Probe CUST_9634_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9634_PI426222305 | JHI_St_60k_v1 | DMT400023732 | AAGTTCTGACAAAGCCTTTCTTCTGTTCCCGAGCTGAAGCTTGGGGTTTTGTGTTTTGTG |
All Microarray Probes Designed to Gene DMG400009180
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9630_PI426222305 | JHI_St_60k_v1 | DMT400023730 | TATTACTTGCTCGGGCAATAGAGTTACTGAAATATGAGGTTGCTGCTGCTGCTGCCGATA |
| CUST_9634_PI426222305 | JHI_St_60k_v1 | DMT400023732 | AAGTTCTGACAAAGCCTTTCTTCTGTTCCCGAGCTGAAGCTTGGGGTTTTGTGTTTTGTG |
Microarray Signals from CUST_9634_PI426222305
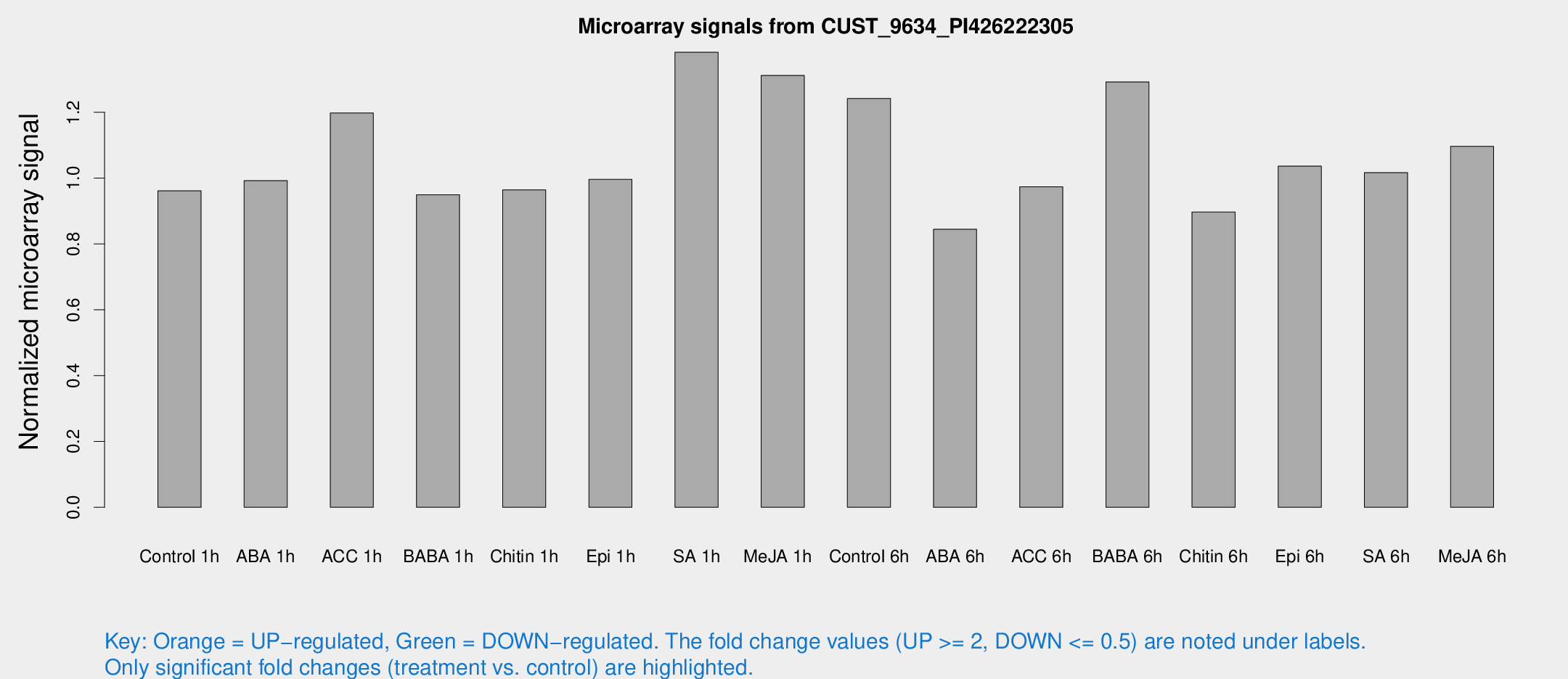
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.31535 | 3.16858 | 0.961547 | 0.500213 |
| ABA 1h | 5.678 | 3.00325 | 0.992268 | 0.530161 |
| ACC 1h | 8.53835 | 3.46457 | 1.1981 | 0.571858 |
| BABA 1h | 5.89504 | 3.34289 | 0.949271 | 0.53826 |
| Chitin 1h | 5.52596 | 3.15331 | 0.964509 | 0.540854 |
| Epi 1h | 5.55247 | 3.13211 | 0.996122 | 0.560369 |
| SA 1h | 10.2973 | 3.61324 | 1.3826 | 0.597846 |
| Me-JA 1h | 7.23685 | 3.2282 | 1.31228 | 0.644285 |
| Control 6h | 9.06572 | 3.35316 | 1.24222 | 0.580973 |
| ABA 6h | 5.78688 | 3.35911 | 0.84483 | 0.489984 |
| ACC 6h | 7.31528 | 3.9479 | 0.974039 | 0.514784 |
| BABA 6h | 9.92461 | 3.62184 | 1.29227 | 0.539782 |
| Chitin 6h | 6.14953 | 3.56279 | 0.897055 | 0.519474 |
| Epi 6h | 7.78088 | 3.73033 | 1.03681 | 0.522202 |
| SA 6h | 6.55121 | 3.38531 | 1.0169 | 0.537667 |
| Me-JA 6h | 7.09923 | 3.21052 | 1.09685 | 0.504475 |
Source Transcript PGSC0003DMT400023732 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G53730.2 | +2 | 0.0 | 775 | 429/691 (62%) | STRUBBELIG-receptor family 6 | chr1:20061771-20065475 FORWARD LENGTH=720 |
