- Transcript BART1_0-u29736.005
- Transcript IDBART1_0-u29736.005
- Gene IDBART1_0-u29736
Exon Structure of BART1_0-u29736.005
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr4H | 1 | 483706267 | 483705915 | + |
| chr4H | 2 | 483706465 | 483706392 | + |
| chr4H | 3 | 483706600 | 483706543 | + |
| chr4H | 4 | 483706780 | 483706679 | + |
| chr4H | 5 | 483707382 | 483707226 | + |
| chr4H | 6 | 483712160 | 483711561 | + |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials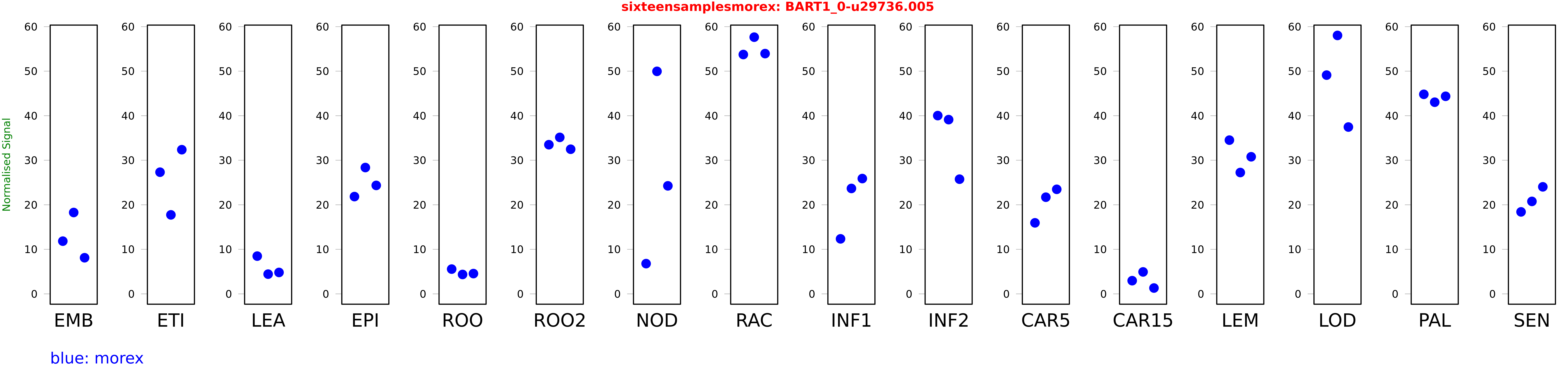
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 11.8342 | 18.2667 | 8.10782 |
| morex | ETI | 27.3176 | 17.7354 | 32.3683 |
| morex | LEA | 8.47519 | 4.42793 | 4.80803 |
| morex | EPI | 21.8394 | 28.3621 | 24.3521 |
| morex | ROO | 5.55836 | 4.35513 | 4.55036 |
| morex | ROO2 | 33.49 | 35.137 | 32.474 |
| morex | NOD | 6.78706 | 49.9637 | 24.2519 |
| morex | RAC | 53.7392 | 57.6352 | 53.9468 |
| morex | INF1 | 12.3504 | 23.6694 | 25.8889 |
| morex | INF2 | 40.0257 | 39.1213 | 25.7499 |
| morex | CAR5 | 15.9463 | 21.7029 | 23.4747 |
| morex | CAR15 | 2.95827 | 4.92564 | 1.3085 |
| morex | LEM | 34.5176 | 27.2451 | 30.7818 |
| morex | LOD | 49.1167 | 58.0283 | 37.46 |
| morex | PAL | 44.806 | 43.0322 | 44.3618 |
| morex | SEN | 18.4055 | 20.7604 | 24.031 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os03g27320.3 | +3 | 5e-136 | 394 | 223/261 (85%) | protein|steroid binding protein, putative, expressed |
| TAIR PP10 | AT5G58220 @ ARAPORT AT5G58220.3 @ TAIR |
+3 | 6e-81 | 253 | 147/272 (54%) | Symbols: TTL | transthyretin-like protein | chr5:23554546-23555861 REVERSE LENGTH=311 |
| BRACH PP3 | Bradi1g60700.3.p | +3 | 9e-144 | 413 | 232/260 (89%) | pacid=32804009 transcript=Bradi1g60700.3 locus=Bradi1g60700 ID=Bradi1g60700.3.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1344 bp)
