- Transcript BART1_0-u19238.003
- Transcript IDBART1_0-u19238.003
- Gene IDBART1_0-u19238
Exon Structure of BART1_0-u19238.003
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr3H | 1 | 133980544 | 133980009 | - |
| chr3H | 2 | 133980745 | 133980629 | - |
| chr3H | 3 | 133980989 | 133980885 | - |
| chr3H | 4 | 133981597 | 133981505 | - |
| chr3H | 5 | 133984485 | 133981692 | - |
| chr3H | 6 | 133984718 | 133984615 | - |
| chr3H | 7 | 133985153 | 133985055 | - |
| chr3H | 8 | 133985370 | 133985247 | - |
| chr3H | 9 | 133985932 | 133985769 | - |
| chr3H | 10 | 133988750 | 133986152 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials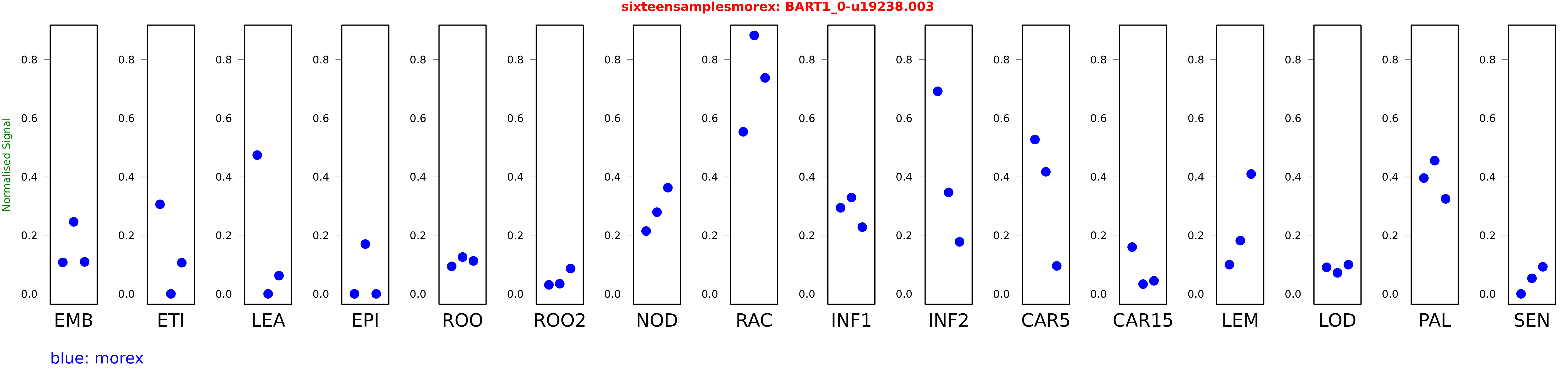
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0.107499 | 0.245829 | 0.109168 |
| morex | ETI | 0.30569 | 0 | 0.10626 |
| morex | LEA | 0.473644 | 0 | 0.0622498 |
| morex | EPI | 0 | 0.169936 | 0 |
| morex | ROO | 0.0940761 | 0.125639 | 0.112549 |
| morex | ROO2 | 0.0307846 | 0.0345488 | 0.0862676 |
| morex | NOD | 0.214445 | 0.278874 | 0.362398 |
| morex | RAC | 0.553287 | 0.88229 | 0.737357 |
| morex | INF1 | 0.293757 | 0.328894 | 0.227832 |
| morex | INF2 | 0.691443 | 0.346395 | 0.17761 |
| morex | CAR5 | 0.526738 | 0.416734 | 0.0954498 |
| morex | CAR15 | 0.159706 | 0.0332394 | 0.0445394 |
| morex | LEM | 0.0994704 | 0.181827 | 0.409108 |
| morex | LOD | 0.090757 | 0.0717243 | 0.0994067 |
| morex | PAL | 0.395314 | 0.454749 | 0.324296 |
| morex | SEN | 0 | 0.0529724 | 0.0924268 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os01g11040.1 | +2 | 0.0 | 1112 | 732/1187 (62%) | protein|SCAR-like protein 2, putative, expressed |
| TAIR PP10 | None | - | - | - | - | - |
| BRACH PP3 | Bradi2g06610.2.p | +2 | 0.0 | 1253 | 773/1196 (65%) | pacid=32774244 transcript=Bradi2g06610.2 locus=Bradi2g06610 ID=Bradi2g06610.2.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (6735 bp)
