- Transcript BART1_0-u20608.003
- Transcript IDBART1_0-u20608.003
- Gene IDBART1_0-u20608
Exon Structure of BART1_0-u20608.003
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr3H | 1 | 358149252 | 358149013 | + |
| chr3H | 2 | 358149679 | 358149564 | + |
| chr3H | 3 | 358149891 | 358149809 | + |
| chr3H | 4 | 358150395 | 358150265 | + |
| chr3H | 5 | 358152519 | 358152355 | + |
| chr3H | 6 | 358152688 | 358152595 | + |
| chr3H | 7 | 358153398 | 358153224 | + |
| chr3H | 8 | 358153832 | 358153493 | + |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials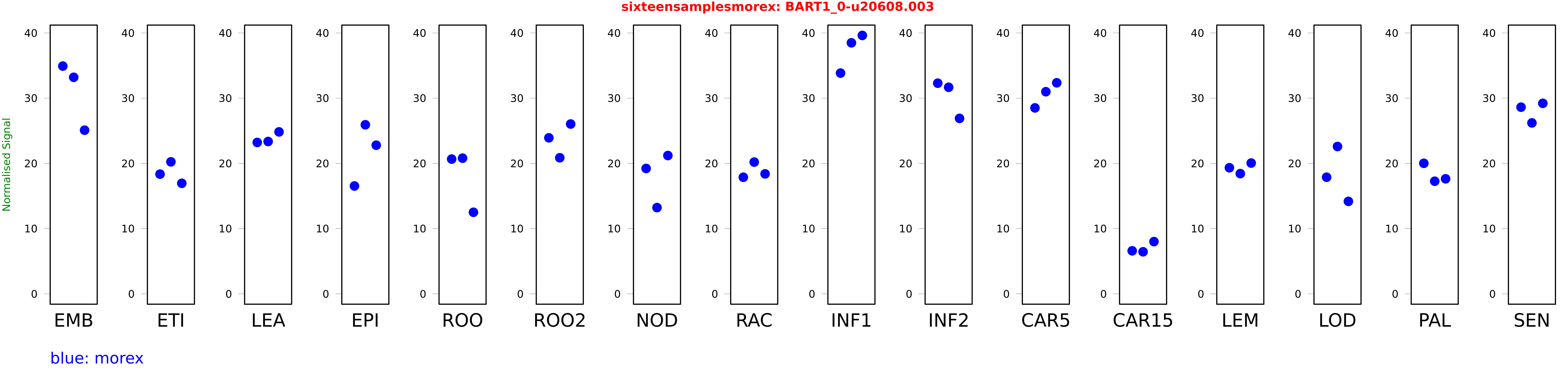
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 34.9176 | 33.1971 | 25.0884 |
| morex | ETI | 18.3532 | 20.2389 | 16.9471 |
| morex | LEA | 23.2085 | 23.3562 | 24.8316 |
| morex | EPI | 16.5361 | 25.92 | 22.7885 |
| morex | ROO | 20.6615 | 20.8011 | 12.5066 |
| morex | ROO2 | 23.9186 | 20.8634 | 26.0345 |
| morex | NOD | 19.2229 | 13.2243 | 21.2048 |
| morex | RAC | 17.8744 | 20.2006 | 18.3937 |
| morex | INF1 | 33.8427 | 38.4854 | 39.6176 |
| morex | INF2 | 32.2939 | 31.6674 | 26.9003 |
| morex | CAR5 | 28.508 | 30.9884 | 32.3574 |
| morex | CAR15 | 6.59708 | 6.44902 | 8.01015 |
| morex | LEM | 19.3327 | 18.4307 | 20.0522 |
| morex | LOD | 17.8793 | 22.5877 | 14.1792 |
| morex | PAL | 20.0136 | 17.266 | 17.6375 |
| morex | SEN | 28.6193 | 26.206 | 29.2097 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os01g39790.1 | +3 | 2e-140 | 404 | 203/276 (74%) | protein|esterase/lipase/thioesterase, putative, expressed |
| TAIR PP10 | AT1G29840 @ ARAPORT AT1G29840.1 @ TAIR |
+3 | 1e-97 | 293 | 143/250 (57%) | Symbols: | alpha/beta-Hydrolases superfamily protein | chr1:10445735-10447422 REVERSE LENGTH=263 |
