- Transcript BART1_0-u20608.004
- Transcript IDBART1_0-u20608.004
- Gene IDBART1_0-u20608
Exon Structure of BART1_0-u20608.004
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr3H | 1 | 358149252 | 358149024 | + |
| chr3H | 2 | 358149679 | 358149564 | + |
| chr3H | 3 | 358149891 | 358149809 | + |
| chr3H | 4 | 358150386 | 358150265 | + |
| chr3H | 5 | 358152519 | 358152442 | + |
| chr3H | 6 | 358152688 | 358152595 | + |
| chr3H | 7 | 358153398 | 358153235 | + |
| chr3H | 8 | 358153926 | 358153493 | + |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials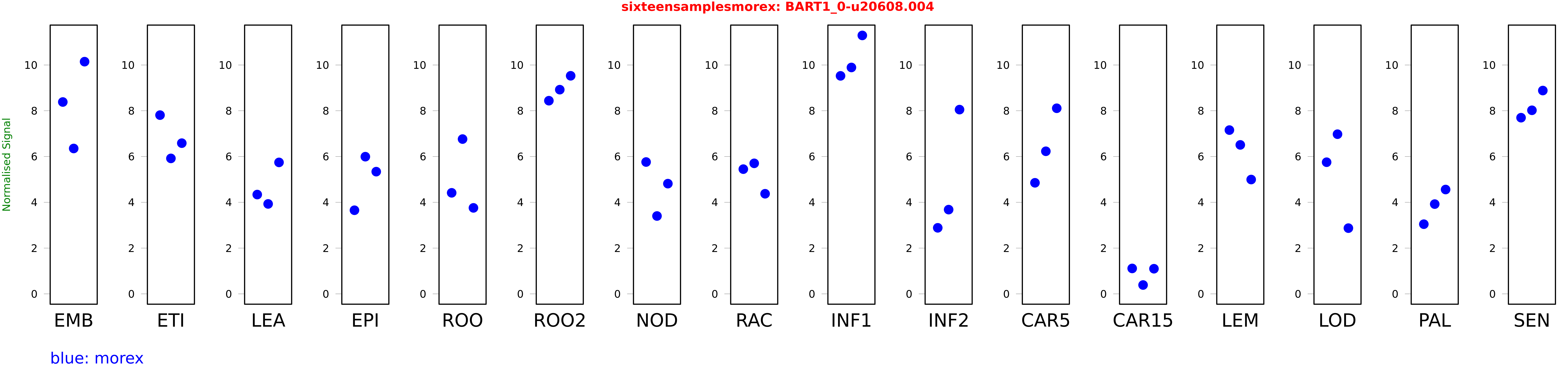
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 8.38247 | 6.35202 | 10.1453 |
| morex | ETI | 7.80934 | 5.91857 | 6.5845 |
| morex | LEA | 4.33841 | 3.92942 | 5.74361 |
| morex | EPI | 3.65329 | 5.99344 | 5.33945 |
| morex | ROO | 4.41443 | 6.76266 | 3.75662 |
| morex | ROO2 | 8.43956 | 8.92161 | 9.52775 |
| morex | NOD | 5.76024 | 3.40161 | 4.81523 |
| morex | RAC | 5.44912 | 5.70269 | 4.37361 |
| morex | INF1 | 9.52469 | 9.89145 | 11.2915 |
| morex | INF2 | 2.88549 | 3.68012 | 8.05011 |
| morex | CAR5 | 4.85217 | 6.22976 | 8.10931 |
| morex | CAR15 | 1.11079 | 0.386313 | 1.09774 |
| morex | LEM | 7.15554 | 6.50848 | 4.99498 |
| morex | LOD | 5.75155 | 6.97597 | 2.87163 |
| morex | PAL | 3.04433 | 3.92178 | 4.55847 |
| morex | SEN | 7.69693 | 8.02048 | 8.88319 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os01g39800.1 | +1 | 2e-78 | 247 | 133/229 (58%) | protein|esterase, putative, expressed |
| TAIR PP10 | AT1G29840 @ ARAPORT AT1G29840.1 @ TAIR |
+1 | 1e-59 | 172 | 91/190 (48%) | Symbols: | alpha/beta-Hydrolases superfamily protein | chr1:10445735-10447422 REVERSE LENGTH=263 |
