- Transcript BART1_0-u19238.006
- Transcript IDBART1_0-u19238.006
- Gene IDBART1_0-u19238
Exon Structure of BART1_0-u19238.006
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr3H | 1 | 133980544 | 133980054 | - |
| chr3H | 2 | 133980745 | 133980629 | - |
| chr3H | 3 | 133980989 | 133980885 | - |
| chr3H | 4 | 133981597 | 133981505 | - |
| chr3H | 5 | 133984488 | 133981692 | - |
| chr3H | 6 | 133984718 | 133984615 | - |
| chr3H | 7 | 133985370 | 133985055 | - |
| chr3H | 8 | 133985932 | 133985769 | - |
| chr3H | 9 | 133988797 | 133988039 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials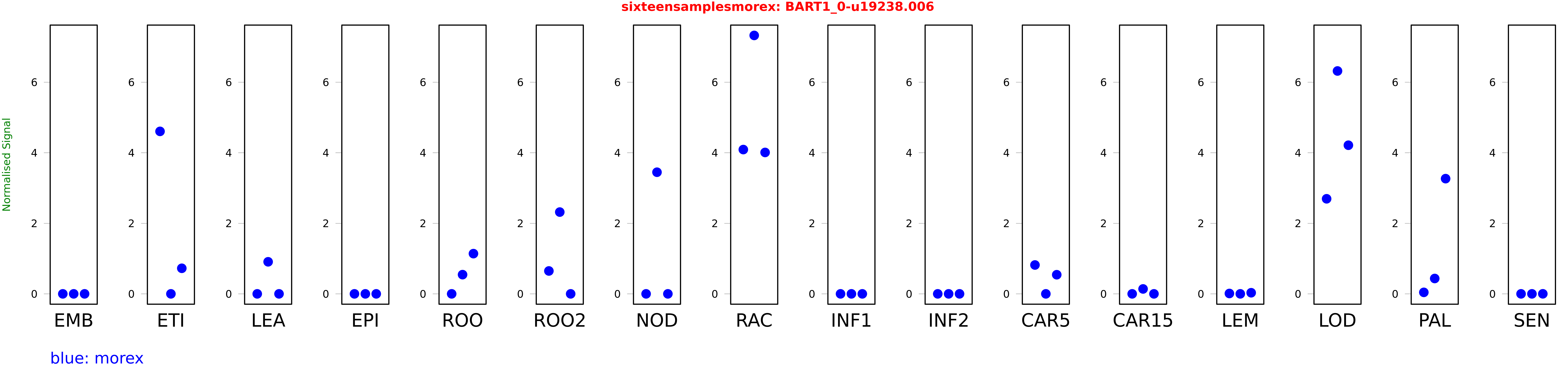
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0 | 0.000000000701924 | 0 |
| morex | ETI | 4.60693 | 0 | 0.724901 |
| morex | LEA | 0.0000000198674 | 0.908832 | 0 |
| morex | EPI | 0 | 0 | 0 |
| morex | ROO | 0 | 0.544745 | 1.14043 |
| morex | ROO2 | 0.649236 | 2.31872 | 0 |
| morex | NOD | 0 | 3.44873 | 0 |
| morex | RAC | 4.09106 | 7.32741 | 4.00946 |
| morex | INF1 | 0 | 0 | 0.00000000368589 |
| morex | INF2 | 0 | 0 | 0 |
| morex | CAR5 | 0.818317 | 0.0000168145 | 0.542221 |
| morex | CAR15 | 0 | 0.139454 | 0.000000444091 |
| morex | LEM | 0.0104765 | 0.000164554 | 0.0314658 |
| morex | LOD | 2.69546 | 6.31946 | 4.21405 |
| morex | PAL | 0.0423594 | 0.435644 | 3.26664 |
| morex | SEN | 0 | 0.0000000785697 | 0.00121985 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os01g11040.1 | +1 | 0.0 | 1131 | 742/1202 (62%) | protein|SCAR-like protein 2, putative, expressed |
| TAIR PP10 | AT2G38440 @ ARAPORT AT2G38440.1 @ TAIR |
+2 | 6e-48 | 159 | 86/161 (53%) | Symbols: ITB1, SCAR2, DIS3, WAVE4, ATSCAR2 | SCAR homolog 2 | chr2:16095550-16100851 FORWARD LENGTH=1399 |
| BRACH PP3 | Bradi2g06610.2.p | +1 | 0.0 | 1273 | 784/1211 (65%) | pacid=32774244 transcript=Bradi2g06610.2 locus=Bradi2g06610 ID=Bradi2g06610.2.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (4946 bp)
