- Transcript BART1_0-u19417.003
- Transcript IDBART1_0-u19417.003
- Gene IDBART1_0-u19417
Exon Structure of BART1_0-u19417.003
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr3H | 1 | 156626354 | 156625511 | - |
| chr3H | 2 | 156627286 | 156627225 | - |
| chr3H | 3 | 156628029 | 156627930 | - |
| chr3H | 4 | 156628966 | 156628728 | - |
| chr3H | 5 | 156629835 | 156629587 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials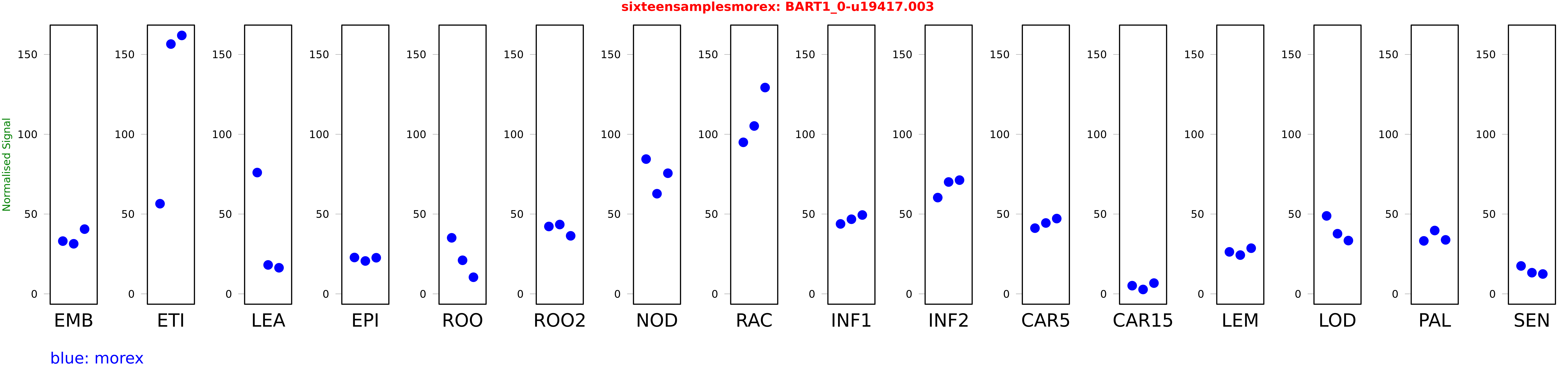
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 33.0488 | 31.3703 | 40.5542 |
| morex | ETI | 56.4725 | 156.505 | 161.91 |
| morex | LEA | 75.9935 | 18.1236 | 16.3434 |
| morex | EPI | 22.7792 | 20.5975 | 22.6252 |
| morex | ROO | 35.1087 | 21.0372 | 10.4077 |
| morex | ROO2 | 42.217 | 43.4337 | 36.3526 |
| morex | NOD | 84.4358 | 62.7414 | 75.5785 |
| morex | RAC | 94.9371 | 105.17 | 129.257 |
| morex | INF1 | 43.8184 | 46.7519 | 49.3969 |
| morex | INF2 | 60.309 | 70.0778 | 71.254 |
| morex | CAR5 | 41.169 | 44.3988 | 47.1626 |
| morex | CAR15 | 5.12749 | 2.75068 | 6.7352 |
| morex | LEM | 26.2994 | 24.2886 | 28.6159 |
| morex | LOD | 48.8445 | 37.6867 | 33.3787 |
| morex | PAL | 33.1859 | 39.6847 | 33.8233 |
| morex | SEN | 17.4894 | 13.2576 | 12.4422 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os05g14180.1 | +2 | 3e-99 | 300 | 168/237 (71%) | protein|OsIAA17 - Auxin-responsive Aux/IAA gene family member, expressed |
| TAIR PP10 | AT4G29080 @ ARAPORT AT4G29080.1 @ TAIR |
+2 | 1e-59 | 198 | 129/269 (48%) | Symbols: PAP2, IAA27 | phytochrome-associated protein 2 | chr4:14323665-14325213 REVERSE LENGTH=305 |
| BRACH PP3 | Bradi2g07770.1.p | +2 | 2e-130 | 379 | 206/227 (91%) | pacid=32771633 transcript=Bradi2g07770.1 locus=Bradi2g07770 ID=Bradi2g07770.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1494 bp)
