- Transcript BART1_0-u19238.002
- Transcript IDBART1_0-u19238.002
- Gene IDBART1_0-u19238
Exon Structure of BART1_0-u19238.002
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr3H | 1 | 133980544 | 133980009 | - |
| chr3H | 2 | 133980745 | 133980629 | - |
| chr3H | 3 | 133980989 | 133980885 | - |
| chr3H | 4 | 133981597 | 133981505 | - |
| chr3H | 5 | 133984485 | 133981692 | - |
| chr3H | 6 | 133985153 | 133985055 | - |
| chr3H | 7 | 133985370 | 133985247 | - |
| chr3H | 8 | 133985932 | 133985769 | - |
| chr3H | 9 | 133988780 | 133988039 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials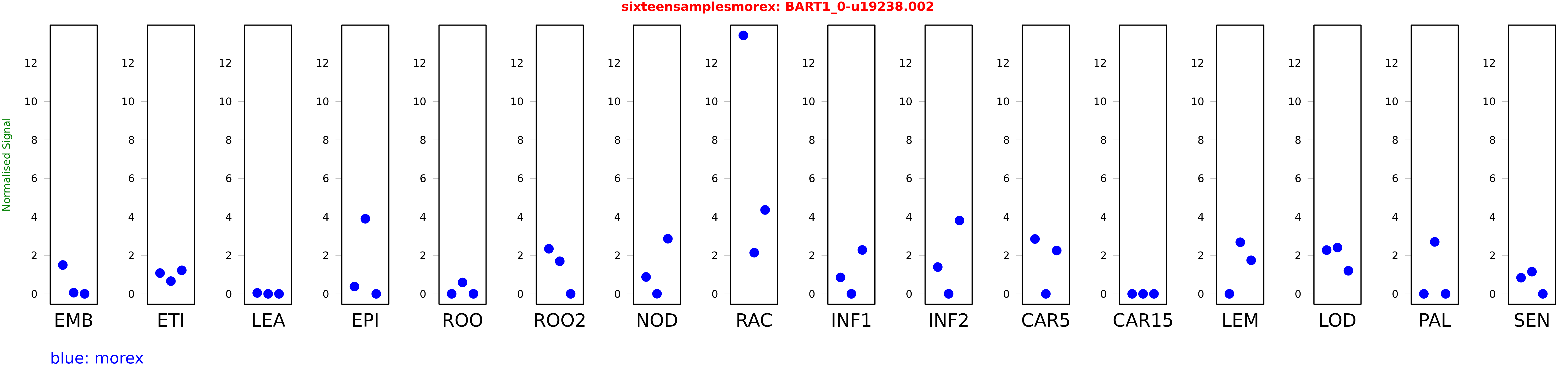
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 1.49779 | 0.0588155 | 0.00000010966 |
| morex | ETI | 1.07874 | 0.66072 | 1.22249 |
| morex | LEA | 0.0485496 | 0.0000000132978 | 0 |
| morex | EPI | 0.376607 | 3.89807 | 0.00000000200111 |
| morex | ROO | 0 | 0.593949 | 0 |
| morex | ROO2 | 2.34148 | 1.69409 | 0.0000000260648 |
| morex | NOD | 0.877167 | 0.00612102 | 2.86213 |
| morex | RAC | 13.4222 | 2.13648 | 4.35834 |
| morex | INF1 | 0.855749 | 0.0000000800483 | 2.27953 |
| morex | INF2 | 1.39139 | 0.00000330325 | 3.80621 |
| morex | CAR5 | 2.84924 | 0 | 2.25072 |
| morex | CAR15 | 0.0000854914 | 0.0000012924 | 0.0000000313852 |
| morex | LEM | 0.000000182784 | 2.6812 | 1.74312 |
| morex | LOD | 2.27217 | 2.40135 | 1.19841 |
| morex | PAL | 0.0000859162 | 2.69936 | 0.0000000559748 |
| morex | SEN | 0.838386 | 1.15172 | 0.0000345919 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os01g11040.1 | +3 | 0.0 | 1226 | 830/1364 (61%) | protein|SCAR-like protein 2, putative, expressed |
| TAIR PP10 | AT4G18600 @ ARAPORT AT4G18600.1 @ TAIR |
+3 | 4e-40 | 162 | 82/161 (51%) | Symbols: WAVE5, ATSCAR-LIKE, SCARL | SCAR family protein | chr4:10239947-10247372 REVERSE LENGTH=2028 |
| BRACH PP3 | Bradi2g06610.1.p | +3 | 0.0 | 1347 | 860/1367 (63%) | pacid=32774243 transcript=Bradi2g06610.1 locus=Bradi2g06610 ID=Bradi2g06610.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (4774 bp)