>BART1_0-u19238.006 4946 150831_barley_pseudomolecules GTGAAAACCAAGAAGGGCCAAGCACGGCCGGAAATGGCAAAGCACCCCCGGCCCCCGCGC GACCCAGCACAGCACAGCACAGCACAGCACAGCAGCCTCCCACCACAACATAGCGGCGGC ACCGGGTATCGAGCATCCAAATCCAAGCCGAGCCCGAGAGCTGAAGCATAATCACGCGGC GCACACCACTCACCAGTCGCCAGGCCGCGCCACTTGCCCACCCCCCTCGTGCCCGCAGCG CTGCGTGCGCTTCCCTTTCCCACCGTCCCTTTCGCCCCGCACCCTGTCGGAGACCGACGC TCCTCCCCTCCCCTCCCCGTCCCCGTCCCCGCCACGCGCCTCGGCGTCGCCTCCACTCCT CCCCCCCTCCCCGCCACTCGCCTCGGCCGCCCCCTTCCGTGCGGAGGCGGCAACCTGTGA CGCCCCAAAGTCGGCGCACCTGCTCCCACTGGGAAAGTAGCCGTTGCGGCGTCGCATTTA TAGCGACCGCCAGGCGTCACGATGGCCGCTGCTCGGCGGTGAGGAGGAGCCCGCTCTCGC GCGCTCGGTGGTTGGTGGAGCGGAGATGCCGCTGGTCAGGTTCGAGGTGCGGAATGAGGT CGGGCTGGGCGACCCCGGGCTGTACGGCGCCGCGGGGGCAGGGAAGCGAGGCGGCGCCGG CGCCGCCGCCGCAGGGGAGGCCGAGCCCAAGGCGCTGCTCGAGGGCGTCGCCGTCGCGGG CCTCGTCGGGATCCTGCGCCAGCTCGGAGATCTCGCCGAATTTGCGGCAGATATTTTTCA TGACCTACACGAGCAAGTTATAACTACATCGTCTAGGGGGCGTAAGGTGTTGACTCGAGT ACAGAACATTGAAGCAGCACTTCCATCTCTTGAAAAAGCTGTGAAGAATCAGAAGAGCCA TATACATTTTGCTTATGTACCAGGTTCTGACTGGCACACACAACTTCAAAATGAGCAAAA TCATCTCCTATCCAGTGATCTGCCTCGGTTTATGATGGATTCCTATGAAGAATGTCGAGA CCCACCACGACTTTACCTTCTCGACAAGTAAGGATGCTTGTTTTTTTTTTTCAAGTGATT TTGGGTCTCTAATGACTTATTTTCAGATTTGATAATGCTGGTGCTGGGGCTTGTTTGAAG AGATATTCTGATCCATCTTACTTCAAGAAATCATGGGATGTGATGAGAGCAGACAAGACT GCACATCTCCAAAAAGAAAGGAGATCTCATAAAATCAAGAGGAAAGGATCACGCCTAAAA GAACCATATCATGGACAAGCCACATCCAGGCATAGGAATGGTGAATTGCAGCGAGCACTC ACTGCTGTTCAACTCACCAGCAGCAGGCAGTGTGCATCTCCCAGCATGGATGGCCAGAGT TTTTCGGAGCATAGATCTACATCTGATGCAAGATCCAACCCTGACAATATAAGCAGATCT TCTTCATTTAGTTCAAAGGCACGACTGAGTTTTGCAGAACAAGCTTCAGATACAAAGCCT TCTGTAATTCCTCATGAGAATGGTCATGCCAAGCTGTCAAATATTGATATGCACAAGCTT AATGATGCCTCTTCCCCCGTCTTGCTTAGCGAGACCAGGGAAGATGATCTGGGTGATGAT TTGAAGCAAGGTTCCCTGTCAGATGAGATGAATGCTAGGTCACCTTCTGTTGAATGGCAT GAAAAGGCTGCAATTGTCATGACTACAAGTTCTGTCTACTGTGATGATGTTGTCATGGAT AGGGCTGAAAATGCAGAAACTAAACATATTAAGCCCGTGCAGCGAGAAGTTGATTATAGG GAGATGCAGACCTTGGAGCAGCAAGGGGCCTTACTTCAGAAGGCAAAGTTGTTATTACTG TCATCAGGCTTAAACCACCGTGATGAAGTCCCCTGTGAAACAGACAACTACATGGATGCA CTTAACACACTTGAATCTGAGACAGAGACTGAAGTTGACTCTCAAACTAAAAAACAAGGG AAGCCAGTACATTCCTTCAATGCTCATGCACCTCAAATAGAGTTGACAGATAATGCCGTT TCACAGCTTCCTGATTCTTCTCCTGTTGAGTTTCCTGATACAAGTCCAAATTCCAGAATG