>BART1_0-u29736.005 1344 150831_barley_pseudomolecules CTTACCCTTCCGGCAAAAACAGGTGTTGCTCTCGTAGCGCTCGCTCTCACCCACCCTTCC CCAACTCGCATCCAGCGCCGCAACCCCTCGTCTCTCCCCCCCTTCCGATCTGAGGCGTCG TTGGAGATCGGCGGCAACCGAAGAGGCGTCCGACGACGGCCGTCGCGGGTGGCACCAGTG CAGTGGAGCAAATCAACCAGTTGAGATGGCGACATTGAGGCCGCTGAGCGTGGAGGAGGT GCTGAGGGTGAACGGCAGCCACCGCTTTGCCGCCGCGCTGGCTGCCGCCTCCCCCTTTGA TTCTCTTGCCAACGCGCTCCTTGCCGCCCGCCGCATCTGGCTGAACGAGGTAGGTGGACG TGACTGGATGGCTGGAGGCCTTCGCTGCCCACCCACCCATCGGAAGCACCTCCTCCTCCG TCTCCAAGTGGAGCAAGGAAGAGCAATCAGCTGCTCTTTCCACAGCCACTGATTCGACTG CGCAGGAGTTGTCCGAGTGGAATGCCAAATACAGGGAGAAGTTTGGGTTCGTGTTCATGA TATGCGCATCTGGGAGGACTGCCCCTGAGGTCTTAGCTGAGCTTAAGAGGCGTTATGCAA ATAGACCAGTTGTTGAGCTTGAGGCTGCGGCAGAGGAAGAACTGAAGATAACTGAGCTGC GTCTTGCAAAGCTTTTCGCATCAGAAACTGCTGCTCCCCCTACTTCAGGTGAAAATCAGA TTAGCCAACCGGATAAAGCAGCAGGCAACTCCAACCGGACACGCCCTCCTATCACAACCC ATGTGCTGGACACGGCTCTTGGATCGCCAGCATCTGGAATTGAAGTTCACCTGGAGATGT GGAAGGACGCGTCAAGTACCCCGTCATTTGACAACAAAGATTTCAAGGGATGGACAACCC TAGGCTCTTCGATTACAAACAATGATGGACGCAGCGGTCAGCTGATGGACATCGTTGACA ATGTCACTCCTGGTTTCTACCGCATAAGTTTCAATACCTCCAAGTATGCACCGTCAGGGT TCTTCCCTTACGTGAGCATCATATTTGAGATAAAAAGAAGCCAGGCGACCGAGCATTTCC ATGTGCCTCTTTTACATTCTCCTTTTTCCTTTACCACTTACCGTGGCAGCTAAGCTGTCA ATTTGTGTGTAAATCACATCGAGTGATGGTATTGTTCTGAAACAAGCATAAATGGTCGTA AATCAGGGTGTTTAGATGTCACTGACATTGTGATATTTGAATAATAATGTTCCATGTAAT AATAAGCTCGAGTATGCAGTGACATTGGATCAGTTTGTTTTCAATACTTTTATCCATTGC TATTTGTGATTGGGTGGCCTTGAG
Protein Sequence (198 aa)
>BART1_0-u29736.005 198 150831_barley_pseudomolecules MICASGRTAPEVLAELKRRYANRPVVELEAAAEEELKITELRLAKLFASETAAPPTSGEN QISQPDKAAGNSNRTRPPITTHVLDTALGSPASGIEVHLEMWKDASSTPSFDNKDFKGWT TLGSSITNNDGRSGQLMDIVDNVTPGFYRISFNTSKYAPSGFFPYVSIIFEIKRSQATEH FHVPLLHSPFSFTTYRGS
