- Transcript BART1_0-u29736.006
- Transcript IDBART1_0-u29736.006
- Gene IDBART1_0-u29736
Exon Structure of BART1_0-u29736.006
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr4H | 1 | 483706263 | 483705921 | + |
| chr4H | 2 | 483706465 | 483706392 | + |
| chr4H | 3 | 483706600 | 483706543 | + |
| chr4H | 4 | 483706780 | 483706679 | + |
| chr4H | 5 | 483707346 | 483707226 | + |
| chr4H | 6 | 483712155 | 483711561 | + |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials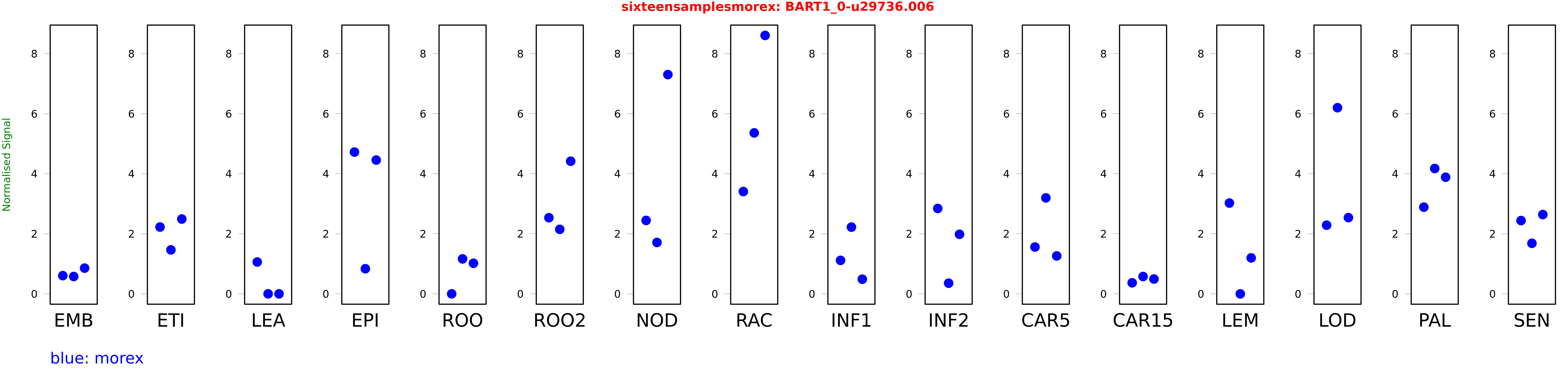
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0.606522 | 0.577947 | 0.856577 |
| morex | ETI | 2.22585 | 1.46425 | 2.49237 |
| morex | LEA | 1.06154 | 0 | 0 |
| morex | EPI | 4.72151 | 0.836647 | 4.45567 |
| morex | ROO | 0 | 1.16396 | 1.01864 |
| morex | ROO2 | 2.53505 | 2.14859 | 4.41772 |
| morex | NOD | 2.44565 | 1.71215 | 7.30134 |
| morex | RAC | 3.40741 | 5.36002 | 8.60611 |
| morex | INF1 | 1.11551 | 2.22201 | 0.487663 |
| morex | INF2 | 2.84347 | 0.354667 | 1.98325 |
| morex | CAR5 | 1.55991 | 3.19631 | 1.26266 |
| morex | CAR15 | 0.367741 | 0.577679 | 0.4941 |
| morex | LEM | 3.02432 | 0 | 1.19778 |
| morex | LOD | 2.28672 | 6.199 | 2.53807 |
| morex | PAL | 2.88719 | 4.17416 | 3.88411 |
| morex | SEN | 2.44031 | 1.68342 | 2.64117 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os03g27320.3 | +2 | 9e-144 | 413 | 258/310 (83%) | protein|steroid binding protein, putative, expressed |
| TAIR PP10 | AT5G58220 @ ARAPORT AT5G58220.3 @ TAIR |
+2 | 6e-86 | 265 | 157/299 (53%) | Symbols: TTL | transthyretin-like protein | chr5:23554546-23555861 REVERSE LENGTH=311 |
| BRACH PP3 | Bradi1g60700.3.p | +2 | 2e-154 | 439 | 263/308 (85%) | pacid=32804009 transcript=Bradi1g60700.3 locus=Bradi1g60700 ID=Bradi1g60700.3.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1293 bp)