>BART1_0-u19238.003 6735 150831_barley_pseudomolecules CCGGCCCCCGCGCGACCCAGCACAGCACAGCACAGCACAGCACAGCAGCCTCCCACCACA ACATAGCGGCGGCACCGGGTATCGAGCATCCAAATCCAAGCCGAGCCCGAGAGCTGAAGC ATAATCACGCGGCGCACACCACTCACCAGTCGCCAGGCCGCGCCACTTGCCCACCCCCCT CGTGCCCGCAGCGCTGCGTGCGCTTCCCTTTCCCACCGTCCCTTTCGCCCCGCACCCTGT CGGAGACCGACGCTCCTCCCCTCCCCTCCCCGTCCCCGTCCCCGCCACGCGCCTCGGCGT CGCCTCCACTCCTCCCCCCCTCCCCGCCACTCGCCTCGGCCGCCCCCTTCCGTGCGGAGG CGGCAACCTGTGACGCCCCAAAGTCGGCGCACCTGCTCCCACTGGGAAAGTAGCCGTTGC GGCGTCGCATTTATAGCGACCGCCAGGCGTCACGATGGCCGCTGCTCGGCGGTGAGGAGG AGCCCGCTCTCGCGCGCTCGGTGGTTGGTGGAGCGGAGATGCCGCTGGTCAGGTTCGAGG TGCGGAATGAGGTCGGGCTGGGCGACCCCGGGCTGTACGGCGCCGCGGGGGCAGGGAAGC GAGGCGGCGCCGGCGCCGCCGCCGCAGGGGAGGCCGAGCCCAAGGCGCTGCTCGAGGGCG TCGCCGTCGCGGGCCTCGTCGGGATCCTGCGCCAGCTCGGAGATCTCGCCGAGTAAGCCG CCCCCTCCCCTCATCTCCGTCGCTCCCTGCCTCGCGGTCTGATTTCAGTTTTTCCTGCGT GATTCGATTCGGCGAGTTGTGTGTGTGTGTGTGTGCTTTTTTTCCAGCCTTCCAGTGTGA GATCTGCTGCTCTCCGTGTGTGGTGATTTTGGCGGGCAACTACGGTTGGGGAAGTGTTTG GTCGCTGGTTTACTGGAGTTGAGGTGTTTGTGCTTGAGATTGGGGGTGATTGGAAGGTAG TGGGATCTGGTGTTGAGGTGGTTGCATTTGATAACGCGAGATTGAGTGCGGGGTTCTAGG AACTATAGGAATTATGGCAGGGTTTCCGCTCGCCGATTATGGACAAAGCAGCAGTGCTGC GTGGCTGGAACTGGATAGTATTTCGTTTTGGGCACTGGTGGCTTTCGAGCTCATTTCCGA GAGAGCATTAGGATTGAGTCGGGTAGAACCAAATTGTGTCTTCAGAGCGTTATACGAGCT AAACTATACAATCAAGTGGGTTTTTACCGCGTTTGCTAGGTTTTTGGAGGTATGCTTTGC TGAATTGGTAGGAACTATCGGTGTACTCTAGGTATGCCATGTTTGGGTAAGATCCCATGT TAGGATGGATTTATTTTATCACACTAGCTATATTATATAGATCGTGATTCTGAAAAGTTC CTGTTCTAGACGGCTTAGTGGGTGACTGGTTGAATTCGTTGAATATGTGGGAATAAAGGA GTGTTTGCTAATTCTTGTGCGACTGTTTCCATGGCTTCTAGCAATTTCTACTGGTTGTGT GATCAAAAATGTCACCACTCTTGGGAAGTCTTTTATTTATGGTAAACGTTTCCACGAACG CTTCGGCCGGTCTCGTCCACTCGCTCGCGGCTATAGCAAATCAAATCGGGAGGGCCCATC ACGTTATCTCTTTCCTCATCCCCGCATCCTCCTCCTTCTTTTTTCCCAACCTCCTGGTTG CCATGGGAAATCCTACCCCGGCGATCTCCTCCAACGAGCCCCCGTCTTCTTCTCCGGCGA GCACCCTCGAAGCCTAGCATCGAACCAGTGTCCTCCCCTCGATTTCCCCTCTTTGAACTG CTCCATGGTGACTGGAGCTGCAACGGGCATGGCCGGCGAAGTTGCATCCATGGCCGAAGG CGTAATTTACTACATGCTTCAACCGGCATCGAAATTTGGTACAACCGGCACCGCATTTTG CTATCACCGGCGTCGCATTTTGCTACATCGCACATGGCATTGCGATTTTGCTACATCCAT CACGGCTGAGCTGCGACCCGTGACGGCGGAACCATGCTCGCGTTTGTTGCAACTGTTGAC