- Transcript BART1_0-u03305.005
- Transcript IDBART1_0-u03305.005
- Gene IDBART1_0-u03305
Exon Structure of BART1_0-u03305.005
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr1H | 1 | 366830525 | 366830093 | - |
| chr1H | 2 | 366831202 | 366831022 | - |
| chr1H | 3 | 366831486 | 366831277 | - |
| chr1H | 4 | 366832334 | 366832002 | - |
| chr1H | 5 | 366833086 | 366832797 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials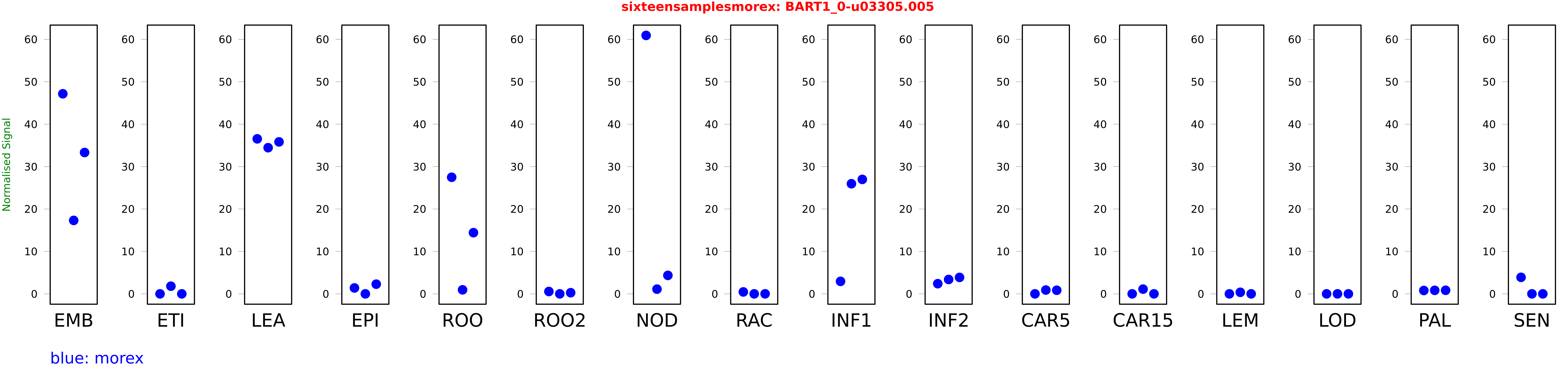
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 47.1663 | 17.3156 | 33.3266 |
| morex | ETI | 0 | 1.80761 | 0 |
| morex | LEA | 36.5428 | 34.4572 | 35.8241 |
| morex | EPI | 1.39452 | 0 | 2.28185 |
| morex | ROO | 27.4884 | 0.948689 | 14.4221 |
| morex | ROO2 | 0.558 | 0 | 0.269857 |
| morex | NOD | 60.9241 | 1.11169 | 4.35264 |
| morex | RAC | 0.455885 | 0 | 0.00958646 |
| morex | INF1 | 2.9459 | 25.9598 | 26.9849 |
| morex | INF2 | 2.39649 | 3.38852 | 3.87872 |
| morex | CAR5 | 0 | 0.90067 | 0.851173 |
| morex | CAR15 | 0 | 1.11371 | 0 |
| morex | LEM | 0 | 0.366249 | 0 |
| morex | LOD | 0 | 0 | 0 |
| morex | PAL | 0.795271 | 0.831625 | 0.841654 |
| morex | SEN | 3.89381 | 0 | 0 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os10g38950.1 | +3 | 1e-152 | 315 | 148/159 (93%) | protein|CGMC_MAPKCMGC_2_ERK.14 - CGMC includes CDA, MAPK, GSK3, and CLKC kinases, expressed |
| TAIR PP10 | AT4G01370 @ ARAPORT AT4G01370.1 @ TAIR |
+3 | 7e-132 | 290 | 130/158 (82%) | Symbols: ATMPK4, MPK4 | MAP kinase 4 | chr4:567219-568889 FORWARD LENGTH=376 |
| BRACH PP3 | Bradi3g32000.1.p | +3 | 7e-157 | 322 | 153/159 (96%) | pacid=32819051 transcript=Bradi3g32000.1 locus=Bradi3g32000 ID=Bradi3g32000.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1447 bp)
