- Transcript BART1_0-u19238.007
- Transcript IDBART1_0-u19238.007
- Gene IDBART1_0-u19238
Exon Structure of BART1_0-u19238.007
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr3H | 1 | 133980544 | 133980056 | - |
| chr3H | 2 | 133980745 | 133980629 | - |
| chr3H | 3 | 133980989 | 133980885 | - |
| chr3H | 4 | 133981597 | 133981505 | - |
| chr3H | 5 | 133984481 | 133981692 | - |
| chr3H | 6 | 133984718 | 133984615 | - |
| chr3H | 7 | 133985153 | 133985055 | - |
| chr3H | 8 | 133985370 | 133985247 | - |
| chr3H | 9 | 133985932 | 133985769 | - |
| chr3H | 10 | 133988750 | 133988039 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials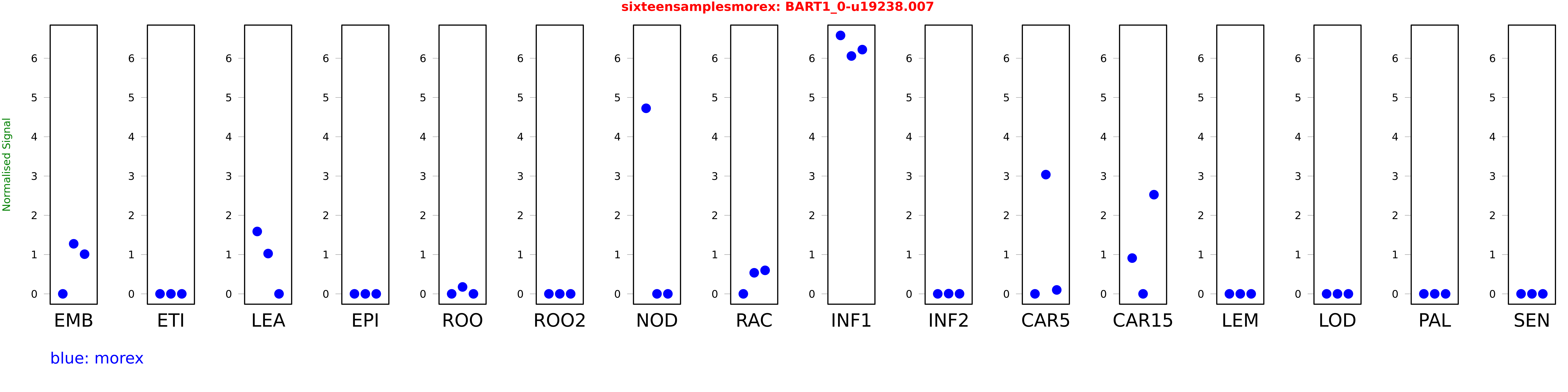
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0.0000017288 | 1.27456 | 1.01036 |
| morex | ETI | 0.000000000149408 | 0 | 0 |
| morex | LEA | 1.58864 | 1.02596 | 0 |
| morex | EPI | 0.000140742 | 0.00000000674792 | 0 |
| morex | ROO | 0 | 0.175863 | 0 |
| morex | ROO2 | 0.000000327348 | 0 | 0 |
| morex | NOD | 4.72504 | 0.00000000963015 | 0.0000000316727 |
| morex | RAC | 0 | 0.536171 | 0.598843 |
| morex | INF1 | 6.58031 | 6.05651 | 6.21903 |
| morex | INF2 | 0.0000507727 | 0.00547554 | 0.00195361 |
| morex | CAR5 | 0.00000000199665 | 3.03638 | 0.0999721 |
| morex | CAR15 | 0.910797 | 0.0000000014685 | 2.52539 |
| morex | LEM | 0 | 0 | 0 |
| morex | LOD | 0 | 0.00000000101845 | 0 |
| morex | PAL | 0 | 0 | 0 |
| morex | SEN | 0 | 0 | 0.0000000252032 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os01g11040.1 | +1 | 0.0 | 990 | 679/1121 (61%) | protein|SCAR-like protein 2, putative, expressed |
| TAIR PP10 | AT4G18600 @ ARAPORT AT4G18600.1 @ TAIR |
+3 | 1e-41 | 162 | 82/161 (51%) | Symbols: WAVE5, ATSCAR-LIKE, SCARL | SCAR family protein | chr4:10239947-10247372 REVERSE LENGTH=2028 |
| BRACH PP3 | Bradi2g06610.2.p | +1 | 0.0 | 1118 | 712/1130 (63%) | pacid=32774244 transcript=Bradi2g06610.2 locus=Bradi2g06610 ID=Bradi2g06610.2.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (4797 bp)