>BART1_0-u19238.002 4774 150831_barley_pseudomolecules CCAAGCACGGCCGGAAATGGCAAAGCACCCCCGGCCCCCGCGCGACCCAGCACAGCACAG CACAGCACAGCACAGCAGCCTCCCACCACAACATAGCGGCGGCACCGGGTATCGAGCATC CAAATCCAAGCCGAGCCCGAGAGCTGAAGCATAATCACGCGGCGCACACCACTCACCAGT CGCCAGGCCGCGCCACTTGCCCACCCCCCTCGTGCCCGCAGCGCTGCGTGCGCTTCCCTT TCCCACCGTCCCTTTCGCCCCGCACCCTGTCGGAGACCGACGCTCCTCCCCTCCCCTCCC CGTCCCCGTCCCCGCCACGCGCCTCGGCGTCGCCTCCACTCCTCCCCCCCTCCCCGCCAC TCGCCTCGGCCGCCCCCTTCCGTGCGGAGGCGGCAACCTGTGACGCCCCAAAGTCGGCGC ACCTGCTCCCACTGGGAAAGTAGCCGTTGCGGCGTCGCATTTATAGCGACCGCCAGGCGT CACGATGGCCGCTGCTCGGCGGTGAGGAGGAGCCCGCTCTCGCGCGCTCGGTGGTTGGTG GAGCGGAGATGCCGCTGGTCAGGTTCGAGGTGCGGAATGAGGTCGGGCTGGGCGACCCCG GGCTGTACGGCGCCGCGGGGGCAGGGAAGCGAGGCGGCGCCGGCGCCGCCGCCGCAGGGG AGGCCGAGCCCAAGGCGCTGCTCGAGGGCGTCGCCGTCGCGGGCCTCGTCGGGATCCTGC GCCAGCTCGGAGATCTCGCCGAATTTGCGGCAGATATTTTTCATGACCTACACGAGCAAG TTATAACTACATCGTCTAGGGGGCGTAAGGTGTTGACTCGAGTACAGAACATTGAAGCAG CACTTCCATCTCTTGAAAAAGCTGTGAAGAATCAGAAGAGCCATATACATTTTGCTTATG TACCAGGTTCTGACTGGCACACACAACTTCAAAATGAGCAAAATCATCTCCTATCCAGTG ATCTGCCTCGGTTTATGATGGATTCCTATGAAGAATGTCGAGACCCACCACGACTTTACC TTCTCGACAAAGATATTCTGATCCATCTTACTTCAAGAAATCATGGGATGTGATGAGAGC AGACAAGACTGCACATCTCCAAAAAGAAAGGAGATCTCATAAAATCAAGGCAGTGTGCAT CTCCCAGCATGGATGGCCAGAGTTTTTCGGAGCATAGATCTACATCTGATGCAAGATCCA ACCCTGACAATATAAGCAGATCTTCTTCATTTAGTTCAAAGGCACGACTGAGTTTTGCAG AACAAGCTTCAGATACAAAGCCTTCTGTAATTCCTCATGAGAATGGTCATGCCAAGCTGT CAAATATTGATATGCACAAGCTTAATGATGCCTCTTCCCCCGTCTTGCTTAGCGAGACCA GGGAAGATGATCTGGGTGATGATTTGAAGCAAGGTTCCCTGTCAGATGAGATGAATGCTA GGTCACCTTCTGTTGAATGGCATGAAAAGGCTGCAATTGTCATGACTACAAGTTCTGTCT ACTGTGATGATGTTGTCATGGATAGGGCTGAAAATGCAGAAACTAAACATATTAAGCCCG TGCAGCGAGAAGTTGATTATAGGGAGATGCAGACCTTGGAGCAGCAAGGGGCCTTACTTC AGAAGGCAAAGTTGTTATTACTGTCATCAGGCTTAAACCACCGTGATGAAGTCCCCTGTG AAACAGACAACTACATGGATGCACTTAACACACTTGAATCTGAGACAGAGACTGAAGTTG ACTCTCAAACTAAAAAACAAGGGAAGCCAGTACATTCCTTCAATGCTCATGCACCTCAAA TAGAGTTGACAGATAATGCCGTTTCACAGCTTCCTGATTCTTCTCCTGTTGAGTTTCCTG ATACAAGTCCAAATTCCAGAATGTTACATACTTTTGAAAGAAGTGCCGATTTCCCCAGTT