- Transcript BART1_0-u19238.010
- Transcript IDBART1_0-u19238.010
- Gene IDBART1_0-u19238
Exon Structure of BART1_0-u19238.010
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr3H | 1 | 133980544 | 133980168 | - |
| chr3H | 2 | 133980745 | 133980629 | - |
| chr3H | 3 | 133981415 | 133980885 | - |
| chr3H | 4 | 133981597 | 133981505 | - |
| chr3H | 5 | 133984485 | 133981692 | - |
| chr3H | 6 | 133984718 | 133984615 | - |
| chr3H | 7 | 133985370 | 133985055 | - |
| chr3H | 8 | 133985932 | 133985769 | - |
| chr3H | 9 | 133988099 | 133988039 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials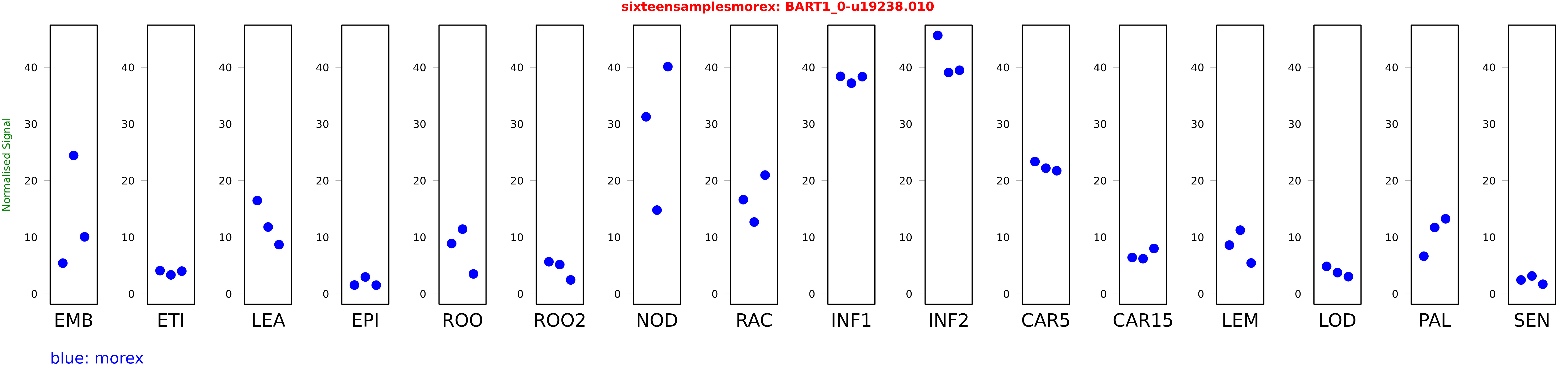
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 5.41978 | 24.433 | 10.067 |
| morex | ETI | 4.10056 | 3.35174 | 4.01069 |
| morex | LEA | 16.473 | 11.8003 | 8.69195 |
| morex | EPI | 1.55114 | 2.98109 | 1.53579 |
| morex | ROO | 8.88208 | 11.4252 | 3.51674 |
| morex | ROO2 | 5.67257 | 5.16925 | 2.46035 |
| morex | NOD | 31.2608 | 14.7842 | 40.115 |
| morex | RAC | 16.6301 | 12.6782 | 20.9667 |
| morex | INF1 | 38.409 | 37.1988 | 38.3439 |
| morex | INF2 | 45.6281 | 39.0807 | 39.4823 |
| morex | CAR5 | 23.354 | 22.186 | 21.7362 |
| morex | CAR15 | 6.42232 | 6.21023 | 8.00977 |
| morex | LEM | 8.60711 | 11.2441 | 5.44515 |
| morex | LOD | 4.85627 | 3.74195 | 3.02605 |
| morex | PAL | 6.64526 | 11.7175 | 13.2483 |
| morex | SEN | 2.45039 | 3.15981 | 1.70152 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os01g11040.1 | +2 | 0.0 | 1046 | 688/1128 (61%) | protein|SCAR-like protein 2, putative, expressed |
| TAIR PP10 | AT2G38440 @ ARAPORT AT2G38440.1 @ TAIR |
+3 | 2e-46 | 154 | 73/116 (63%) | Symbols: ITB1, SCAR2, DIS3, WAVE4, ATSCAR2 | SCAR homolog 2 | chr2:16095550-16100851 FORWARD LENGTH=1399 |
| BRACH PP3 | Bradi2g06610.2.p | +2 | 0.0 | 1159 | 720/1127 (64%) | pacid=32774244 transcript=Bradi2g06610.2 locus=Bradi2g06610 ID=Bradi2g06610.2.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (4557 bp)
