Probe CUST_8944_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8944_PI426222305 | JHI_St_60k_v1 | DMT400033385 | GACCTCAACCTGATAAAGGCCTTGGCAAACTTAGAAAGTCCATTACTCTAATCCAAACTC |
All Microarray Probes Designed to Gene DMG400012826
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8944_PI426222305 | JHI_St_60k_v1 | DMT400033385 | GACCTCAACCTGATAAAGGCCTTGGCAAACTTAGAAAGTCCATTACTCTAATCCAAACTC |
| CUST_9030_PI426222305 | JHI_St_60k_v1 | DMT400033386 | TTATGGCTGGGCCTCGTCCTACTAATAGATTTCTAAAGGACCTCTCTATCAACACTTCTG |
Microarray Signals from CUST_8944_PI426222305
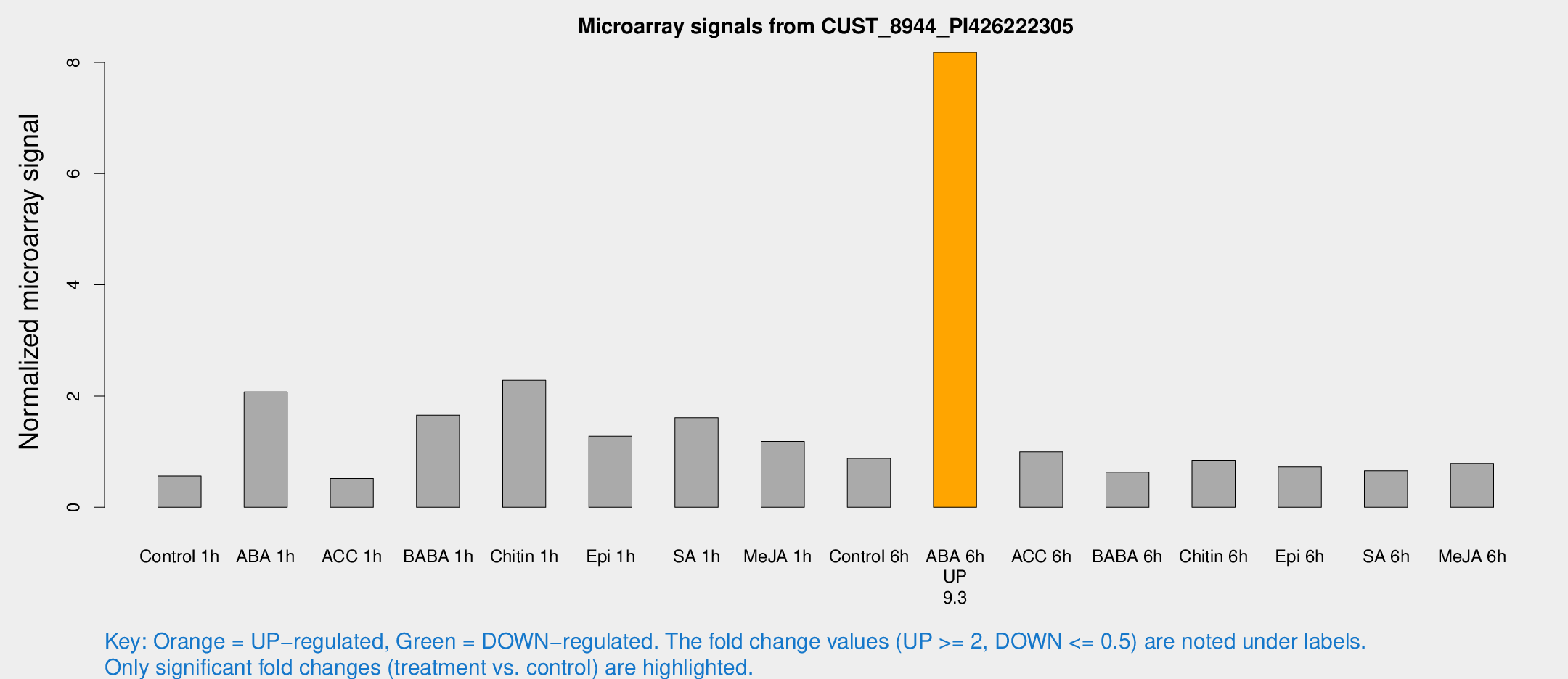
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 19.4865 | 3.60944 | 0.564828 | 0.105951 |
| ABA 1h | 64.6844 | 10.0423 | 2.07551 | 0.207939 |
| ACC 1h | 24.5244 | 12.0179 | 0.520585 | 0.402526 |
| BABA 1h | 60.4009 | 19.3566 | 1.65878 | 0.413192 |
| Chitin 1h | 72.416 | 11.3483 | 2.28235 | 0.410256 |
| Epi 1h | 41.2583 | 10.4531 | 1.27856 | 0.459029 |
| SA 1h | 65.5954 | 25.0389 | 1.61031 | 0.550098 |
| Me-JA 1h | 36.2864 | 9.93152 | 1.18339 | 0.502288 |
| Control 6h | 30.4743 | 4.00914 | 0.879374 | 0.117766 |
| ABA 6h | 309.396 | 53.9565 | 8.18112 | 1.26512 |
| ACC 6h | 39.5294 | 4.96024 | 0.997457 | 0.225193 |
| BABA 6h | 26.7578 | 7.71079 | 0.634828 | 0.183169 |
| Chitin 6h | 31.4435 | 4.34827 | 0.846202 | 0.120648 |
| Epi 6h | 28.4823 | 4.45086 | 0.726384 | 0.117663 |
| SA 6h | 29.5565 | 11.5681 | 0.66013 | 0.382347 |
| Me-JA 6h | 28.7773 | 7.17448 | 0.789694 | 0.137564 |
Source Transcript PGSC0003DMT400033385 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G33830.2 | +3 | 2e-32 | 117 | 60/123 (49%) | Dormancy/auxin associated family protein | chr2:14309768-14310286 REVERSE LENGTH=108 |