AGAAAAAGCTTCAACGGTATATGAAAGATGTTTCAACCGGCAATAGAGAAAGCTTCAACC AGGGGCTCCGTCCATCCATTTAACTCCACTCCGACGAGGCGAACCTCCACCCTGAAAGCT GCAACCAGCTGATCGAAAAGCTGCAGCCGTCGTCTAATTTTGTTACTACTGGCAACGGAT TTTGCTACATCTGTCATGGCGGAGCTGCGGCCTACTACGGTGATGGCTTTTTTTTGTTGC AACCGTCCCTCGGATTTGCTACAACCGGAGAGGTATTTTGCTACATGCATCATGGCGGAG CTGGGATCGATGGCGAGGGCCGCGACCGACGGCGAGGGGGCTGCGACCAGTGATGACTGG CGGTGAGAGAAGATGAGGAGGCGAGGCGTGGGGTGCAGCGTGTGCGGGGGCGGCACCCTC CTTTTTCTCTCTTTCTTCCGTGTGGGAGGTTGAAGATGGGATGCGGAAGAGGAGGGGGGA GATCTGACGGCTACACGCGTGCAAATCGGACTGATCCGGGCTGACCGGCCGAAACATTGG GCCGGCGCGCCGACGCAGAATTTGCGGCAGATATTTTTCATGACCTACACGAGCAAGTTA TAACTACATCGTCTAGGGGGCGTAAGGTGTTGACTCGAGTACAGAACATTGAAGCAGCAC TTCCATCTCTTGAAAAAGCTGTGAAGAATCAGAAGAGCCATATACATTTTGCTTATGTAC CAGGTTCTGACTGGCACACACAACTTCAAAATGAGCAAAATCATCTCCTATCCAGTGATC TGCCTCGGTTTATGATGGATTCCTATGAAGAATGTCGAGACCCACCACGACTTTACCTTC TCGACAAAGATATTCTGATCCATCTTACTTCAAGAAATCATGGGATGTGATGAGAGCAGA CAAGACTGCACATCTCCAAAAAGAAAGGAGATCTCATAAAATCAAGAGGAAAGGATCACG CCTAAAAGAACCATATCATGGACAAGCCACATCCAGGCATAGGAATGGTGAATTGCAGCG AGCACTCACTGCTGTTCAACTCACCAGCAGGCAGTGTGCATCTCCCAGCATGGATGGCCA GAGTTTTTCGGAGCATAGATCTACATCTGATGCAAGATCCAACCCTGACAATATAAGCAG ATCTTCTTCATTTAGTTCAAAGGCACGACTGAGTTTTGCAGAACAAGCTTCAGATACAAA GCCTTCTGTAATTCCTCATGAGAATGGTCATGCCAAGCTGTCAAATATTGATATGCACAA GCTTAATGATGCCTCTTCCCCCGTCTTGCTTAGCGAGACCAGGGAAGATGATCTGGGTGA TGATTTGAAGCAAGGTTCCCTGTCAGATGAGATGAATGCTAGGTCACCTTCTGTTGAATG GCATGAAAAGGCTGCAATTGTCATGACTACAAGTTCTGTCTACTGTGATGATGTTGTCAT GGATAGGGCTGAAAATGCAGAAACTAAACATATTAAGCCCGTGCAGCGAGAAGTTGATTA TAGGGAGATGCAGACCTTGGAGCAGCAAGGGGCCTTACTTCAGAAGGCAAAGTTGTTATT ACTGTCATCAGGCTTAAACCACCGTGATGAAGTCCCCTGTGAAACAGACAACTACATGGA TGCACTTAACACACTTGAATCTGAGACAGAGACTGAAGTTGACTCTCAAACTAAAAAACA AGGGAAGCCAGTACATTCCTTCAATGCTCATGCACCTCAAATAGAGTTGACAGATAATGC CGTTTCACAGCTTCCTGATTCTTCTCCTGTTGAGTTTCCTGATACAAGTCCAAATTCCAG AATGTTACATACTTTTGAAAGAAGTGCCGATTTCCCCAGTTTGTCAAGTGCAGATGCTCC GGATATTTCACAGCATTCATTGTCAGGTTATACAGATATACATCCTAATGAATGGTCATC TGTTACCACCATCCCTGAGAATAATGCAAATAATGCTGCTGGAGACCCCACTGAAATTTC