>BART1_0-u19238.010 4557 150831_barley_pseudomolecules TCGAGGGCGTCGCCGTCGCGGGCCTCGTCGGGATCCTGCGCCAGCTCGGAGATCTCGCCG AATTTGCGGCAGATATTTTTCATGACCTACACGAGCAAGTTATAACTACATCGTCTAGGG GGCGTAAGGTGTTGACTCGAGTACAGAACATTGAAGCAGCACTTCCATCTCTTGAAAAAG CTGTGAAGAATCAGAAGAGCCATATACATTTTGCTTATGTACCAGGTTCTGACTGGCACA CACAACTTCAAAATGAGCAAAATCATCTCCTATCCAGTGATCTGCCTCGGTTTATGATGG ATTCCTATGAAGAATGTCGAGACCCACCACGACTTTACCTTCTCGACAAGTAAGGATGCT TGTTTTTTTTTTTCAAGTGATTTTGGGTCTCTAATGACTTATTTTCAGATTTGATAATGC TGGTGCTGGGGCTTGTTTGAAGAGATATTCTGATCCATCTTACTTCAAGAAATCATGGGA TGTGATGAGAGCAGACAAGACTGCACATCTCCAAAAAGAAAGGAGATCTCATAAAATCAA GAGGAAAGGATCACGCCTAAAAGAACCATATCATGGACAAGCCACATCCAGGCATAGGAA TGGTGAATTGCAGCGAGCACTCACTGCTGTTCAACTCACCAGCAGGCAGTGTGCATCTCC CAGCATGGATGGCCAGAGTTTTTCGGAGCATAGATCTACATCTGATGCAAGATCCAACCC TGACAATATAAGCAGATCTTCTTCATTTAGTTCAAAGGCACGACTGAGTTTTGCAGAACA AGCTTCAGATACAAAGCCTTCTGTAATTCCTCATGAGAATGGTCATGCCAAGCTGTCAAA TATTGATATGCACAAGCTTAATGATGCCTCTTCCCCCGTCTTGCTTAGCGAGACCAGGGA AGATGATCTGGGTGATGATTTGAAGCAAGGTTCCCTGTCAGATGAGATGAATGCTAGGTC ACCTTCTGTTGAATGGCATGAAAAGGCTGCAATTGTCATGACTACAAGTTCTGTCTACTG TGATGATGTTGTCATGGATAGGGCTGAAAATGCAGAAACTAAACATATTAAGCCCGTGCA GCGAGAAGTTGATTATAGGGAGATGCAGACCTTGGAGCAGCAAGGGGCCTTACTTCAGAA GGCAAAGTTGTTATTACTGTCATCAGGCTTAAACCACCGTGATGAAGTCCCCTGTGAAAC AGACAACTACATGGATGCACTTAACACACTTGAATCTGAGACAGAGACTGAAGTTGACTC TCAAACTAAAAAACAAGGGAAGCCAGTACATTCCTTCAATGCTCATGCACCTCAAATAGA GTTGACAGATAATGCCGTTTCACAGCTTCCTGATTCTTCTCCTGTTGAGTTTCCTGATAC AAGTCCAAATTCCAGAATGTTACATACTTTTGAAAGAAGTGCCGATTTCCCCAGTTTGTC AAGTGCAGATGCTCCGGATATTTCACAGCATTCATTGTCAGGTTATACAGATATACATCC TAATGAATGGTCATCTGTTACCACCATCCCTGAGAATAATGCAAATAATGCTGCTGGAGA CCCCACTGAAATTTCAGAACCGGCACTGCAAGCATATACACCTACACCACCCAATCAAAG GTGCCCTCGTGCAAATGAGATTCCTGAGAGTAAGGCAGAAGTTATCCCCAGAGATTCTCC TGTGATCTCAAAACCAGAGCTGTCAACCTATGCAGTTTTCCCTCCCAATAAAGAGTCTAT TGTCAGTCAGATCCCTGAGAATAGTGTAGAAGGTGTGTCAGAAGATGATACCAATGAAAT CACTAGTTGTCTTGTACCGGAACCTTCCATTTCCTTTGCTCCAACTAGTGAGGCGTCTCC