- Transcript BART1_0-u16393.011
- Transcript IDBART1_0-u16393.011
- Gene IDBART1_0-u16393
Exon Structure of BART1_0-u16393.011
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr2H | 1 | 752632673 | 752632188 | - |
| chr2H | 2 | 752633100 | 752632995 | - |
| chr2H | 3 | 752633394 | 752633314 | - |
| chr2H | 4 | 752633633 | 752633497 | - |
| chr2H | 5 | 752634002 | 752633909 | - |
| chr2H | 6 | 752634160 | 752634090 | - |
| chr2H | 7 | 752634845 | 752634684 | - |
| chr2H | 8 | 752635015 | 752634959 | - |
| chr2H | 9 | 752635164 | 752635102 | - |
| chr2H | 10 | 752635679 | 752635387 | - |
| chr2H | 11 | 752636131 | 752635766 | - |
| chr2H | 12 | 752636358 | 752636210 | - |
| chr2H | 13 | 752636704 | 752636676 | - |
| chr2H | 14 | 752637251 | 752636874 | - |
| chr2H | 15 | 752637512 | 752637389 | - |
| chr2H | 16 | 752637750 | 752637597 | - |
| chr2H | 17 | 752637930 | 752637852 | - |
| chr2H | 18 | 752638100 | 752638041 | - |
| chr2H | 19 | 752638282 | 752638181 | - |
| chr2H | 20 | 752638779 | 752638674 | - |
| chr2H | 21 | 752639126 | 752638900 | - |
| chr2H | 22 | 752643671 | 752643212 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials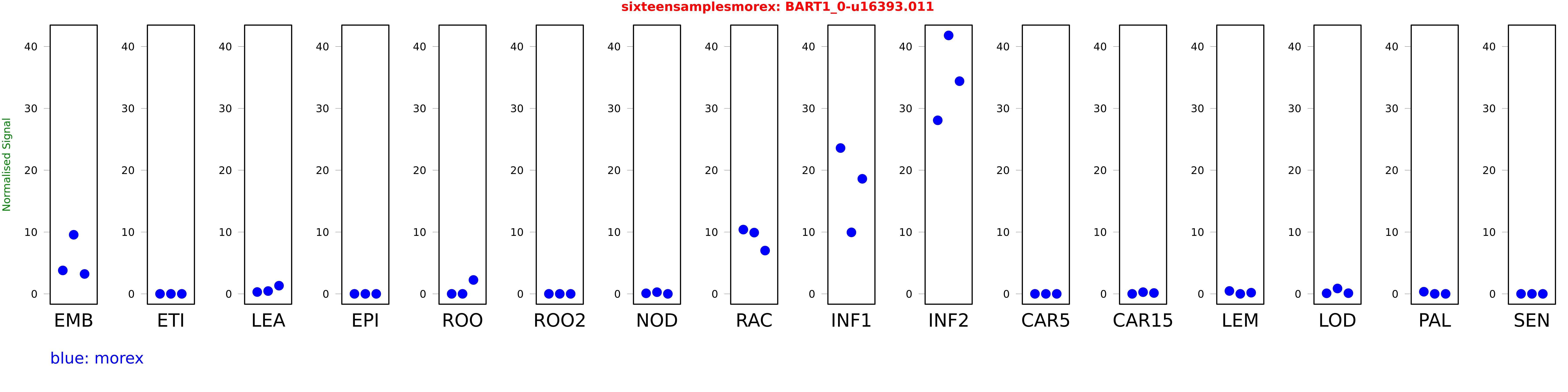
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 3.78952 | 9.5574 | 3.21652 |
| morex | ETI | 0 | 0 | 0 |
| morex | LEA | 0.297157 | 0.454599 | 1.3228 |
| morex | EPI | 0 | 0 | 0 |
| morex | ROO | 0.00000000147729 | 0.00147553 | 2.24776 |
| morex | ROO2 | 0.0028477 | 0 | 0 |
| morex | NOD | 0.0868492 | 0.273233 | 0 |
| morex | RAC | 10.395 | 9.90623 | 7.00212 |
| morex | INF1 | 23.5853 | 9.94442 | 18.6085 |
| morex | INF2 | 28.0708 | 41.799 | 34.4007 |
| morex | CAR5 | 0 | 0 | 0 |
| morex | CAR15 | 0 | 0.275231 | 0.137147 |
| morex | LEM | 0.468662 | 0 | 0.189312 |
| morex | LOD | 0.08826 | 0.871005 | 0.108674 |
| morex | PAL | 0.343327 | 0 | 0 |
| morex | SEN | 0 | 0 | 0.0000555644 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os07g29220.1 | +2 | 8e-162 | 492 | 229/312 (73%) | protein|Cyclopropane-fatty-acyl-phospholipid synthase, putative, expressed |
| TAIR PP10 | AT3G23510 @ ARAPORT AT3G23510.1 @ TAIR |
+2 | 5e-131 | 423 | 197/309 (64%) | Symbols: | Cyclopropane-fatty-acyl-phospholipid synthase | chr3:8428071-8433159 FORWARD LENGTH=867 |
