Probe CUST_6715_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6715_PI426222305 | JHI_St_60k_v1 | DMT400014679 | TCTTTTGACTCTTACCCAGTAGGTGTTGAAGGTGGACTTTAGATGTTAATAAGACTTACA |
All Microarray Probes Designed to Gene DMG402005729
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6379_PI426222305 | JHI_St_60k_v1 | DMT400014682 | AGAAACAGATTTTGCAGAGCCAACTTGCCTTTGTCGGGACCGGACTTATTCCTATTACTG |
| CUST_6715_PI426222305 | JHI_St_60k_v1 | DMT400014679 | TCTTTTGACTCTTACCCAGTAGGTGTTGAAGGTGGACTTTAGATGTTAATAAGACTTACA |
Microarray Signals from CUST_6715_PI426222305
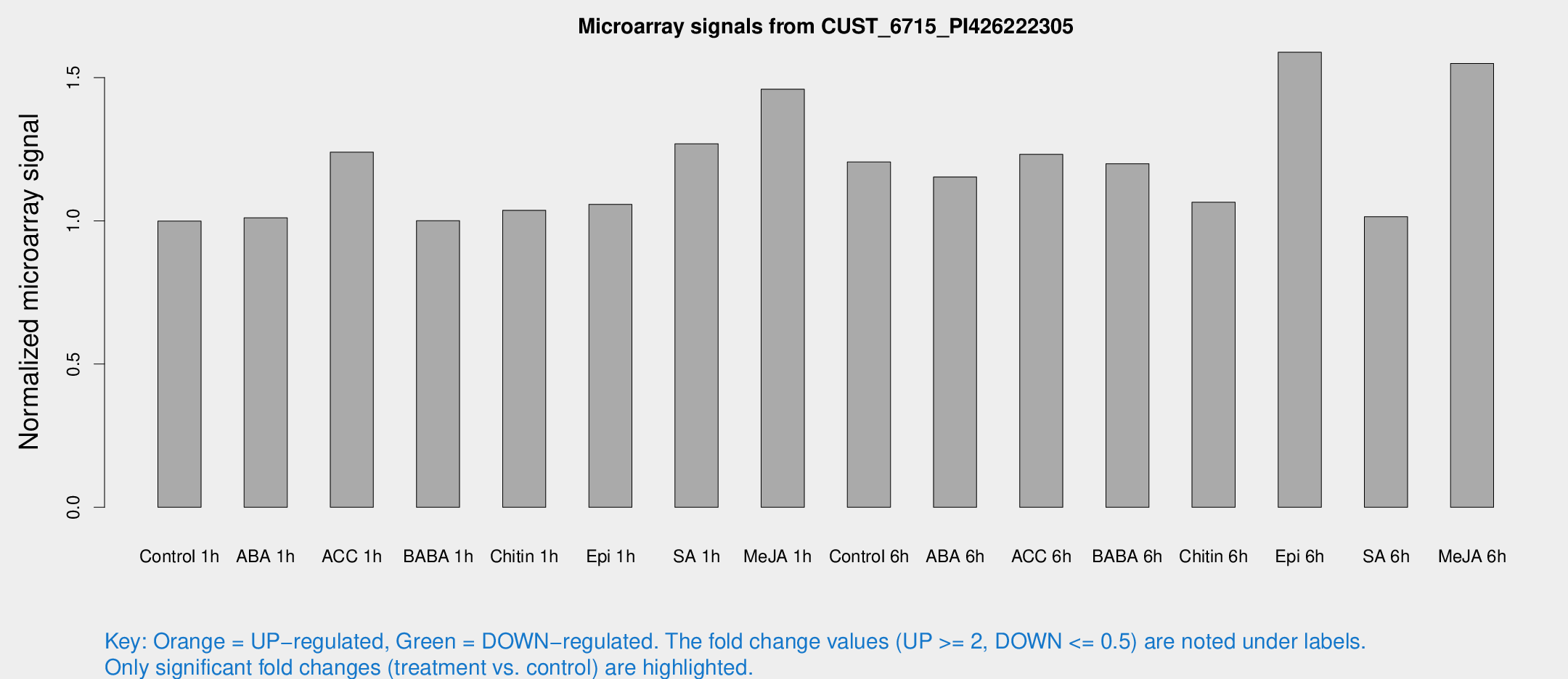
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.8716 | 3.06506 | 0.998845 | 0.530791 |
| ABA 1h | 5.13134 | 2.9761 | 1.01076 | 0.575815 |
| ACC 1h | 7.82993 | 3.35143 | 1.23956 | 0.599459 |
| BABA 1h | 5.62243 | 3.25962 | 1.00054 | 0.579841 |
| Chitin 1h | 5.36348 | 3.11674 | 1.03675 | 0.590856 |
| Epi 1h | 5.34326 | 3.10009 | 1.05748 | 0.612434 |
| SA 1h | 8.69285 | 3.35763 | 1.26904 | 0.593979 |
| Me-JA 1h | 7.23548 | 3.19179 | 1.4594 | 0.707303 |
| Control 6h | 7.51075 | 3.25833 | 1.20526 | 0.591143 |
| ABA 6h | 7.55144 | 3.36806 | 1.153 | 0.5691 |
| ACC 6h | 8.59302 | 3.95038 | 1.23212 | 0.599402 |
| BABA 6h | 8.46981 | 3.65718 | 1.19937 | 0.5889 |
| Chitin 6h | 6.6575 | 3.54531 | 1.06467 | 0.574794 |
| Epi 6h | 11.8945 | 4.52689 | 1.58845 | 0.659204 |
| SA 6h | 5.85138 | 3.39318 | 1.01439 | 0.587797 |
| Me-JA 6h | 11.6567 | 6.14372 | 1.54906 | 0.750973 |
Source Transcript PGSC0003DMT400014679 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G66910.1 | +3 | 2e-20 | 96 | 76/248 (31%) | Protein kinase superfamily protein | chr1:24961634-24963941 REVERSE LENGTH=666 |
