Probe CUST_9630_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9630_PI426222305 | JHI_St_60k_v1 | DMT400023730 | TATTACTTGCTCGGGCAATAGAGTTACTGAAATATGAGGTTGCTGCTGCTGCTGCCGATA |
All Microarray Probes Designed to Gene DMG400009180
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9630_PI426222305 | JHI_St_60k_v1 | DMT400023730 | TATTACTTGCTCGGGCAATAGAGTTACTGAAATATGAGGTTGCTGCTGCTGCTGCCGATA |
| CUST_9634_PI426222305 | JHI_St_60k_v1 | DMT400023732 | AAGTTCTGACAAAGCCTTTCTTCTGTTCCCGAGCTGAAGCTTGGGGTTTTGTGTTTTGTG |
Microarray Signals from CUST_9630_PI426222305
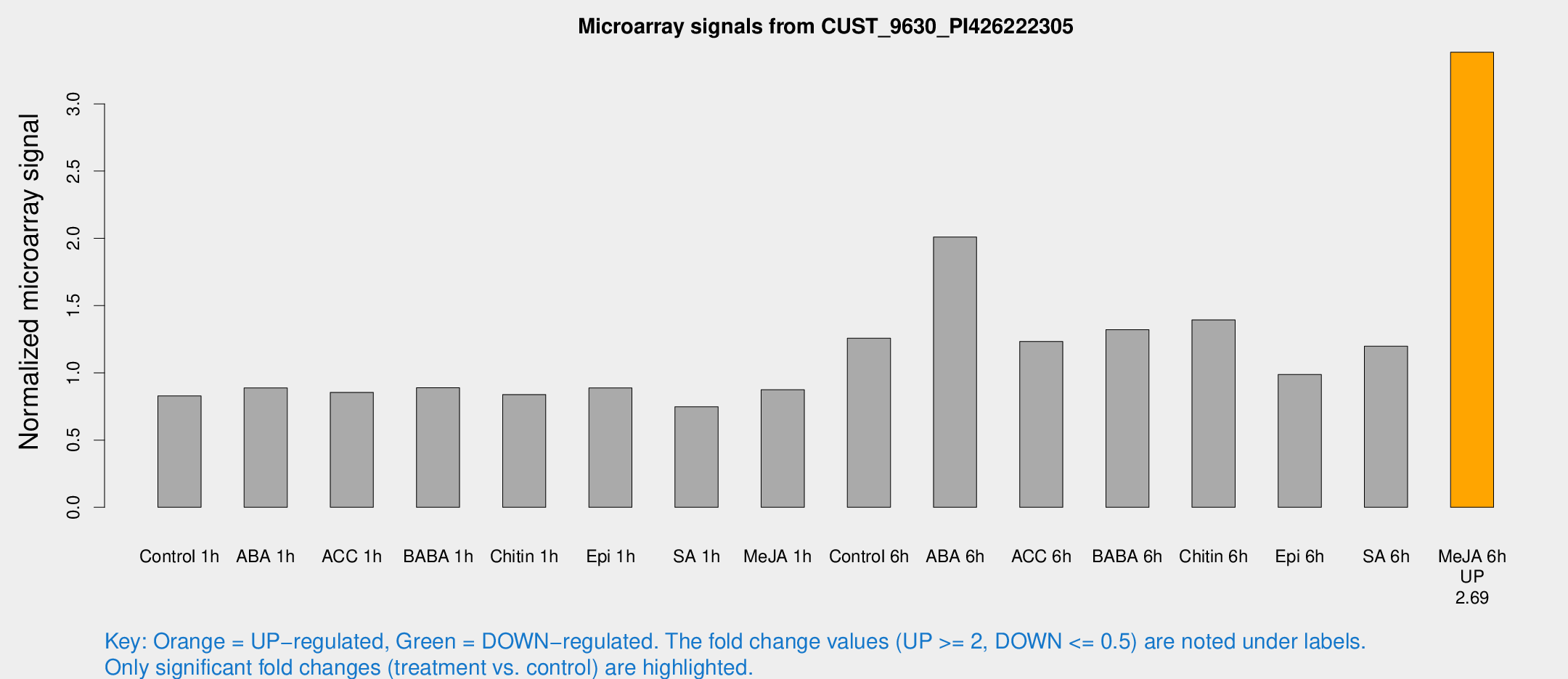
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 148.86 | 15.617 | 0.827857 | 0.0519734 |
| ABA 1h | 140.423 | 9.99218 | 0.887741 | 0.0879036 |
| ACC 1h | 157.003 | 11.7587 | 0.854517 | 0.0694904 |
| BABA 1h | 153.08 | 9.73751 | 0.888739 | 0.0561774 |
| Chitin 1h | 136.455 | 17.3726 | 0.838166 | 0.112602 |
| Epi 1h | 137.58 | 10.8607 | 0.887384 | 0.0637184 |
| SA 1h | 139.53 | 20.5689 | 0.747336 | 0.0889559 |
| Me-JA 1h | 131.4 | 24.4212 | 0.874497 | 0.130167 |
| Control 6h | 232.689 | 44.2502 | 1.25734 | 0.137685 |
| ABA 6h | 385.77 | 44.6846 | 2.00929 | 0.159074 |
| ACC 6h | 251.957 | 15.2992 | 1.23305 | 0.173859 |
| BABA 6h | 267.447 | 33.4008 | 1.32104 | 0.117073 |
| Chitin 6h | 269.712 | 40.4943 | 1.3931 | 0.161596 |
| Epi 6h | 202.596 | 29.3558 | 0.987805 | 0.0721954 |
| SA 6h | 212.027 | 15.5331 | 1.19781 | 0.222123 |
| Me-JA 6h | 604.501 | 55.6202 | 3.38375 | 0.236261 |
Source Transcript PGSC0003DMT400023730 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14350.1 | +1 | 0.0 | 632 | 379/687 (55%) | STRUBBELIG-receptor family 7 | chr3:4783115-4786999 REVERSE LENGTH=717 |
