- Transcript BART1_0-u25435.029
- Transcript IDBART1_0-u25435.029
- Gene IDBART1_0-u25435
Exon Structure of BART1_0-u25435.029
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr3H | 1 | 681734115 | 681733818 | - |
| chr3H | 2 | 681735055 | 681734396 | - |
| chr3H | 3 | 681735390 | 681735231 | - |
| chr3H | 4 | 681735974 | 681735866 | - |
| chr3H | 5 | 681736132 | 681736050 | - |
| chr3H | 6 | 681737010 | 681736477 | - |
| chr3H | 7 | 681737704 | 681737606 | - |
| chr3H | 8 | 681737948 | 681737800 | - |
| chr3H | 9 | 681738516 | 681738306 | - |
| chr3H | 10 | 681738721 | 681738587 | - |
| chr3H | 11 | 681738879 | 681738807 | - |
| chr3H | 12 | 681739133 | 681738961 | - |
| chr3H | 13 | 681740011 | 681739754 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials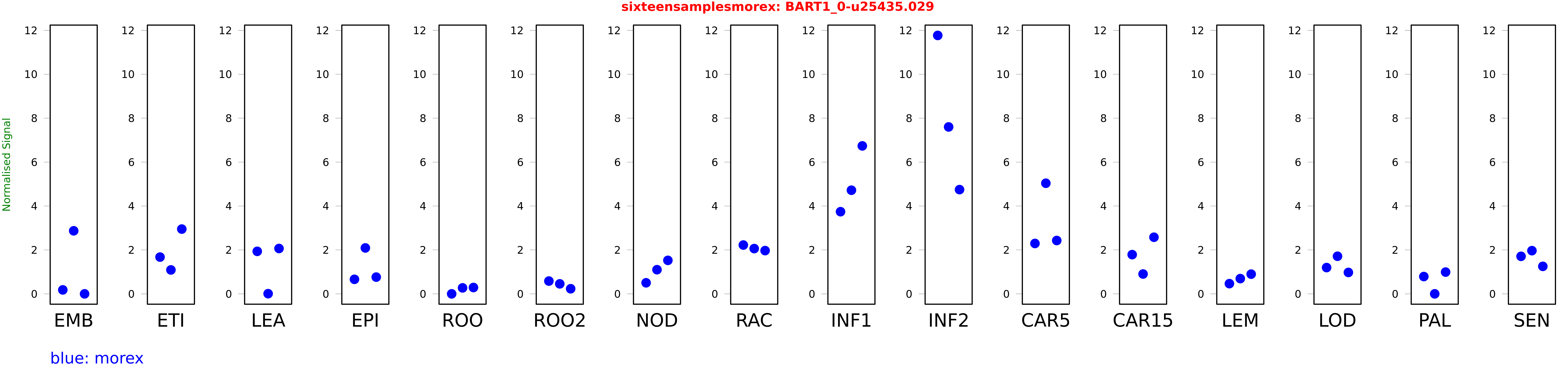
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0.180424 | 2.87094 | 0 |
| morex | ETI | 1.67424 | 1.08933 | 2.94996 |
| morex | LEA | 1.93618 | 0.00780349 | 2.06605 |
| morex | EPI | 0.662373 | 2.09103 | 0.762335 |
| morex | ROO | 0 | 0.272168 | 0.289305 |
| morex | ROO2 | 0.582903 | 0.455254 | 0.23265 |
| morex | NOD | 0.503387 | 1.10003 | 1.52539 |
| morex | RAC | 2.22298 | 2.06124 | 1.96985 |
| morex | INF1 | 3.74137 | 4.71918 | 6.73768 |
| morex | INF2 | 11.768 | 7.60123 | 4.74662 |
| morex | CAR5 | 2.29506 | 5.03725 | 2.4294 |
| morex | CAR15 | 1.78624 | 0.9033 | 2.57826 |
| morex | LEM | 0.464629 | 0.694813 | 0.897313 |
| morex | LOD | 1.19662 | 1.71269 | 0.975174 |
| morex | PAL | 0.788514 | 0 | 0.991163 |
| morex | SEN | 1.70883 | 1.96687 | 1.25258 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os01g72880.1 | +1 | 0.0 | 655 | 331/387 (86%) | protein|DNA mismatch repair protein Mlh1, putative, expressed |
| TAIR PP10 | AT4G09140 @ ARAPORT AT4G09140.1 @ TAIR |
+1 | 2e-178 | 536 | 265/350 (76%) | Symbols: ATMLH1, MLH1 | MUTL-homologue 1 | chr4:5817107-5821035 REVERSE LENGTH=737 |
