Probe CUST_9030_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_9030_PI426222305 | JHI_St_60k_v1 | DMT400033386 | TTATGGCTGGGCCTCGTCCTACTAATAGATTTCTAAAGGACCTCTCTATCAACACTTCTG |
All Microarray Probes Designed to Gene DMG400012826
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8944_PI426222305 | JHI_St_60k_v1 | DMT400033385 | GACCTCAACCTGATAAAGGCCTTGGCAAACTTAGAAAGTCCATTACTCTAATCCAAACTC |
| CUST_9030_PI426222305 | JHI_St_60k_v1 | DMT400033386 | TTATGGCTGGGCCTCGTCCTACTAATAGATTTCTAAAGGACCTCTCTATCAACACTTCTG |
Microarray Signals from CUST_9030_PI426222305
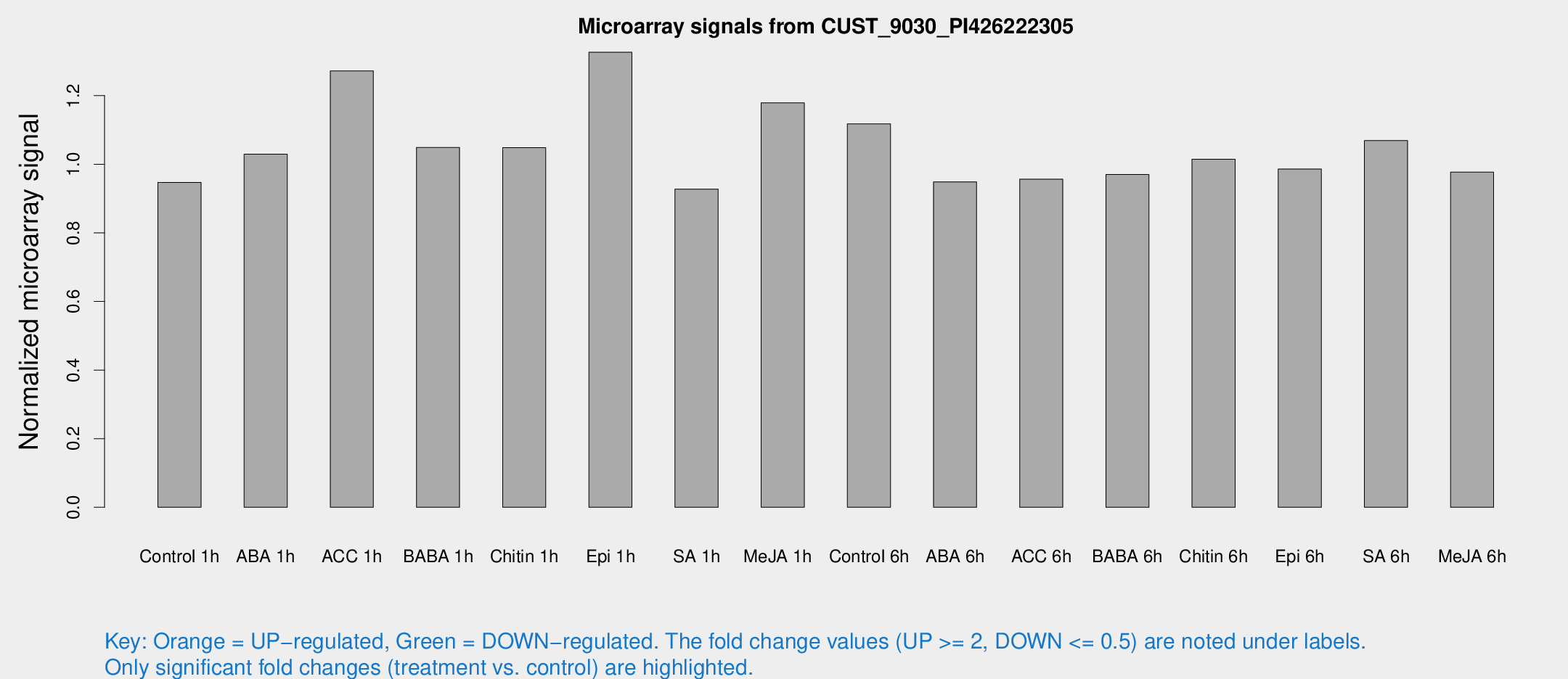
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.74245 | 3.33421 | 0.947121 | 0.548718 |
| ABA 1h | 5.51057 | 3.19261 | 1.02941 | 0.596121 |
| ACC 1h | 8.08884 | 4.1978 | 1.27194 | 0.658271 |
| BABA 1h | 6.13385 | 3.56307 | 1.04885 | 0.607365 |
| Chitin 1h | 5.69287 | 3.32653 | 1.04855 | 0.602266 |
| Epi 1h | 7.11493 | 3.24808 | 1.32658 | 0.624711 |
| SA 1h | 5.77039 | 3.34838 | 0.927544 | 0.537601 |
| Me-JA 1h | 5.86096 | 3.41745 | 1.17892 | 0.682613 |
| Control 6h | 6.9558 | 3.46644 | 1.11777 | 0.579988 |
| ABA 6h | 6.1074 | 3.53994 | 0.948812 | 0.549419 |
| ACC 6h | 6.80056 | 4.04125 | 0.956549 | 0.554258 |
| BABA 6h | 6.58247 | 3.82002 | 0.970164 | 0.561851 |
| Chitin 6h | 6.53851 | 3.78958 | 1.0145 | 0.58742 |
| Epi 6h | 6.81824 | 4.01045 | 0.985888 | 0.571823 |
| SA 6h | 6.39946 | 3.65913 | 1.0689 | 0.611464 |
| Me-JA 6h | 5.88557 | 3.41049 | 0.977 | 0.56609 |
Source Transcript PGSC0003DMT400033386 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G33830.2 | +2 | 2e-25 | 98 | 42/53 (79%) | Dormancy/auxin associated family protein | chr2:14309768-14310286 REVERSE LENGTH=108 |