AGAACCGGCACTGCAAGCATATACACCTACACCACCCAATCAAAGGTGCCCTCGTGCAAA TGAGATTCCTGAGAGTAAGGCAGAAGTTATCCCCAGAGATTCTCCTGTGATCTCAAAACC AGAGCTGTCAACCTATGCAGTTTTCCCTCCCAATAAAGAGTCTATTGTCAGTCAGATCCC TGAGAATAGTGTAGAAGGTGTGTCAGAAGATGATACCAATGAAATCACTAGTTGTCTTGT ACCGGAACCTTCCATTTCCTTTGCTCCAACTAGTGAGGCGTCTCCTGCAAAAATCTTACC TGGTTATAACACTGGTAACTCTGTAATTTCGGAAAGGAGTCCTCAGGATTATCCTGGAGA AAATCATCAGGAATTTGGTGGCTGTGGCATGGCTGAAGTGTCTAATTCACAGACCATGCC TTTGAACGAGTCATCAGAGAATGGATCTGCAACCCAACACCTTCCCACACATGCTCCTAC TAGTTCCCAACACCTTCCCACACATGCTCCGACTAGTTCTGTAGAATTATCTTCTGTTAA GCTCTGGACAAATGCTGGTCTCTTCGGACTTGAACCGTCTAAACCTCCAGTTGGCATATG TCCAGGTTCTGCAAACCAAAGCTACCCAGAGAATAATCAATCAGCCACAAGAACACCTGA TGCAGTTTATAGCCAAACAAACCGGCCCTCAGATTCTTCCACATATTTCGAGCACAGGGA GCACAGCAATTTGAATGGCAAGCAAGCTTCAATAAGTGAGCTCCTAGAATCTGAAGGTAA TGCTGAAATTGGTGCTGAAACATACTCTGGCACTGACTTGGTTGGAAGGAATAACTTGCC CGTGGTGTCTGCATCAAGCTTCTCGAGCATTGCACAAAGATTCCTTGCCAATACACTTCA GAGAAGAACTTCTCCCAAATATAATGATCTTCCTATGTCATCTGGGAGAGTGAATACTGA CACAAATGGGTTTGATGAAGCTACTGTAAATTCTACTCTTGCACCAAAGGAAACAGTATT CGAGGCATCTCAGTTTGAGAAGAAAACAGATTATGGCGTGGATGGACTGCCCAAGTCCTC TAGTTGCCATTACTCTGAGAAATCATCTCCGCCACTAGAGCATATGAAAATATCCTTTCA CCCCATGAGTGCATCTGAAATGTCGAAATTGAACCTAGATTTTTCTGATGGCAATCTTCA TGAAAATGTTGATGATATGATGTTACCAACATTTCAGTTACTTCCAGGGTCTTCTGTCCC ACAGCCTGGCAGTGGTTCTGAATCAGAAGATGACACCTTTGGTAGATCTTATAGTTATTC TTCATATGATGATCTAAGTCCGCGTTTATATTCAAACTCAGAGTTGTGGGATCAAGAAGA TGGAATTGGATTCGAGGATAATGAGCTGTATAATGATTCAAATCAAATAGGATCCTCCAC AACGCCTGTCTCTAGCTATATGGGATTCGAGCAGATGAACTTATCTGGTGAGAAGTCCAC TATTTCACTTTCAGATATTGGAGATAATAACGGACTCGGCTCGTTAGAATCCCATCCTGC TGGAGAACTTCCGAACTTTGACACTTTGATGTCAACAAGTAATCATCAAAATAGAGATGC GCTCATTCCACAAAATCCAATCAATTTATCACCAGTTGTAGATCAGATGCCGCCGCCACC TCCTCTTCCCCCAATGCAGTGGAGGACAATGAGACAAACAACTTCCCTAGAAGAAGAAAG AGACTCTACGGCTAAAGACATGCTTGAGAATGCCTCAAGTCTACCACCTGTACACACTCC TGTGCAGGAACAACATCTTCCACCGTGTGCACCACCATTTCCACAAGGAAATGTTAAGGA AGTGAACCATCAGAAAGTTGATGTGATCAAAGAAACGAGTAATCTTCCTAATATATTTGA GATCAAATCAAGCTTGCTTCAGCAAATCAGGGATAAGCCAGATCTGCACAAGCTGAATGG