>BART1_0-u03305.005 1447 150831_barley_pseudomolecules TCTACGTTTCTAGAAGTGGTATGCCTGAGTGGGTTCTTATTCTTATCATTACAGTTTTAC TGTTGACGGGCTTTGTTGTCCACTCGTGCTCACACTGTCAGGCATTCCTATGATGGGTTT ATTCTGATAGTGGTTTCAAAATTGGATCACAGATTCTTGCCATGAAGGATTTAATACGCC CCCCAAGAAGAGATGATTTTAAGGATGTGTACATTGTTACTGAGTTGATGGACACTGATC TCCATCAGATCATTCGCTCAAATCAATCATTGACTGATGACCATTGCCAGTACTTCTTGT ATCAATTGCTTCGAGGGCTAAAATATGTGCACTCAGCAAATGTCTTGCACCGTGATCTGA AGCCAAGCAATTTGTTCCTAAACGCAAATTGCGATCTCAAGATTGCTGACTTTGGGCTTG CAAGGACCACTTCAGAAACTGATCTCATGACAGAGTATGTGGTCACTCGTTGGTACCGAG CGCCAGAGTTGCTGTTGAACTGTTCACAATATACTGCTGCTATTGATGTCTGGTCAGTTG GGTGCATACTTGGTGAAATTATTACTCGTCAACCCCTGTTTCCTGGAAGAGATTACATCC AACAGTTAAAGTTAATCACCGAGTCCTTTCCGGCATTCTGCTTCAAAAGCTGATAGGATC ACCAGATGATTCAAGCCTGGGATTTCTTCGGAGTGATAATGCACGAAGATACATGAAGCA ATTACCACAGTACCCAAGGCAGGACTTCCGCCTGCGCTTCCGCAACATGTCTGATGGTGC AGTTGATCTGTTAGAGAGGATGCTGGTGTTTGACCCAAGCAGACGCATTACTGTTGATGA GGCTCTGCATCACCCATACTTGGCTTCTCTTCATGACATCAATGAAGAACCCACTTGCCC GGCACCTTTCAGCTTTGATTTTGAGCAACCATCTTTTACAGAAGAACATATGAAAGAGCT CATCTGGAGGGAAACTTTAGCATTTAACCCTGATCCGCCCTACTAAGAGCCAAGACCAAA TTACCAGCTGAGGGCATTGAAGATCTACCTCTAGCTCTAGTGAAGCCGATATTTCCATGT CTTGTGCATCTATTTATTTTATGCTCGCTTATTGGGCGAATGGGCATGGATTATTTGTTG CTGTAACTATTTCTTTTGTGGCCTTTCTGAAGAAACGACATTTGTATGAGATGGCTTGTA TTATACCGGTTGATTGAATAAACTGGTCTTGTGTATCTAAAGCTTGTAATTTTGTGTACA CTTATCTAAACATCCCATGTATATCTGAGTTAACATATTCCCTTTTATTATCTCAAGCAT GTAATTTGGCATTTTGTGATCGGTCGTATCGAAGTTGCCGATATAATTTCGCTCTGAAAT TCAGTTCACTGGTAGGTGTATCCCACATTAAAAGTTTGTACATGCTATTCTATGTATTAC TTACTCC
Protein Sequence (163 aa)
>BART1_0-u03305.005 163 150831_barley_pseudomolecules MKDLIRPPRRDDFKDVYIVTELMDTDLHQIIRSNQSLTDDHCQYFLYQLLRGLKYVHSAN VLHRDLKPSNLFLNANCDLKIADFGLARTTSETDLMTEYVVTRWYRAPELLLNCSQYTAA IDVWSVGCILGEIITRQPLFPGRDYIQQLKLITESFPAFCFKS
