- Transcript BART1_0-u29974.009
- Transcript IDBART1_0-u29974.009
- Gene IDBART1_0-u29974
Exon Structure of BART1_0-u29974.009
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr4H | 1 | 511456072 | 511455912 | + |
| chr4H | 2 | 511457260 | 511457229 | + |
| chr4H | 3 | 511457495 | 511457384 | + |
| chr4H | 4 | 511457741 | 511457592 | + |
| chr4H | 5 | 511458151 | 511458084 | + |
| chr4H | 6 | 511459233 | 511458273 | + |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials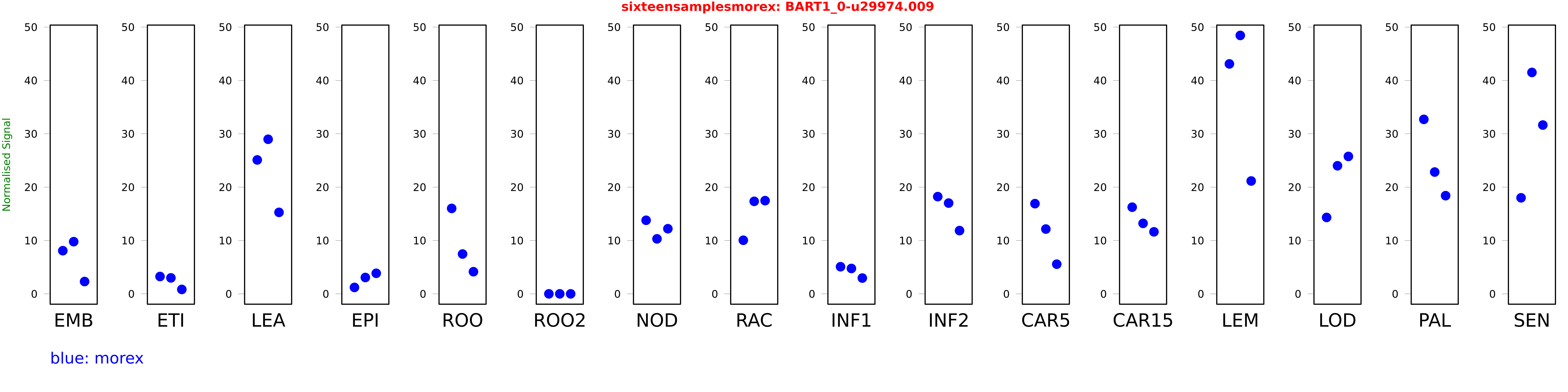
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 8.08984 | 9.7795 | 2.31041 |
| morex | ETI | 3.25214 | 2.9906 | 0.830997 |
| morex | LEA | 25.0981 | 28.9726 | 15.2728 |
| morex | EPI | 1.20583 | 3.06657 | 3.8544 |
| morex | ROO | 16.0132 | 7.46737 | 4.15168 |
| morex | ROO2 | 0 | 0 | 0 |
| morex | NOD | 13.7906 | 10.3094 | 12.2069 |
| morex | RAC | 10.0464 | 17.3369 | 17.4677 |
| morex | INF1 | 5.08391 | 4.7582 | 2.95996 |
| morex | INF2 | 18.2229 | 17.012 | 11.8542 |
| morex | CAR5 | 16.9049 | 12.1454 | 5.56207 |
| morex | CAR15 | 16.2411 | 13.213 | 11.6249 |
| morex | LEM | 43.096 | 48.4333 | 21.153 |
| morex | LOD | 14.3216 | 24.007 | 25.7564 |
| morex | PAL | 32.7024 | 22.8295 | 18.3998 |
| morex | SEN | 18.0012 | 41.4972 | 31.6386 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os03g15890.2 | +3 | 3e-47 | 167 | 90/100 (90%) | protein|RNA recognition motif containing protein, expressed |
| TAIR PP10 | AT1G07350 @ ARAPORT AT1G07350.1 @ TAIR |
+3 | 1e-34 | 134 | 70/101 (69%) | Symbols: | RNA-binding (RRM/RBD/RNP motifs) family protein | chr1:2257732-2260101 REVERSE LENGTH=382 |
| BRACH PP3 | Bradi1g67145.1.p | +3 | 1e-50 | 176 | 95/101 (94%) | pacid=32805382 transcript=Bradi1g67145.1 locus=Bradi1g67145 ID=Bradi1g67145.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1484 bp)