TGCAAAAATCTTACCTGGTTATAACACTGGTAACTCTGTAATTTCGGAAAGGAGTCCTCA GGATTATCCTGGAGAAAATCATCAGGAATTTGGTGGCTGTGGCATGGCTGAAGTGTCTAA TTCACAGACCATGCCTTTGAACGAGTCATCAGAGAATGGATCTGCAACCCAACACCTTCC CACACATGCTCCTACTAGTTCCCAACACCTTCCCACACATGCTCCGACTAGTTCTGTAGA ATTATCTTCTGTTAAGCTCTGGACAAATGCTGGTCTCTTCGGACTTGAACCGTCTAAACC TCCAGTTGGCATATGTCCAGGTTCTGCAAACCAAAGCTACCCAGAGAATAATCAATCAGC CACAAGAACACCTGATGCAGTTTATAGCCAAACAAACCGGCCCTCAGATTCTTCCACATA TTTCGAGCACAGGGAGCACAGCAATTTGAATGGCAAGCAAGCTTCAATAAGTGAGCTCCT AGAATCTGAAGGTAATGCTGAAATTGGTGCTGAAACATACTCTGGCACTGACTTGGTTGG AAGGAATAACTTGCCCGTGGTGTCTGCATCAAGCTTCTCGAGCATTGCACAAAGATTCCT TGCCAATACACTTCAGAGAAGAACTTCTCCCAAATATAATGATCTTCCTATGTCATCTGG GAGAGTGAATACTGACACAAATGGGTTTGATGAAGCTACTGTAAATTCTACTCTTGCACC AAAGGAAACAGTATTCGAGGCATCTCAGTTTGAGAAGAAAACAGATTATGGCGTGGATGG ACTGCCCAAGTCCTCTAGTTGCCATTACTCTGAGAAATCATCTCCGCCACTAGAGCATAT GAAAATATCCTTTCACCCCATGAGTGCATCTGAAATGTCGAAATTGAACCTAGATTTTTC TGATGGCAATCTTCATGAAAATGTTGATGATATGATGTTACCAACATTTCAGTTACTTCC AGGGTCTTCTGTCCCACAGCCTGGCAGTGGTTCTGAATCAGAAGATGACACCTTTGGTAG ATCTTATAGTTATTCTTCATATGATGATCTAAGTCCGCGTTTATATTCAAACTCAGAGTT GTGGGATCAAGAAGATGGAATTGGATTCGAGGATAATGAGCTGTATAATGATTCAAATCA AATAGGATCCTCCACAACGCCTGTCTCTAGCTATATGGGATTCGAGCAGATGAACTTATC TGGTGAGAAGTCCACTATTTCACTTTCAGATATTGGAGATAATAACGGACTCGGCTCGTT AGAATCCCATCCTGCTGGAGAACTTCCGAACTTTGACACTTTGATGTCAACAAGTAATCA TCAAAATAGAGATGCGCTCATTCCACAAAATCCAATCAATTTATCACCAGTTGTAGATCA GATGCCGCCGCCACCTCCTCTTCCCCCAATGCAGTGGAGGACAATGAGACAAACAACTTC CCTAGAAGAAGAAAGAGACTCTACGGCTAAAGACATGCTTGAGAATGCCTCAAGTCTACC ACCTGTACACACTCCTGTGCAGGAACAACATCTTCCACCGTGTGCACCACCATTTCCACA AGGAAATGTTAAGGAAGTGAACCATCAGAAAGTTGATGTGATCAAAGAAACGAGTAATCT TCCTAATATATTTGAGATCAAATCAAGCTTGCTTCAGCAAATCAGGGATAAGGAAAAAGT GCAAGTGAAGAACCCACTTCCTCTTCCCCAGTCTGAAAGTTCGTTTTTGGATAATTATTT GGTAATTCTATTGATACAAGTGACGAGTTGGCTAAATTTGTTAAAAATGAAACTTCGTGT GTGTGATATGTGTAGTCAGACCAATAGAATCAAAGTTATGGGTTCATCACTGCGAGTGAG TTAGCATTTTATGTTCATAAAATTAATGCCATAACAAGTACTCGCTCTGTAACTTAATAC