TGTCAAGTGCAGATGCTCCGGATATTTCACAGCATTCATTGTCAGGTTATACAGATATAC ATCCTAATGAATGGTCATCTGTTACCACCATCCCTGAGAATAATGCAAATAATGCTGCTG GAGACCCCACTGAAATTTCAGAACCGGCACTGCAAGCATATACACCTACACCACCCAATC AAAGGTGCCCTCGTGCAAATGAGATTCCTGAGAGTAAGGCAGAAGTTATCCCCAGAGATT CTCCTGTGATCTCAAAACCAGAGCTGTCAACCTATGCAGTTTTCCCTCCCAATAAAGAGT CTATTGTCAGTCAGATCCCTGAGAATAGTGTAGAAGGTGTGTCAGAAGATGATACCAATG AAATCACTAGTTGTCTTGTACCGGAACCTTCCATTTCCTTTGCTCCAACTAGTGAGGCGT CTCCTGCAAAAATCTTACCTGGTTATAACACTGGTAACTCTGTAATTTCGGAAAGGAGTC CTCAGGATTATCCTGGAGAAAATCATCAGGAATTTGGTGGCTGTGGCATGGCTGAAGTGT CTAATTCACAGACCATGCCTTTGAACGAGTCATCAGAGAATGGATCTGCAACCCAACACC TTCCCACACATGCTCCTACTAGTTCCCAACACCTTCCCACACATGCTCCGACTAGTTCTG TAGAATTATCTTCTGTTAAGCTCTGGACAAATGCTGGTCTCTTCGGACTTGAACCGTCTA AACCTCCAGTTGGCATATGTCCAGGTTCTGCAAACCAAAGCTACCCAGAGAATAATCAAT CAGCCACAAGAACACCTGATGCAGTTTATAGCCAAACAAACCGGCCCTCAGATTCTTCCA CATATTTCGAGCACAGGGAGCACAGCAATTTGAATGGCAAGCAAGCTTCAATAAGTGAGC TCCTAGAATCTGAAGGTAATGCTGAAATTGGTGCTGAAACATACTCTGGCACTGACTTGG TTGGAAGGAATAACTTGCCCGTGGTGTCTGCATCAAGCTTCTCGAGCATTGCACAAAGAT TCCTTGCCAATACACTTCAGAGAAGAACTTCTCCCAAATATAATGATCTTCCTATGTCAT CTGGGAGAGTGAATACTGACACAAATGGGTTTGATGAAGCTACTGTAAATTCTACTCTTG CACCAAAGGAAACAGTATTCGAGGCATCTCAGTTTGAGAAGAAAACAGATTATGGCGTGG ATGGACTGCCCAAGTCCTCTAGTTGCCATTACTCTGAGAAATCATCTCCGCCACTAGAGC ATATGAAAATATCCTTTCACCCCATGAGTGCATCTGAAATGTCGAAATTGAACCTAGATT TTTCTGATGGCAATCTTCATGAAAATGTTGATGATATGATGTTACCAACATTTCAGTTAC TTCCAGGGTCTTCTGTCCCACAGCCTGGCAGTGGTTCTGAATCAGAAGATGACACCTTTG GTAGATCTTATAGTTATTCTTCATATGATGATCTAAGTCCGCGTTTATATTCAAACTCAG AGTTGTGGGATCAAGAAGATGGAATTGGATTCGAGGATAATGAGCTGTATAATGATTCAA ATCAAATAGGATCCTCCACAACGCCTGTCTCTAGCTATATGGGATTCGAGCAGATGAACT TATCTGGTGAGAAGTCCACTATTTCACTTTCAGATATTGGAGATAATAACGGACTCGGCT CGTTAGAATCCCATCCTGCTGGAGAACTTCCGAACTTTGACACTTTGATGTCAACAAGTA ATCATCAAAATAGAGATGCGCTCATTCCACAAAATCCAATCAATTTATCACCAGTTGTAG ATCAGATGCCGCCGCCACCTCCTCTTCCCCCAATGCAGTGGAGGACAATGAGACAAACAA CTTCCCTAGAAGAAGAAAGAGACTCTACGGCTAAAGACATGCTTGAGAATGCCTCAAGTC TACCACCTGTACACACTCCTGTGCAGGAACAACATCTTCCACCGTGTGCACCACCATTTC