TTACATACTTTTGAAAGAAGTGCCGATTTCCCCAGTTTGTCAAGTGCAGATGCTCCGGAT ATTTCACAGCATTCATTGTCAGGTTATACAGATATACATCCTAATGAATGGTCATCTGTT ACCACCATCCCTGAGAATAATGCAAATAATGCTGCTGGAGACCCCACTGAAATTTCAGAA CCGGCACTGCAAGCATATACACCTACACCACCCAATCAAAGGTGCCCTCGTGCAAATGAG ATTCCTGAGAGTAAGGCAGAAGTTATCCCCAGAGATTCTCCTGTGATCTCAAAACCAGAG CTGTCAACCTATGCAGTTTTCCCTCCCAATAAAGAGTCTATTGTCAGTCAGATCCCTGAG AATAGTGTAGAAGGTGTGTCAGAAGATGATACCAATGAAATCACTAGTTGTCTTGTACCG GAACCTTCCATTTCCTTTGCTCCAACTAGTGAGGCGTCTCCTGCAAAAATCTTACCTGGT TATAACACTGGTAACTCTGTAATTTCGGAAAGGAGTCCTCAGGATTATCCTGGAGAAAAT CATCAGGAATTTGGTGGCTGTGGCATGGCTGAAGTGTCTAATTCACAGACCATGCCTTTG AACGAGTCATCAGAGAATGGATCTGCAACCCAACACCTTCCCACACATGCTCCTACTAGT TCCCAACACCTTCCCACACATGCTCCGACTAGTTCTGTAGAATTATCTTCTGTTAAGCTC TGGACAAATGCTGGTCTCTTCGGACTTGAACCGTCTAAACCTCCAGTTGGCATATGTCCA GGTTCTGCAAACCAAAGCTACCCAGAGAATAATCAATCAGCCACAAGAACACCTGATGCA GTTTATAGCCAAACAAACCGGCCCTCAGATTCTTCCACATATTTCGAGCACAGGGAGCAC AGCAATTTGAATGGCAAGCAAGCTTCAATAAGTGAGCTCCTAGAATCTGAAGGTAATGCT GAAATTGGTGCTGAAACATACTCTGGCACTGACTTGGTTGGAAGGAATAACTTGCCCGTG GTGTCTGCATCAAGCTTCTCGAGCATTGCACAAAGATTCCTTGCCAATACACTTCAGAGA AGAACTTCTCCCAAATATAATGATCTTCCTATGTCATCTGGGAGAGTGAATACTGACACA AATGGGTTTGATGAAGCTACTGTAAATTCTACTCTTGCACCAAAGGAAACAGTATTCGAG GCATCTCAGTTTGAGAAGAAAACAGATTATGGCGTGGATGGACTGCCCAAGTCCTCTAGT TGCCATTACTCTGAGAAATCATCTCCGCCACTAGAGCATATGAAAATATCCTTTCACCCC ATGAGTGCATCTGAAATGTCGAAATTGAACCTAGATTTTTCTGATGGCAATCTTCATGAA AATGTTGATGATATGATGTTACCAACATTTCAGTTACTTCCAGGGTCTTCTGTCCCACAG CCTGGCAGTGGTTCTGAATCAGAAGATGACACCTTTGGTAGATCTTATAGTTATTCTTCA TATGATGATCTAAGTCCGCGTTTATATTCAAACTCAGAGTTGTGGGATCAAGAAGATGGA ATTGGATTCGAGGATAATGAGCTGTATAATGATTCAAATCAAATAGGATCCTCCACAACG CCTGTCTCTAGCTATATGGGATTCGAGCAGATGAACTTATCTGGTGAGAAGTCCACTATT TCACTTTCAGATATTGGAGATAATAACGGACTCGGCTCGTTAGAATCCCATCCTGCTGGA GAACTTCCGAACTTTGACACTTTGATGTCAACAAGTAATCATCAAAATAGAGATGCGCTC ATTCCACAAAATCCAATCAATTTATCACCAGTTGTAGATCAGATGCCGCCGCCACCTCCT CTTCCCCCAATGCAGTGGAGGACAATGAGACAAACAACTTCCCTAGAAGAAGAAAGAGAC TCTACGGCTAAAGACATGCTTGAGAATGCCTCAAGTCTACCACCTGTACACACTCCTGTG CAGGAACAACATCTTCCACCGTGTGCACCACCATTTCCACAAGGAAATGTTAAGGAAGTG