AAGATGTTTTTGACACTGCCATAGACTCTAAAAACGTCTTATAAAAAGTTACAGAGGTAG TAACATTTATAATAACGGTTAGTTGACTCTTAACTTGCAAGGTCATTACCTAGTTTGTAG CATTTGACTGTCATCGATACTTACGTTTCTCCTCACTTTACATATCAAACTATTGCAGCC AGATCTGCACAAGCTGAATGGACATGAAAAGTCCAAAGCAGTATTCAGTGATGTTAACAG TTTGGATGAAAGGGGGGAGTTGCTTCAGCAAATCAGGAGCAAGACATTCAACTTAAGACG AACAAATGGATCTAAGACAAACACCTCATCGCAGTCAACCGAGTCCACTGCCAATTCCAG CGTTGTAGCAATTTTGGAGAAGGCGAATGCAATCCGACAGGCTGTGGCCAGTGACGAGGG GGGTGATGATGATAACTGGAGCGATATCAACTGAGAATGCATATGCCAAACCTTTTCTGT ATATGCAACTAGTACAAGATGAAGAGTGAAAATGTGAATAACACCTTTTTCTTCATTGTA ATTAGATATTGCAGCTCTGTTTTTTAACCTTTTTTTAGTTCATGGTAGTGTATCTTAGAT CCCTATGGTTGTAGAGATGTTAAATCATTTCAACATCATGTAAATTTATCTTATTATTTA TTAATGGTAGTGAGATGGACTAATAAAATAAGTATGTTAAGCTATCTTTTGTAGCATTGT ACTAAAAATTGTCTCTTTGCATTGTTAATAAGATGGGGGCATACTGATAAAAAAGTT
Protein Sequence (1079 aa)
>BART1_0-u19238.010 1079 150831_barley_pseudomolecules MRADKTAHLQKERRSHKIKRKGSRLKEPYHGQATSRHRNGELQRALTAVQLTSRQCASPS MDGQSFSEHRSTSDARSNPDNISRSSSFSSKARLSFAEQASDTKPSVIPHENGHAKLSNI DMHKLNDASSPVLLSETREDDLGDDLKQGSLSDEMNARSPSVEWHEKAAIVMTTSSVYCD DVVMDRAENAETKHIKPVQREVDYREMQTLEQQGALLQKAKLLLLSSGLNHRDEVPCETD NYMDALNTLESETETEVDSQTKKQGKPVHSFNAHAPQIELTDNAVSQLPDSSPVEFPDTS PNSRMLHTFERSADFPSLSSADAPDISQHSLSGYTDIHPNEWSSVTTIPENNANNAAGDP TEISEPALQAYTPTPPNQRCPRANEIPESKAEVIPRDSPVISKPELSTYAVFPPNKESIV SQIPENSVEGVSEDDTNEITSCLVPEPSISFAPTSEASPAKILPGYNTGNSVISERSPQD YPGENHQEFGGCGMAEVSNSQTMPLNESSENGSATQHLPTHAPTSSQHLPTHAPTSSVEL SSVKLWTNAGLFGLEPSKPPVGICPGSANQSYPENNQSATRTPDAVYSQTNRPSDSSTYF EHREHSNLNGKQASISELLESEGNAEIGAETYSGTDLVGRNNLPVVSASSFSSIAQRFLA NTLQRRTSPKYNDLPMSSGRVNTDTNGFDEATVNSTLAPKETVFEASQFEKKTDYGVDGL PKSSSCHYSEKSSPPLEHMKISFHPMSASEMSKLNLDFSDGNLHENVDDMMLPTFQLLPG SSVPQPGSGSESEDDTFGRSYSYSSYDDLSPRLYSNSELWDQEDGIGFEDNELYNDSNQI GSSTTPVSSYMGFEQMNLSGEKSTISLSDIGDNNGLGSLESHPAGELPNFDTLMSTSNHQ NRDALIPQNPINLSPVVDQMPPPPPLPPMQWRTMRQTTSLEEERDSTAKDMLENASSLPP VHTPVQEQHLPPCAPPFPQGNVKEVNHQKVDVIKETSNLPNIFEIKSSLLQQIRDKEKVQ VKNPLPLPQSESSFLDNYLVILLIQVTSWLNLLKMKLRVCDMCSQTNRIKVMGSSLRVS