>BART1_0-u29736.006 1293 150831_barley_pseudomolecules CTTCCGGCAAAAACAGGTGTTGCTCTCGTAGCGCTCGCTCTCACCCACCCTTCCCCAACT CGCATCCAGCGCCGCAACCCCTCGTCTCTCCCCCCCTTCCGATCTGAGGCGTCGTTGGAG ATCGGCGGCAACCGAAGAGGCGTCCGACGACGGCCGTCGCGGGTGGCACCAGTGCAGTGG AGCAAATCAACCAGTTGAGATGGCGACATTGAGGCCGCTGAGCGTGGAGGAGGTGCTGAG GGTGAACGGCAGCCACCGCTTTGCCGCCGCGCTGGCTGCCGCCTCCCCCTTTGATTCTCT TGCCAACGCGCTCCTTGCCGCCCGCCGCATCTGGCTGAACGAGGTGGACGTGACTGGATG GCTGGAGGCCTTCGCTGCCCACCCACCCATCGGAAGCACCTCCTCCTCCGTCTCCAAGTG GAGCAAGGAAGAGCAATCAGCTGCTCTTTCCACAGCCACTGATTCGACTGCGCAGGAGTT GTCCGAGTGGAATGCCAAATACAGGGAGAAGTTTGGGTTCGTGTTCATGATATGCGCATC TGGGAGGACTGCCCCTGAGGTCTTAGCTGAGCTTAAGAGGCGTTATGCAAATAGACCAGT TGTTGAGCTTGAGGCTGCGGCAGAGGAAGAACTGAAGATAACTGAGCTGCGTCTTGCAAA GCTTTTCGCATCAGAAACTGCTGCTCCCCCTACTTCAGGCAACTCCAACCGGACACGCCC TCCTATCACAACCCATGTGCTGGACACGGCTCTTGGATCGCCAGCATCTGGAATTGAAGT TCACCTGGAGATGTGGAAGGACGCGTCAAGTACCCCGTCATTTGACAACAAAGATTTCAA GGGATGGACAACCCTAGGCTCTTCGATTACAAACAATGATGGACGCAGCGGTCAGCTGAT GGACATCGTTGACAATGTCACTCCTGGTTTCTACCGCATAAGTTTCAATACCTCCAAGTA TGCACCGTCAGGGTTCTTCCCTTACGTGAGCATCATATTTGAGATAAAAAGAAGCCAGGC GACCGAGCATTTCCATGTGCCTCTTTTACATTCTCCTTTTTCCTTTACCACTTACCGTGG CAGCTAAGCTGTCAATTTGTGTGTAAATCACATCGAGTGATGGTATTGTTCTGAAACAAG CATAAATGGTCGTAAATCAGGGTGTTTAGATGTCACTGACATTGTGATATTTGAATAATA ATGTTCCATGTAATAATAAGCTCGAGTATGCAGTGACATTGGATCAGTTTGTTTTCAATA CTTTTATCCATTGCTATTTGTGATTGGGTGGCC
Protein Sequence (295 aa)
>BART1_0-u29736.006 295 150831_barley_pseudomolecules MATLRPLSVEEVLRVNGSHRFAAALAAASPFDSLANALLAARRIWLNEVDVTGWLEAFAA HPPIGSTSSSVSKWSKEEQSAALSTATDSTAQELSEWNAKYREKFGFVFMICASGRTAPE VLAELKRRYANRPVVELEAAAEEELKITELRLAKLFASETAAPPTSGNSNRTRPPITTHV LDTALGSPASGIEVHLEMWKDASSTPSFDNKDFKGWTTLGSSITNNDGRSGQLMDIVDNV TPGFYRISFNTSKYAPSGFFPYVSIIFEIKRSQATEHFHVPLLHSPFSFTTYRGS