>BART1_0-u19238.007 4797 150831_barley_pseudomolecules CCGGCCCCCGCGCGACCCAGCACAGCACAGCACAGCACAGCACAGCAGCCTCCCACCACA ACATAGCGGCGGCACCGGGTATCGAGCATCCAAATCCAAGCCGAGCCCGAGAGCTGAAGC ATAATCACGCGGCGCACACCACTCACCAGTCGCCAGGCCGCGCCACTTGCCCACCCCCCT CGTGCCCGCAGCGCTGCGTGCGCTTCCCTTTCCCACCGTCCCTTTCGCCCCGCACCCTGT CGGAGACCGACGCTCCTCCCCTCCCCTCCCCGTCCCCGTCCCCGCCACGCGCCTCGGCGT CGCCTCCACTCCTCCCCCCCTCCCCGCCACTCGCCTCGGCCGCCCCCTTCCGTGCGGAGG CGGCAACCTGTGACGCCCCAAAGTCGGCGCACCTGCTCCCACTGGGAAAGTAGCCGTTGC GGCGTCGCATTTATAGCGACCGCCAGGCGTCACGATGGCCGCTGCTCGGCGGTGAGGAGG AGCCCGCTCTCGCGCGCTCGGTGGTTGGTGGAGCGGAGATGCCGCTGGTCAGGTTCGAGG TGCGGAATGAGGTCGGGCTGGGCGACCCCGGGCTGTACGGCGCCGCGGGGGCAGGGAAGC GAGGCGGCGCCGGCGCCGCCGCCGCAGGGGAGGCCGAGCCCAAGGCGCTGCTCGAGGGCG TCGCCGTCGCGGGCCTCGTCGGGATCCTGCGCCAGCTCGGAGATCTCGCCGAATTTGCGG CAGATATTTTTCATGACCTACACGAGCAAGTTATAACTACATCGTCTAGGGGGCGTAAGG TGTTGACTCGAGTACAGAACATTGAAGCAGCACTTCCATCTCTTGAAAAAGCTGTGAAGA ATCAGAAGAGCCATATACATTTTGCTTATGTACCAGGTTCTGACTGGCACACACAACTTC AAAATGAGCAAAATCATCTCCTATCCAGTGATCTGCCTCGGTTTATGATGGATTCCTATG AAGAATGTCGAGACCCACCACGACTTTACCTTCTCGACAAAGATATTCTGATCCATCTTA CTTCAAGAAATCATGGGATGTGATGAGAGCAGACAAGACTGCACATCTCCAAAAAGAAAG GAGATCTCATAAAATCAAGAGGAAAGGATCACGCCTAAAAGAACCATATCATGGACAAGC CACATCCAGGCATAGGAATGGTGAATTGCAGCGAGCACTCACTGCTGTTCAACTCACCAG CAGTGTGCATCTCCCAGCATGGATGGCCAGAGTTTTTCGGAGCATAGATCTACATCTGAT GCAAGATCCAACCCTGACAATATAAGCAGATCTTCTTCATTTAGTTCAAAGGCACGACTG AGTTTTGCAGAACAAGCTTCAGATACAAAGCCTTCTGTAATTCCTCATGAGAATGGTCAT GCCAAGCTGTCAAATATTGATATGCACAAGCTTAATGATGCCTCTTCCCCCGTCTTGCTT AGCGAGACCAGGGAAGATGATCTGGGTGATGATTTGAAGCAAGGTTCCCTGTCAGATGAG ATGAATGCTAGGTCACCTTCTGTTGAATGGCATGAAAAGGCTGCAATTGTCATGACTACA AGTTCTGTCTACTGTGATGATGTTGTCATGGATAGGGCTGAAAATGCAGAAACTAAACAT ATTAAGCCCGTGCAGCGAGAAGTTGATTATAGGGAGATGCAGACCTTGGAGCAGCAAGGG GCCTTACTTCAGAAGGCAAAGTTGTTATTACTGTCATCAGGCTTAAACCACCGTGATGAA GTCCCCTGTGAAACAGACAACTACATGGATGCACTTAACACACTTGAATCTGAGACAGAG ACTGAAGTTGACTCTCAAACTAAAAAACAAGGGAAGCCAGTACATTCCTTCAATGCTCAT GCACCTCAAATAGAGTTGACAGATAATGCCGTTTCACAGCTTCCTGATTCTTCTCCTGTT