| BRACH PP3 | Bradi2g41780.1.p | +1 | 8e-90 | 274 | 148/223 (66%) | pacid=32782108 transcript=Bradi2g41780.1 locus=Bradi2g41780 ID=Bradi2g41780.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1320 bp)
>BART1_0-u20608.004 1320 150831_barley_pseudomolecules CCAATGTAGCGCCTTCCTAGTGCGTTCTTTTCTTTGTACACCTTAAATTCCACGCCCTCC TCTGGCGTCCTCCACTTGCCCGGCGAATCTAACCTTCATCCTGCTGCTGCCACCGCCGGC GGGTCGACTAGACTCGGGTTCCTCCCACCCCAAACAGCACGAGGCTGCCGCCGCCATCTG TCTTCCATGTCACAGCCGCTCAATTCCTCCCGGCATGCTTCCAATCCCCATGCTACACAG AGGGTATTGATAACGAACAAGCGTGGAGAGAAACTTGTGGGGCTACTGCATCCCACGGGT TTGAACAAGATTGTAGTCTTGTGCCATGGCTTTACGGCCTCCAAGAGTTCTGGCATTATT GTTGATCTGGCAGATGCAATAATAAAAAAAGGGATTAGTGTCTTTCACTTTGATTTCAGT GGAAACGGAGAGAGCGAAGGCGAATTCCAGTATGGCAACTACAGGAAAGAGGCTGAAGAC TTGCATTCTGTTGTCTCATATCTGTATCAGGAGAAGTATGATGTAGCAGCTGTTGTTGGT CACAGCAAAGGCATTGAGGAACGCCTAGGGAAGGAATTTCTTGACAGAATAAATAAGGAA GGTTACATTGATGTCACAAACAAGTCAGGAAATATGTTATACAGGGTAACAAAAGAGAGC TTGATGGAACGACTCAACACTGACATGCGTGCAACGAGCCTCTCCATTAGCAAGGAATGC AGTTCATGGTTCTGCTGATGACATCATTCCGGTGGAAGATGCATATGAGTTTGCAAAGCA TATACCAACCCACAAACTCTGTGTCGTCGAGGGAGCAGATCACTGCTACACCAGACATCG TAAAGAACTTTCAGATGCTGTAGTGGGTTTCATTACATCCAACAAGGCCAGGGATACCAC GCTAGAAGATCAATGAATTCTGATGGATTTATGCTGTAGGATTTTGTACAGCAGTTCTCT TTACAGAAGCTCCCCTCAGAACAAGGCAATTCTAGATCATGGTCATTCTACTTCTTAGTT CCGGGATGTAATAAGCAATAAAGCATTGCAAGGCTTTAGGTTTGCGTTATTTTATGGGGA CTGCTATGCGTCGGTCCCTTCCAGTGGGATGCACATTTTTCTATTTTGCAACAAAGACCT TGTTGCAGAAACATTTATAACGGAGATTGTGTTGGAGAATTTTTATGTAACAGAGGTTGT GTTGCGATTTTTTTTTGCAAGAGAGACCGTGTTATAATTTTTTTGCAACATGAGTGTTGT TTTGGAAAAAAACACAAACGATGTTGCAGAAAATATTTGCAGTTTTTTTTTGKCAACGTA
Protein Sequence (183 aa)
>BART1_0-u20608.004 183 150831_barley_pseudomolecules MSQPLNSSRHASNPHATQRVLITNKRGEKLVGLLHPTGLNKIVVLCHGFTASKSSGIIVD LADAIIKKGISVFHFDFSGNGESEGEFQYGNYRKEAEDLHSVVSYLYQEKYDVAAVVGHS KGIEERLGKEFLDRINKEGYIDVTNKSGNMLYRVTKESLMERLNTDMRATSLSISKECSS WFC
