- Transcript BART1_0-u33637.016
- Transcript IDBART1_0-u33637.016
- Gene IDBART1_0-u33637
Exon Structure of BART1_0-u33637.016
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr5H | 1 | 84473806 | 84473606 | + |
| chr5H | 2 | 84474019 | 84473884 | + |
| chr5H | 3 | 84476028 | 84475935 | + |
| chr5H | 4 | 84477467 | 84477275 | + |
| chr5H | 5 | 84478810 | 84477550 | + |
| chr5H | 6 | 84479142 | 84478916 | + |
| chr5H | 7 | 84480117 | 84479843 | + |
| chr5H | 8 | 84480655 | 84480237 | + |
| chr5H | 9 | 84480831 | 84480728 | + |
| chr5H | 10 | 84483365 | 84483169 | + |
| chr5H | 11 | 84483994 | 84483785 | + |
| chr5H | 12 | 84490180 | 84489767 | + |
| chr5H | 13 | 84491799 | 84491518 | + |
| chr5H | 14 | 84493204 | 84492482 | + |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials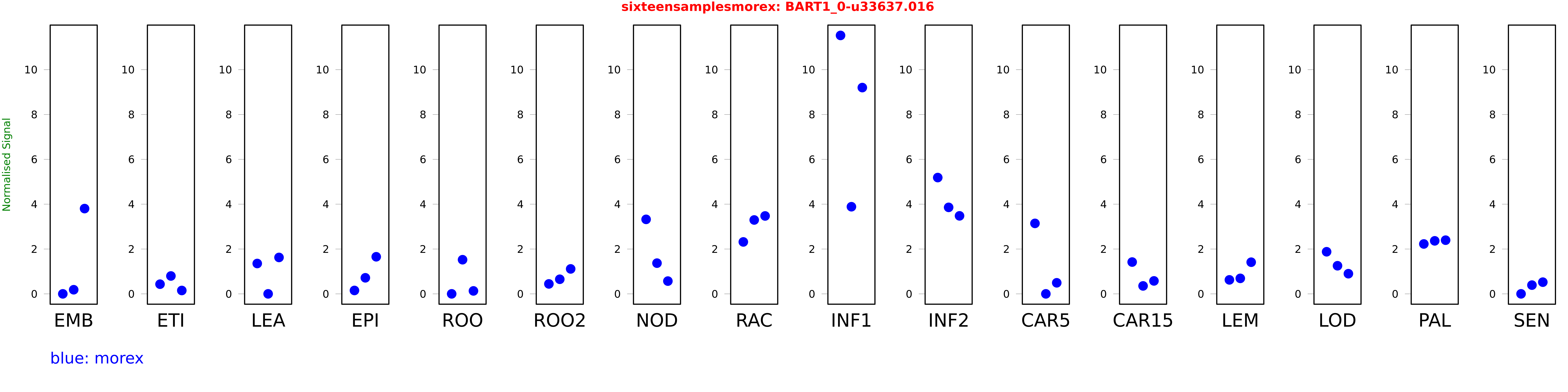
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0 | 0.183859 | 3.80353 |
| morex | ETI | 0.431624 | 0.797358 | 0.150465 |
| morex | LEA | 1.35401 | 0 | 1.62424 |
| morex | EPI | 0.151884 | 0.716128 | 1.65427 |
| morex | ROO | 0 | 1.52284 | 0.13166 |
| morex | ROO2 | 0.443124 | 0.651278 | 1.1127 |
| morex | NOD | 3.32323 | 1.37006 | 0.569267 |
| morex | RAC | 2.31653 | 3.29576 | 3.47506 |
| morex | INF1 | 11.5278 | 3.88661 | 9.20112 |
| morex | INF2 | 5.18728 | 3.85814 | 3.47936 |
| morex | CAR5 | 3.1435 | 0 | 0.492999 |
| morex | CAR15 | 1.41886 | 0.354395 | 0.577763 |
| morex | LEM | 0.624096 | 0.691084 | 1.41178 |
| morex | LOD | 1.87787 | 1.25369 | 0.898646 |
| morex | PAL | 2.22579 | 2.36452 | 2.39427 |
| morex | SEN | 0 | 0.38988 | 0.521737 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os03g38740.1 | +3 | 0.0 | 1268 | 626/874 (72%) | protein|Dicer, putative, expressed |
| TAIR PP10 | AT3G03300 @ ARAPORT AT3G03300.3 @ TAIR |
+3 | 0.0 | 629 | 361/856 (42%) | Symbols: DCL2 | dicer-like 2 | chr3:768020-774833 REVERSE LENGTH=1388 |