| BRACH PP3 | Bradi2g61370.1.p | +1 | 0.0 | 700 | 354/388 (91%) | pacid=32771966 transcript=Bradi2g61370.1 locus=Bradi2g61370 ID=Bradi2g61370.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (2942 bp)
>BART1_0-u25435.029 2942 150831_barley_pseudomolecules NNGGTCCCCAGACGCGGCGCGGCGGGCATGGAGGTCGACGACGCGGCGCCGCGCGGCGGG GAGCCGCCCCGCATCCGGCGGCTGGAGGAGTCGGTGGTGAACCGCATCGCCGCGGGGGAG GTGATCCAGCGGCCCTCGTCGGCGGTGAAGGAACTCGTCGAGAACAGCATCGACGCCGGC TCCTCCACCATCTCCGTCACCGTCAAGGACGGGGGCCTCAAGCTCATCCAGGTCTCGGAC GACGGCCACGGCATCCGGTGTGAGGACTTGCCAATATTGTGTGAGAGGCATACTACCTCA AAGTTGTCTGCATATGAGGATCTACAGACAATAAAGTCCATGGGGTTCAGAGGGGAGGCT TTGGCCAGTATGACTTATGTTGGCCATGTTACCGTGACAACAATAACAGAAGGCCAACTG CATGGCTACAGAGTTTCTTATAGGGATGGAGTAATGGAGAATGACCCAAAGCCCTGTGCT GCGGTTAAAGGAACTCAAATCATGGTTGAAAATTTATTCTACAATATGGTAGCTCGTAGG AAAACACTGCAGAACTCCAATGATGACTATCCAAAGATAGTGGACTTCATCAGTCGGTTT GCAGTCCATCACATCAATGTGAACTTCTCTTGCAGAAAGCATGGAGCCAATAGAGCAGAT GTTCACAGTGGGAGCACATCTTCAAGGCTAGATGCTATCAGGAATGTCTATGGGGCTTCG GTTGTTCGTGATCTGATGGAAATACAGGTTTCAGATGAGAATGCTGTAGATGAAATATTC AAGATGGATGGTTTCATCTCAAATGCAAATTATGTGGCAAAGAAGACTACTATGATTCTT TTCATAAATGATCGACTTGTAGATTGCACTTCTTTGAAAAGAGCTACTGAATTTGTGTAC TCTGCAATACTGCCTCAAGCATCGAAACCCTTCATATACATGTCCATCAATCTTCCACCA GAACACGTGGATGTCAATATACATCCAACCAAAAAGGAGGTTAGCCTCTTAAATCAAGAG CGTATTATTGAAAAAATTAAAGATGCCATCGAGGAAAAACTGGTGAACTGTAATAACACA AGGATATTCCAAACTCAGGCACTAAACCCTTCAGCACTTACTCAAGCTAACACACGAAAG GACATGGGTACCGAGGTCAGCACGCCTACTGGTATGTACTCCCTCCGTTCCAAAATATAA GACCTTTTAGGAATTTCACTAAGAGACTACATACGGAGCAAAATGAGTGAATCTATACTC TAAAATATGTCTATGTACATCCGTATGTAGTTTATAGTACAATCTCTAAAAGGTCTTATA TTTAGGAACGGAGGGAGTAGCATATATCATATGCCTATTTCAAGCTAAAATGTTTCTTAA ATACACTATACTGCCACATCACATTACTTAACTGTGTATTATTGAGCTATCTAGTAAAGG GCGTGTGAAATCACCACCATCCAGTGAATAATCTTTAGTCTAATTATATCATACTGTAGG AGAAAAATCTCAAAAGATTCCTGTGAGCCAAATTGTCAGAACAGATCCTCGCGATCCATC