ACATGAAAAGTCCAAAGCAGTATTCAGTGATGTTAACAGTTTGGATGAAAGGGGGGAGTT GCTTCAGCAAATCAGGAGCAAGACATTCAACTTAAGACGAACAAATGGATCTAAGACAAA CACCTCATCGCAGTCAACCGAGTCCACTGCCAATTCCAGCGTTGTAGCAATTTTGGAGAA GGCGAATGCAATCCGACAGGCTGTGGCCAGTGACGAGGGGGGTGATGATGATAACTGGAG CGATATCAACTGAGAATGCATATGCCAAACCTTTTCTGTATATGCAACTAGTACAAGATG AAGAGTGAAAATGTGAATAACACCTTTTTCTTCATTGTAATTAGATATTGCAGCTCTGTT TTTTAACCTTTTTTTAGTTCATGGTAGTGTATCTTAGATCCCTATGGTTGTAGAGATGTT AAATCATTTCAACATCATGTAAATTTATCTTATTATTTATTAATGGTAGTGAGATGGACT AATAAAATAAGTATGTTAAGCTATCTTTTGTAGCATTGTACTAAAAATTGTCTCTTTGCA TTGTTAATAAGATGGGGGCATACTGATAAAAAAGTTATGCTGTCATTTGCAAGCATGCTA TTTTTTTTGGCGAAGGCAAAATAATTCTTAAGAAATCTCGCTGCATTACCTCCATGCTTT GTATTTGAGTAGCAAAATCTAAATCAGGAGATGCAATTGATTTTTCTTCAAGTTAGTCAA TATATCATGTAAGAG
Protein Sequence (1133 aa)
>BART1_0-u19238.003 1133 150831_barley_pseudomolecules MSRPTTTLPSRQRYSDPSYFKKSWDVMRADKTAHLQKERRSHKIKRKGSRLKEPYHGQAT SRHRNGELQRALTAVQLTSRQCASPSMDGQSFSEHRSTSDARSNPDNISRSSSFSSKARL SFAEQASDTKPSVIPHENGHAKLSNIDMHKLNDASSPVLLSETREDDLGDDLKQGSLSDE MNARSPSVEWHEKAAIVMTTSSVYCDDVVMDRAENAETKHIKPVQREVDYREMQTLEQQG ALLQKAKLLLLSSGLNHRDEVPCETDNYMDALNTLESETETEVDSQTKKQGKPVHSFNAH APQIELTDNAVSQLPDSSPVEFPDTSPNSRMLHTFERSADFPSLSSADAPDISQHSLSGY TDIHPNEWSSVTTIPENNANNAAGDPTEISEPALQAYTPTPPNQRCPRANEIPESKAEVI PRDSPVISKPELSTYAVFPPNKESIVSQIPENSVEGVSEDDTNEITSCLVPEPSISFAPT SEASPAKILPGYNTGNSVISERSPQDYPGENHQEFGGCGMAEVSNSQTMPLNESSENGSA TQHLPTHAPTSSQHLPTHAPTSSVELSSVKLWTNAGLFGLEPSKPPVGICPGSANQSYPE NNQSATRTPDAVYSQTNRPSDSSTYFEHREHSNLNGKQASISELLESEGNAEIGAETYSG TDLVGRNNLPVVSASSFSSIAQRFLANTLQRRTSPKYNDLPMSSGRVNTDTNGFDEATVN STLAPKETVFEASQFEKKTDYGVDGLPKSSSCHYSEKSSPPLEHMKISFHPMSASEMSKL NLDFSDGNLHENVDDMMLPTFQLLPGSSVPQPGSGSESEDDTFGRSYSYSSYDDLSPRLY SNSELWDQEDGIGFEDNELYNDSNQIGSSTTPVSSYMGFEQMNLSGEKSTISLSDIGDNN GLGSLESHPAGELPNFDTLMSTSNHQNRDALIPQNPINLSPVVDQMPPPPPLPPMQWRTM RQTTSLEEERDSTAKDMLENASSLPPVHTPVQEQHLPPCAPPFPQGNVKEVNHQKVDVIK ETSNLPNIFEIKSSLLQQIRDKPDLHKLNGHEKSKAVFSDVNSLDERGELLQQIRSKTFN LRRTNGSKTNTSSQSTESTANSSVVAILEKANAIRQAVASDEGGDDDNWSDIN