| BRACH PP3 | Bradi1g15440.2.p | +3 | 0.0 | 1355 | 683/873 (78%) | pacid=32792174 transcript=Bradi1g15440.2 locus=Bradi1g15440 ID=Bradi1g15440.2.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (4736 bp)
>BART1_0-u33637.016 4736 150831_barley_pseudomolecules GTTTTTTGGATATGCATGGCCTTGTCTTCTAAGTTTTAACACCTTCAATAACCTAACATA TTTACATTCTCTTAGCATTTAAAAGTCCTCAAGGCAAATCTACATGGATCGTCTCGGAAA AATGCGAAGAAAGGGATATCAAAATTACACAAGGCATTCTTATATTGCACGGCTAATCTT GGAGTATGGCTAGCGGCAAAGGCTGCTGAGATACACTCAACAACTAACGAACAATTCTTA TCTTTCTGGGGTGAAGAACTGGATAAGAATGTTGAAGGCTTTGTTAGAAGCTATAGTGAA GAAGTATACAGGGAACTCTCATGTTTTTCAGAGAGAGGTCATATTGGTAAAGATTTTGAA GCTGATTTGCAAGATGGGTTATTGACACCTAAAGTTCATTGTCTTGTTCAGTTTCTTCTG GAATACAGGTGGCACATGCAAGACATAAGATGCATTGTATTTGTCGAGAGAGTTGTTACA AGCATTGTGTTGGAGTCTCTCCTGTCCATTATCAATCAAATGCCTGGGTGGATTATTAAA CACATTGCAGGAAACAGACCTATGTTGCACAACCAGAGTAGAAATAAACAAACGGAAATT GTTGACGCCTTTAAAGGAGGAAAGGTACATATAATTGTGACCACACAAGTATTGGAGGAG GGGCTGAATGTGCCAGGTTGTCACTTAGTAATTCGCTTTGATCCACCAACGACTGGGCGC AGCTTTATACAGTCACGTGGTCGAGCTCGGATGCCAAACTCTGAATATGTTTTACTTGTT AGAAGGCACGTCCTTCGATTTTTGTTGCTATTTTTTTGTTTCCAGCGCAAATTTTATTTA ATTGTTCATCTTCATGCAAATTAAAATCTTTGTCTTCTGTATTGTGGACATTATCCACCT CTTTTCCCACTTCATCAGTTGAATATGATGTGCTCAGTATATAATCCTTGGTCACTAGTA CCAGTAAATTGTTGTTTACCTATATTACTCAGTGTGTTCCGTTTATCAGTTGAATTGTTT ATATTGGATTAAGCATACTGGGCTGGGTGTTAAATGCTAGTGTTCACTTACTATTGTGAG ATGACTTGATGTGACTGCCGGTATGACTCCAAATCCTGATATGGTCAAACCTGAAACAAT TGATAAATGCCAAGTTCAAAAGCCACTTGTTTGCTTTTTCATTAAGAATCTAAAATTGAC TAGTGTTTCCTAGTGTTCACTTAATATTGCAAGTTGGCGTGATGTGAGTGCCGTTATGAT ACCAAATCCTGATATGTTTTCCTGTCAGATCTGTATGGAAACCGCAGGTAACCATAACAT GATAAGGATCTGTTTTCAATCGAGACAACTGTTCATATTTGAGTAGAAAACCGTCCGACT TTTTGCTCGGTTATTGGTGGAAACTAGTCAGCACTTATGATTATTGTCAGTGAGCTGTGA ATTCTATTCCTACTGATTACATTTCTCTATTTAGCTATACAAAAAATGTGGACTCGTGCT ACAGTGGAAAAATCGCTATACTCAAGTTGTCTTAAGTTGATTTATTAGTACAGTTTCCTC TTGTATTGCAAAAAAACCCTACTTTACCTTTGTTCTCATCATCACTTCTAGCTCTTTTTG GCATGCTACGTAACATGATGATGCAATGCTTGTGTTTCTGTATCAGGGGGGATGCAGAGG CCCTCTCTAAGACAGTGAAGTTCCTTGCAAGCGGGCAAATGATGCGGGAAGCATCTCTGA AGCTTGCATCTACTATGTGTCAACCACTAGAAAACACCTTGTTGCAGGGAGAATATTACC