCAGATTATTCATGAAAAGTAT TTGAGTCATCTCTGATTATTATGATTCTTAATTGCATGATCATCATAGCTTCATAATGCT CTTCGATCTATTGAGATAGTTT
Protein Sequence (228 aa)
>BART1_0-u35721.006 228 150831_barley_pseudomolecules MGEFSKESCPSVKNILLLDSEGKRVAVKYFSDDWPNNASRLTFEKSIFTKTLKTNARSEA EITLLDGNIVVYKFVQDLHFFVTAGDDENELIIANVLQGFADSVGLLLRGDVEKRTALEN LDLILLCIDEIIDGGIILETDANTIAGKVATNAADGSVPFSEQTISQALATAREHFARSL LKFTLPNEFQMPQNFNMIFSGLYEDLQASEVDERPNEVGTSQQGAAAP