| BRACH PP3 | Bradi5g02490.1.p | +2 | 4e-153 | 483 | 227/324 (70%) | pacid=32809829 transcript=Bradi5g02490.1 locus=Bradi5g02490 ID=Bradi5g02490.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (3784 bp)
>BART1_0-u16393.011 3784 150831_barley_pseudomolecules TACAAAGTCAAATAAACACAACGGCCAGCATTCAGAGGTAGGTGGTTGGTCGATCCAAGG ACGCACTGCGGATGGTGGCTATTTTCAGTAGTATTGGAATGCATTCCGTCCCTGAGATGT AGCACTACTGCACTAGTAGTATATTCTGCAAGTTGTGACGCGCGCTACAGCCTACCGCAG CCGCTCCGGCTTAAATAGAGGGGCAGGCCTGAGAGCGCAGAGAGCAGAGAGCAGAGAGCA GAGCGGAGCATAGACACCGAGAAGAATGAGGGTGGCGGTGGTTGGCGGCGGGGTGAGCGG GCTGGCGGCGGCGCATGAGCTCGCCGCCAGCGGGGGAGACGGCGTGCGAGTGACCCTGTA CGAGGAGGAGGCCAGCCTCGGCGGGCACGCCAGGACGGTGGCCGTCGATGACGGCGCCGG CTGCGTCAAGCTCGACCTCGGCTTCATGGTCTTCAACCAGGTAACATATCCGCAAATGAT GGAATGGTTAGAAGGGCTCGGGGTTGAGATGGAGAGGTCAGACATGTCTTTCTCTGTGAG CACACAATCTAAAGGAGGCGGTGGAGGATGTGAGTGGGGAAATGGTAACGGTATATCGAG CCTCTTGGCGCAAAAAACCAACATACTGAAACCCAGTTTTTGGCACATGGTCTGTGAGAT ACTCAAGTTCAAGAACAATGCCCTGACGTACCTGGAGGACCATGAACGTAACCCTGATCT GGATCGTAATGAGACACTGGGGCAGTTCATTCAGTCGCATGGATATTCCCTATCGTTTCA AGAGGCTTATCTTATTCCAGTCTGCACAGGCACGTGGTCATGTTCATCACAAGATGTTTT GAGCCTGTCTGCTTTCTTGGTGCTGTCATTTTGCCGTAACCATGGCCTTCTTCAGCTGTT TCGTCACTCCCAGTTGCCCACTGTCAAGCCTAGTTCGCAGTCCTATGTAAACAAGGTAAA AGGAGAATTGGAGAGCATTGGCTGTCGAATTAAAACCAGTTGTCGGGTCAAATCTGTTTC AAGTCTTGATGGAGCAGCTGGCTACAGAGTCTTGGAGAATGATGGTTCAGTGGAAACATA CGACAGTGTCATCTTTGGGGTCCATGCATCCAATGCTCTCAAATTATTAGGAGCTGAAGC ATCACATCACGAACGAAGAATTCTAGGTGCCTGTCAGTATGTCCAAAGAGATATATACCT CCACTGCGATCAAAATTTGATGCCCCGAAAATCAGCAGCATGGAGTGCGTGGAATTTCCT TGGGACAACTAGTAGGGGTTTTTCTATTACTTACTGGCTAAACCAAATTCAGGACTAGGC ATGCCTCCTCGGCCAAAAGGAAGAAACCAAATGAAGAAAAAAATATTTTCATCATGTTAG AACTCAGAACACCCTGTAGAGTAGGTTCACACAATAACATCAAGACGGAAGCTTCAGCCT AACCCCTCCAGCAGCATGAATGTATTACTTCATTATTAACTGTGGCATATTTGTTCTTTT TTGCTCTTCTAGAAATTTGAATCGGGCAGGCCTTTCCTGGTGACACTCAATCCCCCTTGT GTTCCGGATCATGTACTACTTAAATGGAATACAAGCCTTCCTGTTCCGTCTGTGGCCGCG GCGAAGGCTTATCTTCAGCTTGATCAGATCCAGGGAAAGAGGGGAATATGGTTCTGTGGG GCATATCAAGGTCATGGCTTCCATGAAGACGGATTAAAGTCTGGGAAAGCAGCGGCTCAA GGTTTGCTTGGAAAGAAATGTGAGCTTCTGCTGAACCCAAAGAATATGATTCCATCATGG ATTGAGGCTGGGGCACGCCTTCTAGTTGCAAGATTTTTCAACCAATACATATCCATTGGT AACTTGATATTGGTCGAAGAAGGAGGTAGTGTGTTCAGCTTTGGTAAAGCTTGCGACAAA TGCCGTGTAAAATCTGTCATCCAAGTTCATGACCCCTTGTTCTATTGGAAGGTTATCTTC