GTGTTGAATCTACTGGAGCAACCGTGACTATGAACTCAAGTGTCCAGTTGATTTATTTTT TCTGCTCGAAGCTCCCATCTGATGAATACTTTACACCTCTTCCGAGATTCATTATTGACA AGGAACTACGTACTTGCACCTTGTACCTTCCCAATAGCAGCCCTGTGCAGACTGTTAATG CAGAAGGAGAAGTTTCTGCCCTCAAAAAGGCAGTTTGTCTTAAGGCCTGCCGAGAATTAC ACGCTGTCGGCGCCCTAACAGATTATCTGTTACCAGAATTTGGTTTTCCATGTGAAGAAG AACCCGACATTGTTGTTGAGAAATACCAACACGAGCAGCCTGTGTACTTCCCAGAGGAAT TTGTGTACAATTGGTCATCCTTTTCTCGTCTTGGAATATATTATTGCTATAAGATTTCAG TAGAAGGGTGCTTGAAAACAACTTATTGTCCAAATGACATACTTCTAGCTGTGAAATGTG ATTTGGGACCAGAGTTTGTCTCTACCTCACTTCAATTGTTTGGTGCGCAAGATAATGCTA GTCTTGCCATGAAATATGTGGGAATCATTCATCTGAATCAAGAACAGGTAGTTATGGCCA GAAGGTTTCAGACTACTATTCTTTCTTTGCTTATTAATAAAGACCATTCTGAGGTCAGTA ATGCTGTTAAATACTCCCATGAGATGCAAGTGTCCATTGGAATTGTCTACCTGCTGCTAC CCTTAGTTTCTGGCAAAGTTGATTGGTGCAGTATCAAGTTCTCGACCTCGCAAGTGTATG TTGGCAGTAACAAGGATATCAGACATTGCCATTCTTGTAAACAAGTTGATTTACTGCAAA CAAAAGATGGTCCTCTGTGCAGATGCATGCTTGAAAGTTCCATTGTTTGCACCCCTCATA ATAGCAAATTCTATGCTGTTAATGGGTTCCTTGACTTAAATTCCAAATCTCTTCTGCACC TTCGTGATGGCAGTGCCCTTACCTACATAAACTATTTCAAGACTAGGCATGGACTCAGTC TAACACATGAAAATCAACCCCTGTTGGCAGCCAGGAGTCCTGTTGAAGTGCGGAATTTCC TCCACAAGCGCCATTATAAAAATAAAAAAGAATCGTGCACTTCACATGGTGTTGAGTTGC CACCTGAGCTCTGCAGACTAGTCATGTCACCAGTCTCTACTAATACTTTGTACAGCTTCT CGATTATTCCTTCTGTAATGTACCGCATCCAGTGTCTGCTTCTTTCTGCAAAGTTGAAAG ATCAACTGGGCCCAATAATGCAGCAGTTTGCCATACCAGCTCTGAAGATTCTAGAAGCAG TAACAACCAAGGAATGCCAAGAGGAATTTTCACAGGAATCATTAGAAACCCTTGGGGATT CTTTCCTCAAGTACATCGCCACGCAACATCTGTATGTCAAACACAAGCTTCAGCATGAGG GGACATTAACCAAGATGAAGAAAAACTTAATCTCGAATGCAGCTCTATGCCAGCTGGCCT GCAACAACAATCTTGTGGGTTACATTCAAGGGGAAGAGTTTAACCCAAAAGGTTGGATCA TTCCAGGGCTTGGTTATGACATGTGTGGTAACAGCAAATTTTCCTTTTTAAGCTCCAATG ATATGCACACTTTGAGGAAAATGTCTGTTAGAAGTAAGAGAATAGCAGATACAGTTGAGG CATTGATTGGAGCCTACCTTGGTGCTGCGGGAGAACAGGCTGCATTTGTTTTCCTAAAAT CATTAGGATTGGATATAGGATTTCACAGCACTATTCCACTTGAAAGGAAGATTGTATTAA AATCTGAAAAGTTCATCAACGTGAGAAGTTTGGAAATGATTCTTGGTTATGAGTTCAAGG ATACTTCATTGTTGGTGGAAGCATTGACACATGGGTCATACCAGACTGCTGGCACTACTG CATGCTATCAGAGGCTTGAGTTTCTCGGAGACGCTGTGTTGGACCATCTCTTCACTATAT