CACAAGGAAATGTTAAGGAAGTGAACCATCAGAAAGTTGATGTGATCAAAGAAACGAGTA ATCTTCCTAATATATTTGAGATCAAATCAAGCTTGCTTCAGCAAATCAGGGATAAGCCAG ATCTGCACAAGCTGAATGGACATGAAAAGTCCAAAGCAGTATTCAGTGATGTTAACAGTT TGGATGAAAGGGGGGAGTTGCTTCAGCAAATCAGGAGCAAGACATTCAACTTAAGACGAA CAAATGGATCTAAGACAAACACCTCATCGCAGTCAACCGAGTCCACTGCCAATTCCAGCG TTGTAGCAATTTTGGAGAAGGCGAATGCAATCCGACAGGCTGTGGCCAGTGACGAGGGGG GTGATGATGATAACTGGAGCGATATCAACTGAGAATGCATATGCCAAACCTTTTCTGTAT ATGCAACTAGTACAAGATGAAGAGTGAAAATGTGAATAACACCTTTTTCTTCATTGTAAT TAGATATTGCAGCTCTGTTTTTTAACCTTTTTTTAGTTCATGGTAGTGTATCTTAGATCC CTATGGTTGTAGAGATGTTAAATCATTTCAACATCATGTAAATTTATCTTATTATTTATT AATGGTAGTGAGATGGACTAATAAAATAAGTATGTTAAGCTATCTTTTGTAGCATTGTAC TAAAAATTGTCTCTTTGCATTGTTAATAAGATGGGGGCATACTGATAAAAAAGTTATGCT GTCATTTGCAAGCATGCTATTTTTTTTGGCGAAGGCAAAATAATTCTTAAGAAATCTCGC TGCATTACCTCCATGCTTTGTATTTGAGTAGCAAAATCTAAATCAGGAGATGCAATTGAT TTTTCTTCAAGTTAGTCAATATATCATGTAAGAG
Protein Sequence (1047 aa)
>BART1_0-u19238.002 1047 150831_barley_pseudomolecules MDGQSFSEHRSTSDARSNPDNISRSSSFSSKARLSFAEQASDTKPSVIPHENGHAKLSNI DMHKLNDASSPVLLSETREDDLGDDLKQGSLSDEMNARSPSVEWHEKAAIVMTTSSVYCD DVVMDRAENAETKHIKPVQREVDYREMQTLEQQGALLQKAKLLLLSSGLNHRDEVPCETD NYMDALNTLESETETEVDSQTKKQGKPVHSFNAHAPQIELTDNAVSQLPDSSPVEFPDTS PNSRMLHTFERSADFPSLSSADAPDISQHSLSGYTDIHPNEWSSVTTIPENNANNAAGDP TEISEPALQAYTPTPPNQRCPRANEIPESKAEVIPRDSPVISKPELSTYAVFPPNKESIV SQIPENSVEGVSEDDTNEITSCLVPEPSISFAPTSEASPAKILPGYNTGNSVISERSPQD YPGENHQEFGGCGMAEVSNSQTMPLNESSENGSATQHLPTHAPTSSQHLPTHAPTSSVEL SSVKLWTNAGLFGLEPSKPPVGICPGSANQSYPENNQSATRTPDAVYSQTNRPSDSSTYF EHREHSNLNGKQASISELLESEGNAEIGAETYSGTDLVGRNNLPVVSASSFSSIAQRFLA NTLQRRTSPKYNDLPMSSGRVNTDTNGFDEATVNSTLAPKETVFEASQFEKKTDYGVDGL PKSSSCHYSEKSSPPLEHMKISFHPMSASEMSKLNLDFSDGNLHENVDDMMLPTFQLLPG SSVPQPGSGSESEDDTFGRSYSYSSYDDLSPRLYSNSELWDQEDGIGFEDNELYNDSNQI GSSTTPVSSYMGFEQMNLSGEKSTISLSDIGDNNGLGSLESHPAGELPNFDTLMSTSNHQ NRDALIPQNPINLSPVVDQMPPPPPLPPMQWRTMRQTTSLEEERDSTAKDMLENASSLPP VHTPVQEQHLPPCAPPFPQGNVKEVNHQKVDVIKETSNLPNIFEIKSSLLQQIRDKPDLH KLNGHEKSKAVFSDVNSLDERGELLQQIRSKTFNLRRTNGSKTNTSSQSTESTANSSVVA ILEKANAIRQAVASDEGGDDDNWSDIN