AACCATCAGAAAGTTGATGTGATCAAAGAAACGAGTAATCTTCCTAATATATTTGAGATC AAATCAAGCTTGCTTCAGCAAATCAGGGATAAGCCAGATCTGCACAAGCTGAATGGACAT GAAAAGTCCAAAGCAGTATTCAGTGATGTTAACAGTTTGGATGAAAGGGGGGAGTTGCTT CAGCAAATCAGGAGCAAGACATTCAACTTAAGACGAACAAATGGATCTAAGACAAACACC TCATCGCAGTCAACCGAGTCCACTGCCAATTCCAGCGTTGTAGCAATTTTGGAGAAGGCG AATGCAATCCGACAGGCTGTGGCCAGTGACGAGGGGGGTGATGATGATAACTGGAGCGAT ATCAACTGAGAATGCATATGCCAAACCTTTTCTGTATATGCAACTAGTACAAGATGAAGA GTGAAAATGTGAATAACACCTTTTTCTTCATTGTAATTAGATATTGCAGCTCTGTTTTTT AACCTTTTTTTAGTTCATGGTAGTGTATCTTAGATCCCTATGGTTGTAGAGATGTTAAAT CATTTCAACATCATGTAAATTTATCTTATTATTTATTAATGGTAGTGAGATGGACTAATA AAATAAGTATGTTAAGCTATCTTTTGTAGCATTGTACTAAAAATTGTCTCTTTGCATTGT TAATAAGATGGGGGCATACTGATAAAAAAGTTATGCTGTCATTTGCAAGCATGCTATTTT TTTTGGCGAAGGCAAAATAATTCTTAAGAAATCTCGCTGCATTACCTCCATGCTTTGTAT TTGAGTAGCAAAATCTAAATCAGGAG
Protein Sequence (1108 aa)
>BART1_0-u19238.006 1108 150831_barley_pseudomolecules MRADKTAHLQKERRSHKIKRKGSRLKEPYHGQATSRHRNGELQRALTAVQLTSSRQCASP SMDGQSFSEHRSTSDARSNPDNISRSSSFSSKARLSFAEQASDTKPSVIPHENGHAKLSN IDMHKLNDASSPVLLSETREDDLGDDLKQGSLSDEMNARSPSVEWHEKAAIVMTTSSVYC DDVVMDRAENAETKHIKPVQREVDYREMQTLEQQGALLQKAKLLLLSSGLNHRDEVPCET DNYMDALNTLESETETEVDSQTKKQGKPVHSFNAHAPQIELTDNAVSQLPDSSPVEFPDT SPNSRMLHTFERSADFPSLSSADAPDISQHSLSGYTDIHPNEWSSVTTIPENNANNAAGD PTEISEPALQAYTPTPPNQRCPRANEIPESKAEVIPRDSPVISKPELSTYAVFPPNKESI VSQIPENSVEGVSEDDTNEITSCLVPEPSISFAPTSEASPAKILPGYNTGNSVISERSPQ DYPGENHQEFGGCGMAEVSNSQTMPLNESSENGSATQHLPTHAPTSSQHLPTHAPTSSVE LSSVKLWTNAGLFGLEPSKPPVGICPGSANQSYPENNQSATRTPDAVYSQTNRPSDSSTY FEHREHSNLNGKQASISELLESEGNAEIGAETYSGTDLVGRNNLPVVSASSFSSIAQRFL ANTLQRRTSPKYNDLPMSSGRVNTDTNGFDEATVNSTLAPKETVFEASQFEKKTDYGVDG LPKSSSCHYSEKSSPPLEHMKISFHPMSASEMSKLNLDFSDGNLHENVDDMMLPTFQLLP GSSVPQPGSGSESEDDTFGRSYSYSSYDDLSPRLYSNSELWDQEDGIGFEDNELYNDSNQ IGSSTTPVSSYMGFEQMNLSGEKSTISLSDIGDNNGLGSLESHPAGELPNFDTLMSTSNH QNRDALIPQNPINLSPVVDQMPPPPPLPPMQWRTMRQTTSLEEERDSTAKDMLENASSLP PVHTPVQEQHLPPCAPPFPQGNVKEVNHQKVDVIKETSNLPNIFEIKSSLLQQIRDKPDL HKLNGHEKSKAVFSDVNSLDERGELLQQIRSKTFNLRRTNGSKTNTSSQSTESTANSSVV AILEKANAIRQAVASDEGGDDDNWSDIN