ATTTCTACAATAAGTACCCTGAATGTACTCCAGAATTGTTGACAGATCTTAGATCTGCTT CCGTTAACAACAATTGCTATGCGCATGCAGCTGCTAAAGCAGGACTAAACAAGCACATTC TTCATTCGTCATTGCCATTGCACGGAAGGATGGCTTACTACTTTGAGAATTTTACTAAGC ACCGATTTACAGGTCCATCACATGGTTGGGAAGCTGGCATTGGTCTACCCAAGGTTCTTG GGGATGTTATTGAATCATTAGCCGGTGCCATTTATCTTGACAGCAAGTATGACAAAGAGG TGGTTTGGAGAAGCATGAAAAAGCTATTGGAGCCTCTTGCGACACCGGAGACGGTTGAGC GTGACCCAGTGAAACTGCTGCAGGAATTTTGTGATCGCAGGTCTTATAGCAGGTCTTACA CAAAGGTCCAGAAAGATGGAGTGTCTTCGGTGGTTGGGGAGGTACAGGTAGAAGGGACTA CCTACTCAGCAACTGGAACTGGCCCTGATAAGGTTGTCGCAAAAAAGCTTGCAGCGAAGT TGCTGCTAGAAGATCTGAAGGCTATAGTACCATGACACGCTTGCATTTGCAGATGATGCA ATACTGTATAGCTCCTTCCGACTTCACTTGTATTGTGACTAGCCTGTAGGTAGCCTTTTG AAGTTTTTGTGGGAGTATGCGTGTACTTGACAAATGTGCAGCTCAATCATTGCTTGTGAA TATTCAGACTATTGAACTGCACATCTGTGAAGTACGGGCATCGTCTCTCAGAACCATGGT ACAGACGATAATGCTACAGTATCCTTAGACTTCTACTTGTACAGTTGTAAGCACAATCTT GAGTGGAAAGCTGGAATATCTTTTTCTCTCCATCGGGCAGTGCAGTGCATGTATCTTTTA TTTTTTGAGCGGGGGTACCTTTTGTAGGGCTGGTAATTGTTTTTTTTAAATGGCAG
Protein Sequence (910 aa)
>BART1_0-u33637.016 910 150831_barley_pseudomolecules MLRNMMMQCLCFCIRGDAEALSKTVKFLASGQMMREASLKLASTMCQPLENTLLQGEYYR VESTGATVTMNSSVQLIYFFCSKLPSDEYFTPLPRFIIDKELRTCTLYLPNSSPVQTVNA EGEVSALKKAVCLKACRELHAVGALTDYLLPEFGFPCEEEPDIVVEKYQHEQPVYFPEEF VYNWSSFSRLGIYYCYKISVEGCLKTTYCPNDILLAVKCDLGPEFVSTSLQLFGAQDNAS LAMKYVGIIHLNQEQVVMARRFQTTILSLLINKDHSEVSNAVKYSHEMQVSIGIVYLLLP LVSGKVDWCSIKFSTSQVYVGSNKDIRHCHSCKQVDLLQTKDGPLCRCMLESSIVCTPHN SKFYAVNGFLDLNSKSLLHLRDGSALTYINYFKTRHGLSLTHENQPLLAARSPVEVRNFL HKRHYKNKKESCTSHGVELPPELCRLVMSPVSTNTLYSFSIIPSVMYRIQCLLLSAKLKD QLGPIMQQFAIPALKILEAVTTKECQEEFSQESLETLGDSFLKYIATQHLYVKHKLQHEG TLTKMKKNLISNAALCQLACNNNLVGYIQGEEFNPKGWIIPGLGYDMCGNSKFSFLSSND MHTLRKMSVRSKRIADTVEALIGAYLGAAGEQAAFVFLKSLGLDIGFHSTIPLERKIVLK SEKFINVRSLEMILGYEFKDTSLLVEALTHGSYQTAGTTACYQRLEFLGDAVLDHLFTIY FYNKYPECTPELLTDLRSASVNNNCYAHAAAKAGLNKHILHSSLPLHGRMAYYFENFTKH RFTGPSHGWEAGIGLPKVLGDVIESLAGAIYLDSKYDKEVVWRSMKKLLEPLATPETVER DPVKLLQEFCDRRSYSRSYTKVQKDGVSSVVGEVQVEGTTYSATGTGPDKVVAKKLAAKL LLEDLKAIVP