>BART1_0-u29974.009 1484 150831_barley_pseudomolecules CTCCAGAACCTTCCGGCAGGTCCTCTCTATTTATTCACACGAGCGCGCGTGCACCATCCA ACGGGAAAAGAGAGGCGGTGGTCGCAGGTCGGAACTGACGGAGGCAGAGGCGAAGACGAA GACAGAGACTCGAGAGGCCTCGCAGCGGCGACGAATCAAAGATGTCGTATTCGAGGTACA GGAGTCGTTCAAGGAGTGTGGACTCGAGTGATGTTGAGAACCCTGGGAACAATCTCTTTG TGACTGGTTTATCATCTCGTCTAACTGATCGAGATCTGGAGAAGCATTTCTCTACAGAGG GAGAGGTGATTGATGCAAGTATTGTACTTGATCCATGGACAAGGGAATCACGGGGATTTG GATTTGTTACCATGGCCAATCTTAAGGAGGCAGAACGCTGCATCAAATATCTCGACAGTT CAGTGCTGGAAGGTCGTGTCATTACTGTTGAGAAGGCAAAGAGAAGACGAGGTCGAACCC CAACACCGGGAAGGTACCTTGGCAGCAAATCATCGCGTCGTAAGAAGGAGGTATTCCCGA AGCAGGTCACCTGTTAGGAGAGACCGTTACGGCTCACGCTACCCGCCCGAGCTAGAACGT TCTTATTCTCCTTATCGCAGAGGACGATCATACTCTCGTCATGACAGGCATAGATCATAC TCCCCCTATGGAAGGAGGCCATCCTACTCCCCCTATGAAAGGAGGCCATCCTACTCTCCC TATGGCAGAAGGAGATCATACTCTCCTTCCGACAGGTCGGAGTCTCCCTACCACAGGCGT AGATCCTACTCACCCTACGACAGGCGTCACTCACCCCGCTATGGTCATCGCCACCGTTCA AGATCCCCATACAGTTACAGAAGGCGGAGGTCATGCTCCCATGACCGTTATGTTTCAGCA TACTACAGCAGGCGCTATTCTCCAAGGAACAGAAGACGGAGTTACTCTCGCAGCGTATCA CCACGCAGGAGCTACTCCCGTAGCTGCTCCCTGGCGTCAGAAGAATCAAGGAGCTGTTCT CCGCAGAAAGGATATGCTGAAAGCAAGGCATCGCGCAGCAGGTCACCTGCGATTGCGAAG AGGCGTTCTAGAGAAAGCTATGCGCACAGCCGCAGTTCATGCTCGAGGTCCGTCTCCAGG GAGCGCTCAACCTCATCAAGCACCTGATGGTTTACAAAGTGCCTTGATGACTTTGGATGT TTGTAAACAATGTGGATGGTATGATCGTGTACCTAGTGAAGCAGCTGTATTTCGTCATCG TTAACATCTTAAATTGCAGTCTGCTTCCATACTATAGGAGAAAAATGCAGAAAATTTCCT ACAGATTGCCTATTCCACAAGGATTTTATAGCATACTTCTACAACGGAAAAAACGCAGAA ATTTTCATAAATATTGGCGAGGTCATTTCTTTGTTTCAGATGACCCAGTGCGGATTTTTC TATAGGAATTCGATACTGCAATTTTCCTTTGTTTATAAGATCTC
Protein Sequence (131 aa)
>BART1_0-u29974.009 131 150831_barley_pseudomolecules MSYSRYRSRSRSVDSSDVENPGNNLFVTGLSSRLTDRDLEKHFSTEGEVIDASIVLDPWT RESRGFGFVTMANLKEAERCIKYLDSSVLEGRVITVEKAKRRRGRTPTPGRYLGSKSSRR KKEVFPKQVTC