>BART1_0-u19417.003 1494 150831_barley_pseudomolecules NNNNNNccgCGCGGACGACGACCTGAAAGGCACCGAGCTCCGCCTCGGACTTCCCGGCTG CGAGTCGCCGGACCGCCGCCCGGTCGCCGCCACCACCACACTGGAGCTTCTGCCCGCCAA GGGCGCCAAGCGAGGGTTCTCGGACGAGGTCCTGCCGCCTGCGCCGTCTGCCGCCGGCGG GAAAGGCAAGGAGACGTCCGGGGATGAGAAGGACAAGAAGGTCGCCGCACCGCCGCAGCC TGCTGCCAAGGCTCAGGTGGTGGGATGGCCACCAATCCGCAGCTACCGCAAGAACACGAT GGCAACCACCACCAACCAGCTGAAGAGCAGCAAGGAGGATTCTGACGCGAAGCAGGGGCA AGAGTTTCTGTATGTCAAAGTCAGCATGGATGGCGCGCCTTACCTGAGGAAGGTTGACCT CAAGACATACAAGAACTATAAGGACCTGTCTCTGGGGCTAGAGAAAATGTTCATTGGCTT CAGCACCGGTAAAGATGGTGTATCTGAGAACAGAAAGGATGGTGAATACGTGCTGACCTA CGAAGACAAGGACGGAGACTGGATGCTGGTTGGTGACGTCCCATGGGAGATGTTCACCGA GTCTTGCCGAAGGCTGAGGGTCATGAAAGGTTCAGATGCAGTTGGACTTGTTCCAAGGGC AGGGGATAAGTCCAAGAATAAGAACTAAAGCGGTTGCTGCCCACAGTTTGAAGACGGGGT CGCACATACCGTCTGTACTACCGCATGGTTTTTCTATCCCGCAAGTAAAATCACTCCGTT ACTCTACTGTTGTAAGATTTGTGATGTCTCCTAGAGACTTATGCTTTATATTTTGTCGTG CCGAAGTTGTTGTACGCATGCTATTTCCATCCGTGTAAATGGTTGGTAAGCAGTGCGAGA GCTACTGCTGTGATTTGATGACTTCTATGTATTTTGCCAGTGTTGTCCTGAACCGGCCGT TCGTCGTAAATGTGAGGGTTGCAAGACAATTATTTACCTGATGAACTTTGCTTCCGTACA CCCTCCTAGAAAAGTTTACAGATTTCGATGTAGGTTAGAAGTTACTGCACTTTAATTATT AAATAGAAAAAAGGGCAACATAGTCTTTCGTATCCAAATCAATCTATTCGGTATGTTTGA TCTGCAGGCTAATATGACGACAGTCTTTTTAAGTAAATTTAGGCTGCGTTTGGATTGAGT GTAAAATAATACAGAGGTATTTTTTTCTACACGTGTAACTTACCGGCCAATGTATAATTT CGGGCAGAAAATGAAACGACCGATCGACGCATGCATCTTTTACGGAGGTATTTCCAGCTG AAACGCTGATCCAAACAGGCCATTAAGATTCATGAAAACTTTTAAGGAGGGTGTACCGAA GCAAAACTCCTGACTTGTTCAAAAGTTTTTCCCGTTCCATTTATTATACTGTTTGCGGGT GAGTGGGTTGTTGCTTTGTTCTCCATTTCTTCTGAAACATATGCAATTGCATGG
Protein Sequence (228 aa)
>BART1_0-u19417.003 228 150831_barley_pseudomolecules XXRADDDLKGTELRLGLPGCESPDRRPVAATTTLELLPAKGAKRGFSDEVLPPAPSAAGG KGKETSGDEKDKKVAAPPQPAAKAQVVGWPPIRSYRKNTMATTTNQLKSSKEDSDAKQGQ EFLYVKVSMDGAPYLRKVDLKTYKNYKDLSLGLEKMFIGFSTGKDGVSENRKDGEYVLTY EDKDGDWMLVGDVPWEMFTESCRRLRVMKGSDAVGLVPRAGDKSKNKN