TGGAAGGTTGCACACCTATTGGCAAGGTCAAACTTCAAATCTTGAAAAGAAGTCTGACCT TGTTGCCGTAAGATCAAGAAGGAATCCAAAAGATGCTGGTGATTTGTCCAGTCGCCATGA GCTTGTTACAGAAATAGATTCTAACTTACATCCTGGCCTTTTCGATATTGTTAAGAATTG CACATATGTTGGAGTTGCGGATGAAGTTTTCGCTTTGATACAACACAATACCCTCTTATA CCTTGTCAATGTGGTAAATGTTAGCAAAGAACTCATGTACCAACAAGCCTTGTGCCGTTT TGGGAACTTCAATGCTATAAAACTCAGTGAACCAGCTCCACTTCTGGAGTTGCTGAGAAT GGCACTGAAAGATGACGAGTCAATGAGTGATGTAAATGAGAAGGAGAAACTAGAAATTGC AGAAGTGAACACTGAGATACTGAAAGAAAATGCTGAGATGATTAATGAGTACTTCTCTAT TCACATTGATCAAGGTGGCAACCTGACCAGACTTCCTGTTGTACTTGATCAGTACACTCC TGATATGGATCGCCTTCCAGAGTTCATGTTGAGTCTAGGAAATGATGTAGGTTTCTATTA TGCTGCCGCGTTCATTACATATGGTATTTTAAATTTGATCATGATATTTGTTTATCCATT CCCTGCTGAGATGATGGCAAGTAGGTTTTCAGGCTGATACCGTGTTCAAATTTTAATTTT GGTGTTTCATGTAGATATGGCAGCCAGCTGTCCAGAAGTAACTTATGTTTCTTATTTTGT ACTGTATAAGTCACTTGGTAATTTCAGTTCCAGGAGTTGTTGCATTATGACATATTTTGT TCTAGATTGTACTTTAATTCTGCAAACTGTCAGTCCTAATTTTTAGTGATTACCATCTTA TAATAGATTACCTGGGACGTTGAGAAAGAGTGCTTCAGAACGGCAGCAGCTGCTATCGGA AACTTCTATGCGCTTCATCCTCCCATCCTTCCAAATCCATCTGGCAAAGGCATTCAGTTA TACAAGAAAAATAAAGATTGCATGGGAAGTGCTGAACAGGCTGATAACAATTTACCAAGT ACAGATGAAGATGACATCGATCAAGAGCTGCTTGCGGAAGCAGAGGCAGCATGGGCTCAA CGTGAGTGGACCATCCAGCACGTCTTGTTCCCGTCGATGCGACTTTTCCTCAAGCCCCCT AAGTCGATGGCAACTGATGGAACGTTTGTCCAGGTGCCCCTCCTAACCTCGGTTCAGCCC TCCTTCATAGTATATGTTTGTAGCTATACGTGCTTCAGTGTTTTACTTATGCTGGTGCAC CATAGTCTGTTACTGTGCTCCTGTCTATTTTTTACTGCTCAATGTCGGCACCACAATGCT GC
Protein Sequence (390 aa)
>BART1_0-u25435.029 390 150831_barley_pseudomolecules MEVDDAAPRGGEPPRIRRLEESVVNRIAAGEVIQRPSSAVKELVENSIDAGSSTISVTVK DGGLKLIQVSDDGHGIRCEDLPILCERHTTSKLSAYEDLQTIKSMGFRGEALASMTYVGH VTVTTITEGQLHGYRVSYRDGVMENDPKPCAAVKGTQIMVENLFYNMVARRKTLQNSNDD YPKIVDFISRFAVHHINVNFSCRKHGANRADVHSGSTSSRLDAIRNVYGASVVRDLMEIQ VSDENAVDEIFKMDGFISNANYVAKKTTMILFINDRLVDCTSLKRATEFVYSAILPQASK PFIYMSINLPPEHVDVNIHPTKKEVSLLNQERIIEKIKDAIEEKLVNCNNTRIFQTQALN PSALTQANTRKDMGTEVSTPTGMYSLRSKI