GAGTTTCCTGATACAAGTCCAAATTCCAGAATGTTACATACTTTTGAAAGAAGTGCCGAT TTCCCCAGTTTGTCAAGTGCAGATGCTCCGGATATTTCACAGCATTCATTGTCAGGTTAT ACAGATATACATCCTAATGAATGGTCATCTGTTACCACCATCCCTGAGAATAATGCAAAT AATGCTGCTGGAGACCCCACTGAAATTTCAGAACCGGCACTGCAAGCATATACACCTACA CCACCCAATCAAAGGTGCCCTCGTGCAAATGAGATTCCTGAGAGTAAGGCAGAAGTTATC CCCAGAGATTCTCCTGTGATCTCAAAACCAGAGCTGTCAACCTATGCAGTTTTCCCTCCC AATAAAGAGTCTATTGTCAGTCAGATCCCTGAGAATAGTGTAGAAGGTGTGTCAGAAGAT GATACCAATGAAATCACTAGTTGTCTTGTACCGGAACCTTCCATTTCCTTTGCTCCAACT AGTGAGGCGTCTCCTGCAAAAATCTTACCTGGTTATAACACTGGTAACTCTGTAATTTCG GAAAGGAGTCCTCAGGATTATCCTGGAGAAAATCATCAGGAATTTGGTGGCTGTGGCATG GCTGAAGTGTCTAATTCACAGACCATGCCTTTGAACGAGTCATCAGAGAATGGATCTGCA ACCCAACACCTTCCCACACATGCTCCTACTAGTTCCCAACACCTTCCCACACATGCTCCG ACTAGTTCTGTAGAATTATCTTCTGTTAAGCTCTGGACAAATGCTGGTCTCTTCGGACTT GAACCGTCTAAACCTCCAGTTGGCATATGTCCAGGTTCTGCAAACCAAAGCTACCCAGAG AATAATCAATCAGCCACAAGAACACCTGATGCAGTTTATAGCCAAACAAACCGGCCCTCA GATTCTTCCACATATTTCGAGCACAGGGAGCACAGCAATTTGAATGGCAAGCAAGCTTCA ATAAGTGAGCTCCTAGAATCTGAAGGTAATGCTGAAATTGGTGCTGAAACATACTCTGGC ACTGACTTGGTTGGAAGGAATAACTTGCCCGTGGTGTCTGCATCAAGCTTCTCGAGCATT GCACAAAGATTCCTTGCCAATACACTTCAGAGAAGAACTTCTCCCAAATATAATGATCTT CCTATGTCATCTGGGAGAGTGAATACTGACACAAATGGGTTTGATGAAGCTACTGTAAAT TCTACTCTTGCACCAAAGGAAACAGTATTCGAGGCATCTCAGTTTGAGAAGAAAACAGAT TATGGCGTGGATGGACTGCCCAAGTCCTCTAGTTGCCATTACTCTGAGAAATCATCTCCG CCACTAGAGCATATGAAAATATCCTTTCACCCCATGAGTGCATCTGAAATGTCGAAATTG AACCTAGATTTTTCTGATGGCAATCTTCATGAAAATGTTGATGATATGATGTTACCAACA TTTCAGTTACTTCCAGGGTCTTCTGTCCCACAGCCTGGCAGTGGTTCTGAATCAGAAGAT GACACCTTTGGTAGATCTTATAGTTATTCTTCATATGATGATCTAAGTCCGCGTTTATAT TCAAACTCAGAGTTGTGGGATCAAGAAGATGGAATTGGATTCGAGGATAATGAGCTGTAT AATGATTCAAATCAAATAGGATCCTCCACAACGCCTGTCTCTAGCTATATGGGATTCGAG CAGATGAACTTATCTGGTGAGAAGTCCACTATTTCACTTTCAGATATTGGAGATAATAAC GGACTCGGCTCGTTAGAATCCCATCCTGCTGGAGAACTTCCGAACTTTGACACTTTGATG TCAACAAGTAATCATCAAAATAGAGATGCGCTCATTCCACAAAATCCAATCAATTTATCA CCAGTTGTAGATCAGATGCCGCCGCCACCTCCTCTTCCCCCAATGCAGTGGAGGACAATG AGACAAACAACTTCCCTAGAAGAAGAAAGAGACTCTACGGCTAAAGACATGCTTGAGAAT GCCTCAAGTCTACCACCTGTACACACTCCTGTGCAGGAACAACATCTTCCACCGTGTGCA