| BRACH PP3 | Bradi2g41780.1.p | +3 | 1e-159 | 452 | 222/270 (82%) | pacid=32782108 transcript=Bradi2g41780.1 locus=Bradi2g41780 ID=Bradi2g41780.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1344 bp)
>BART1_0-u20608.003 1344 150831_barley_pseudomolecules ATTGGGTCCATCCAATGTAGCGCCTTCCTAGTGCGTTCTTTTCTTTGTACACCTTAAATT CCACGCCCTCCTCTGGCGTCCTCCACTTGCCCGGCGAATCTAACCTTCATCCTGCTGCTG CCACCGCCGGCGGGTCGACTAGACTCGGGTTCCTCCCACCCCAAACAGCACGAGGCTGCC GCCGCCATCTGTCTTCCATGTCACAGCCGCTCAATTCCTCCCGGCATGCTTCCAATCCCC ATGCTACACAGAGGGTATTGATAACGAACAAGCGTGGAGAGAAACTTGTGGGGCTACTGC ATCCCACGGGTTTGAACAAGATTGTAGTCTTGTGCCATGGCTTTACGGCCTCCAAGAGTT CTGGCATTATTGTTGATCTGGCAGATGCAATAATAAAAAAAGGGATTAGTGTCTTTCACT TTGATTTCAGTGGAAACGGAGAGAGCGAAGGCGAATTCCAGTATGGCAACTACAGGAAAG AGGCTGAAGACTTGCATTCTGTTGTCTCATATCTGTATCAGGAGAAGTATGATGTAGCAG CTGTTGTTGGTCACAGCAAAGGTTGGTGATGAGGAGATGTGGTTGTTCTTTATGCTTCCC TATATGACGATGTCCACATGGTTGTTAACCTATCTGGCCGATTTTATTTGGAGAAAGGCA TTGAGGAACGCCTAGGGAAGGAATTTCTTGACAGAATAAATAAGGAAGGTTACATTGATG TCACAAACAAGTCAGGAAATATGTTATACAGGGTAACAAAAGAGAGCTTGATGGAACGAC TCAACACTGACATGCGTGCAACGAGCCTCTCCATTAGCAAGGAATGCAGCTTCTTCACAG TTCATGGTTCTGCTGATGACATCATTCCGGTGGAAGATGCATATGAGTTTGCAAAGCATA TACCAACCCACAAACTCTGTGTCGTCGAGGGAGCAGATCACTGCTACACCAGACATCGTA AAGAACTTTCAGATGCTGTAGTGGGTTTCATTACATCCAACAAGGCCAGGGATACCACGC TAGAAGATCAATGAATTCTGATGGATTTATGCTGTAGGATTTTGTACAGCAGTTCTCTTT ACAGAAGCTCCCCTCAGAACAAGGCAATTCTAGATCATGGTCATTCTACTTCTTAGTTCC GGGATGTAATAAGCAATAAAGCATTGCAAGGCTTTAGGTTTGCGTTATTTTATGGGGACT GCTATGCGTCGGTCCCTTCCAGTGGGATGCACATTTTTCTATTTTGCAACAAAGACCTTG TTGCAGAAACATTTATAACGGAGATTGTGTTGGAGAATTTTTATGTAACAGAGGTTGTGT TGCGATTTTTTTTTGCAAGAGAGA
Protein Sequence (138 aa)
>BART1_0-u20608.003 138 150831_barley_pseudomolecules MVVNLSGRFYLEKGIEERLGKEFLDRINKEGYIDVTNKSGNMLYRVTKESLMERLNTDMR ATSLSISKECSFFTVHGSADDIIPVEDAYEFAKHIPTHKLCVVEGADHCYTRHRKELSDA VVGFITSNKARDTTLEDQ
