Probe CUST_6379_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6379_PI426222305 | JHI_St_60k_v1 | DMT400014682 | AGAAACAGATTTTGCAGAGCCAACTTGCCTTTGTCGGGACCGGACTTATTCCTATTACTG |
All Microarray Probes Designed to Gene DMG402005729
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6379_PI426222305 | JHI_St_60k_v1 | DMT400014682 | AGAAACAGATTTTGCAGAGCCAACTTGCCTTTGTCGGGACCGGACTTATTCCTATTACTG |
| CUST_6715_PI426222305 | JHI_St_60k_v1 | DMT400014679 | TCTTTTGACTCTTACCCAGTAGGTGTTGAAGGTGGACTTTAGATGTTAATAAGACTTACA |
Microarray Signals from CUST_6379_PI426222305
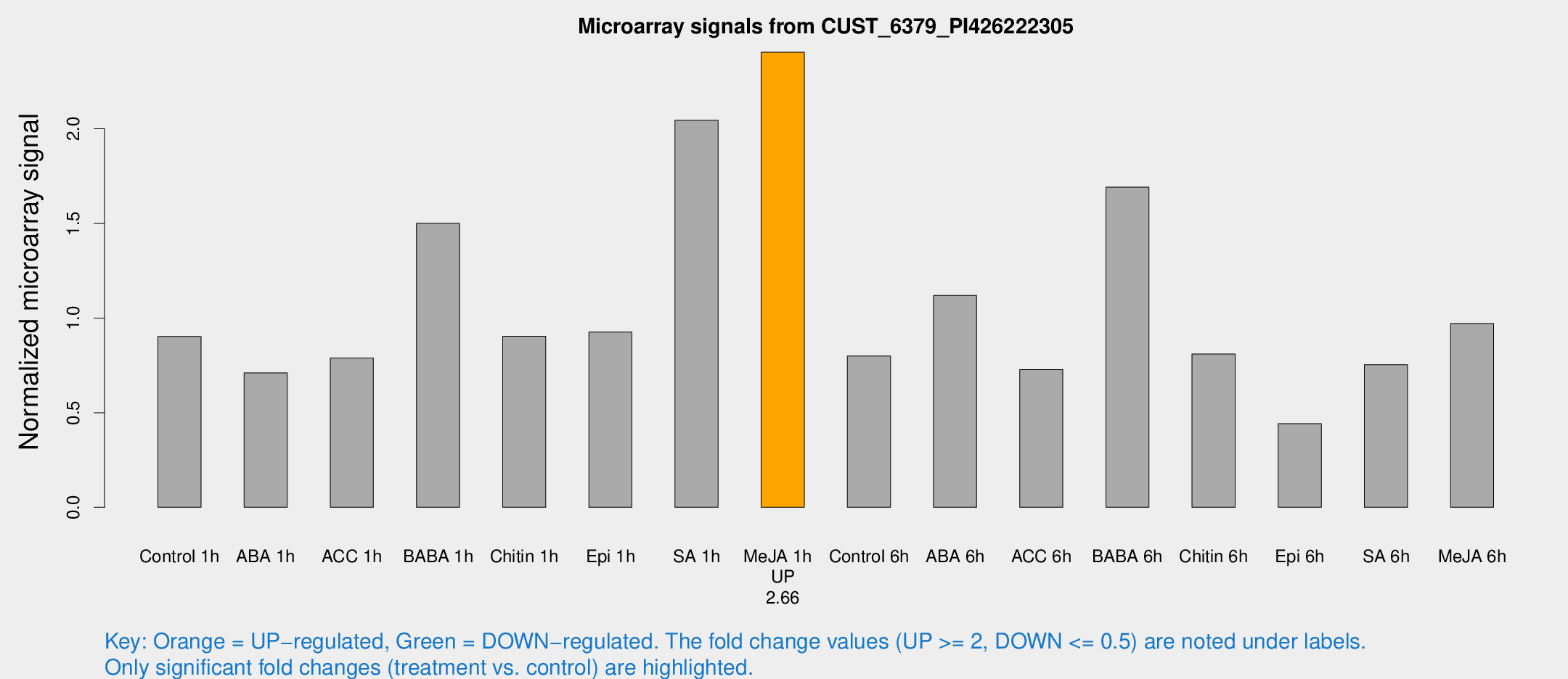
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 32.5279 | 4.05328 | 0.903215 | 0.117451 |
| ABA 1h | 22.8074 | 3.69092 | 0.71017 | 0.117641 |
| ACC 1h | 35.0463 | 12.3233 | 0.789281 | 0.368965 |
| BABA 1h | 53.0119 | 8.66265 | 1.50125 | 0.144367 |
| Chitin 1h | 29.4336 | 3.95475 | 0.904275 | 0.125126 |
| Epi 1h | 28.6205 | 4.05586 | 0.925716 | 0.134399 |
| SA 1h | 78.466 | 18.0269 | 2.04564 | 0.490486 |
| Me-JA 1h | 70.4153 | 6.41961 | 2.40464 | 0.203006 |
| Control 6h | 37.2015 | 14.37 | 0.799874 | 0.479236 |
| ABA 6h | 50.0496 | 16.9304 | 1.11978 | 0.518492 |
| ACC 6h | 35.1152 | 11.797 | 0.727663 | 0.291074 |
| BABA 6h | 74.5497 | 24.1623 | 1.69187 | 0.496793 |
| Chitin 6h | 32.633 | 8.44119 | 0.809769 | 0.228524 |
| Epi 6h | 22.7019 | 9.85058 | 0.441682 | 0.256332 |
| SA 6h | 40.9086 | 26.667 | 0.753233 | 0.940277 |
| Me-JA 6h | 38.414 | 12.3487 | 0.970726 | 0.258364 |
Source Transcript PGSC0003DMT400014682 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G18390.2 | +2 | 2e-168 | 506 | 298/626 (48%) | Protein kinase superfamily protein | chr1:6327463-6329935 FORWARD LENGTH=654 |