CCACCATTTCCACAAGGAAATGTTAAGGAAGTGAACCATCAGAAAGTTGATGTGATCAAA GAAACGAGTAATCTTCCTAATATATTTGAGATCAAATCAAGCTTGCTTCAGCAAATCAGG GATAAGCCAGATCTGCACAAGCTGAATGGACATGAAAAGTCCAAAGCAGTATTCAGTGAT GTTAACAGTTTGGATGAAAGGGGGGAGTTGCTTCAGCAAATCAGGAGCAAGACATTCAAC TTAAGACGAACAAATGGATCTAAGACAAACACCTCATCGCAGTCAACCGAGTCCACTGCC AATTCCAGCGTTGTAGCAATTTTGGAGAAGGCGAATGCAATCCGACAGGCTGTGGCCAGT GACGAGGGGGGTGATGATGATAACTGGAGCGATATCAACTGAGAATGCATATGCCAAACC TTTTCTGTATATGCAACTAGTACAAGATGAAGAGTGAAAATGTGAATAACACCTTTTTCT TCATTGTAATTAGATATTGCAGCTCTGTTTTTTAACCTTTTTTTAGTTCATGGTAGTGTA TCTTAGATCCCTATGGTTGTAGAGATGTTAAATCATTTCAACATCATGTAAATTTATCTT ATTATTTATTAATGGTAGTGAGATGGACTAATAAAATAAGTATGTTAAGCTATCTTTTGT AGCATTGTACTAAAAATTGTCTCTTTGCATTGTTAATAAGATGGGGGCATACTGATAAAA AAGTTATGCTGTCATTTGCAAGCATGCTATTTTTTTTGGCGAAGGCAAAATAATTCTTAA GAAATCTCGCTGCATTACCTCCATGCTTTGTATTTGAGTAGCAAAATCTAAATCAGG
Protein Sequence (1047 aa)
>BART1_0-u19238.007 1047 150831_barley_pseudomolecules MDGQSFSEHRSTSDARSNPDNISRSSSFSSKARLSFAEQASDTKPSVIPHENGHAKLSNI DMHKLNDASSPVLLSETREDDLGDDLKQGSLSDEMNARSPSVEWHEKAAIVMTTSSVYCD DVVMDRAENAETKHIKPVQREVDYREMQTLEQQGALLQKAKLLLLSSGLNHRDEVPCETD NYMDALNTLESETETEVDSQTKKQGKPVHSFNAHAPQIELTDNAVSQLPDSSPVEFPDTS PNSRMLHTFERSADFPSLSSADAPDISQHSLSGYTDIHPNEWSSVTTIPENNANNAAGDP TEISEPALQAYTPTPPNQRCPRANEIPESKAEVIPRDSPVISKPELSTYAVFPPNKESIV SQIPENSVEGVSEDDTNEITSCLVPEPSISFAPTSEASPAKILPGYNTGNSVISERSPQD YPGENHQEFGGCGMAEVSNSQTMPLNESSENGSATQHLPTHAPTSSQHLPTHAPTSSVEL SSVKLWTNAGLFGLEPSKPPVGICPGSANQSYPENNQSATRTPDAVYSQTNRPSDSSTYF EHREHSNLNGKQASISELLESEGNAEIGAETYSGTDLVGRNNLPVVSASSFSSIAQRFLA NTLQRRTSPKYNDLPMSSGRVNTDTNGFDEATVNSTLAPKETVFEASQFEKKTDYGVDGL PKSSSCHYSEKSSPPLEHMKISFHPMSASEMSKLNLDFSDGNLHENVDDMMLPTFQLLPG SSVPQPGSGSESEDDTFGRSYSYSSYDDLSPRLYSNSELWDQEDGIGFEDNELYNDSNQI GSSTTPVSSYMGFEQMNLSGEKSTISLSDIGDNNGLGSLESHPAGELPNFDTLMSTSNHQ NRDALIPQNPINLSPVVDQMPPPPPLPPMQWRTMRQTTSLEEERDSTAKDMLENASSLPP VHTPVQEQHLPPCAPPFPQGNVKEVNHQKVDVIKETSNLPNIFEIKSSLLQQIRDKPDLH KLNGHEKSKAVFSDVNSLDERGELLQQIRSKTFNLRRTNGSKTNTSSQSTESTANSSVVA ILEKANAIRQAVASDEGGDDDNWSDIN