TCTTTTCTTATTCTGACTACACATGCATTTAAAAAAATTAAGAGTGGTGATGTGCCCATG CATGATTCATAATTTCCTATATACTCCACTGCATTTGCTTTGTACAACTTTTTCATTCGA ACCACATGTGTGTACCCTGGCTCTCTCATGGCTTTCAGGTTGCAATAGAAGGAGGCATGG GCTTGGCAGAATCCTATATTGACGGTTGTTTCTCTGTTCTTGACAAGAGAGAAGGCCTTC TAAATCTTATCCTGATTCTCATTGCTAACAGAGACGAGCGTAGGAACCGCCGCATTGCCA GAAAAGGGTTCTGAAACCTTCAATTGTCCACTAGATTCCTATGAATTGTTTGATAAAAAT GAAGTAGTACCTATGAGTATGCGTTTGTGATTAATCCTGTACATGCTGTTTTTCGGTTTG CAGGTTTTGGTTGTCACCCTTCCATGTAATAGCTCAATTGGCATATGCTAAACACTTTTT GCGCCAGGCAACAAGGAAGAACACTGCAACACAAACTCGTCGAAACATCTCTGAGCACTA TGATCTTAGTAACGACTTTTTTTCGCTTTTTCTGGATAAATCGATGACTTACTCCTGTGC AGTTTTCAAGGAGAACGAAAGCTTAGAAGCAGCCCAACAACGTAAACTTAGCCTTCTAAT CAACAAGGCTAAAGTCAAGAGGGAGCATCATGTTCTTGATATCGGTAGCGGCTGGGGAAG TTTAGCAATAGAAACAGTTAAGCAAACTGGCTGCAAATACACTGGAGTCACTTTATCGGC GGAGCAGCATAAATACGCAGAAAGAAAAGTAAGAGAAGCTGGCTTAGAGGACCGCATAAA TTTTCTGCTGTGCGACTACCGTAAAATACCACCTTTTAAGTATGACGCAATCATAAGCTG TGGTATGATTGAACATGTTGGCCATGAATACATGGACGAATTTTTTGCTTGCTGTGAGTC TTACTTGGCTGAAGATGGGATATTGGTCCTCCAGTTCATCTCAGTTCCAGAGGAACGGTA CGAGCAATACAGAAAAAGGCCAGACTTCATAAAAGAGTACATATTCCCTGGTGGTTGCCT TCCTTCTTTGGCCCGTATAATGTCTGCCATGACCGCGTCATCAAGGTTCTGCATAGAGCA TGTCGAGAATATTGGGCCCAATTACTACACAACTCTGATGCACTGGAGGGACAACTTCAT GGCCAACAGAGACCAAGTTTTGGCCCTGGGCTTTGATGAAAGGTTCCTCCGTATATGGGA GTACTACCTCATATTCTCTGCGGCTGGTTTCAAGTCACGAGCAATTGGAGATTACCAGGT TGTTTTCTCTCGTCCAGGCAATCGTCGTCTAGGCCAGCCTTGATGGCCGGAGACAACATA TCCTTCTCCGTTCAGCGGCCGAATGGTTACTTGTCAGTACCGTACGAGATTTTAATAAGT CAGTCCTGGAATGATTCGCTCATGCCGGAATCCTACTCAGGTGGTTTGGTGATCTATCTT GTTGTTTAACCATGCAATGGTTAGTTTTTCGTGGGAGTGTCTTGTTATCCATCAATTAAC TAGAATTTTGAGAATTGTATGTAAGGAACTGGTTTGTTTGCTTTAATATCTATGAGGAAG AGTTGATGTCCTTGCTTGTAGTTTCTCATGATCTGATCTATTCGGTGTTTGCCGTTTTCA ATGCTTGCTTGCGAAGAGAATCAATATATGTTCCTGAGTATCATGTTTCACCAATGACTG AGGAGGACTTTTGCTAACAGAAAAGATGAGATCTCTTTGGCTAGTTTTTGTTAAAGGGGA TGGC
Protein Sequence (350 aa)
>BART1_0-u16393.011 350 150831_barley_pseudomolecules MRVAVVGGGVSGLAAAHELAASGGDGVRVTLYEEEASLGGHARTVAVDDGAGCVKLDLGF MVFNQVTYPQMMEWLEGLGVEMERSDMSFSVSTQSKGGGGGCEWGNGNGISSLLAQKTNI LKPSFWHMVCEILKFKNNALTYLEDHERNPDLDRNETLGQFIQSHGYSLSFQEAYLIPVC TGTWSCSSQDVLSLSAFLVLSFCRNHGLLQLFRHSQLPTVKPSSQSYVNKVKGELESIGC RIKTSCRVKSVSSLDGAAGYRVLENDGSVETYDSVIFGVHASNALKLLGAEASHHERRIL GACQYVQRDIYLHCDQNLMPRKSAAWSAWNFLGTTSRGFSITYWLNQIQD
