Probe CUST_6622_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6622_PI426222305 | JHI_St_60k_v1 | DMT400014362 | AATCAATCTTCGCCATGGACTAATGGAATCCGACCCCATTTCAATCCTGATGACTACTGG |
All Microarray Probes Designed to Gene DMG400005633
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6622_PI426222305 | JHI_St_60k_v1 | DMT400014362 | AATCAATCTTCGCCATGGACTAATGGAATCCGACCCCATTTCAATCCTGATGACTACTGG |
| CUST_6425_PI426222305 | JHI_St_60k_v1 | DMT400014363 | ACTGGCCCGATTTCTCGGTATTTTATTCTTCTTCCTCCACAATTATTGTTCGTGTTGTTG |
Microarray Signals from CUST_6622_PI426222305
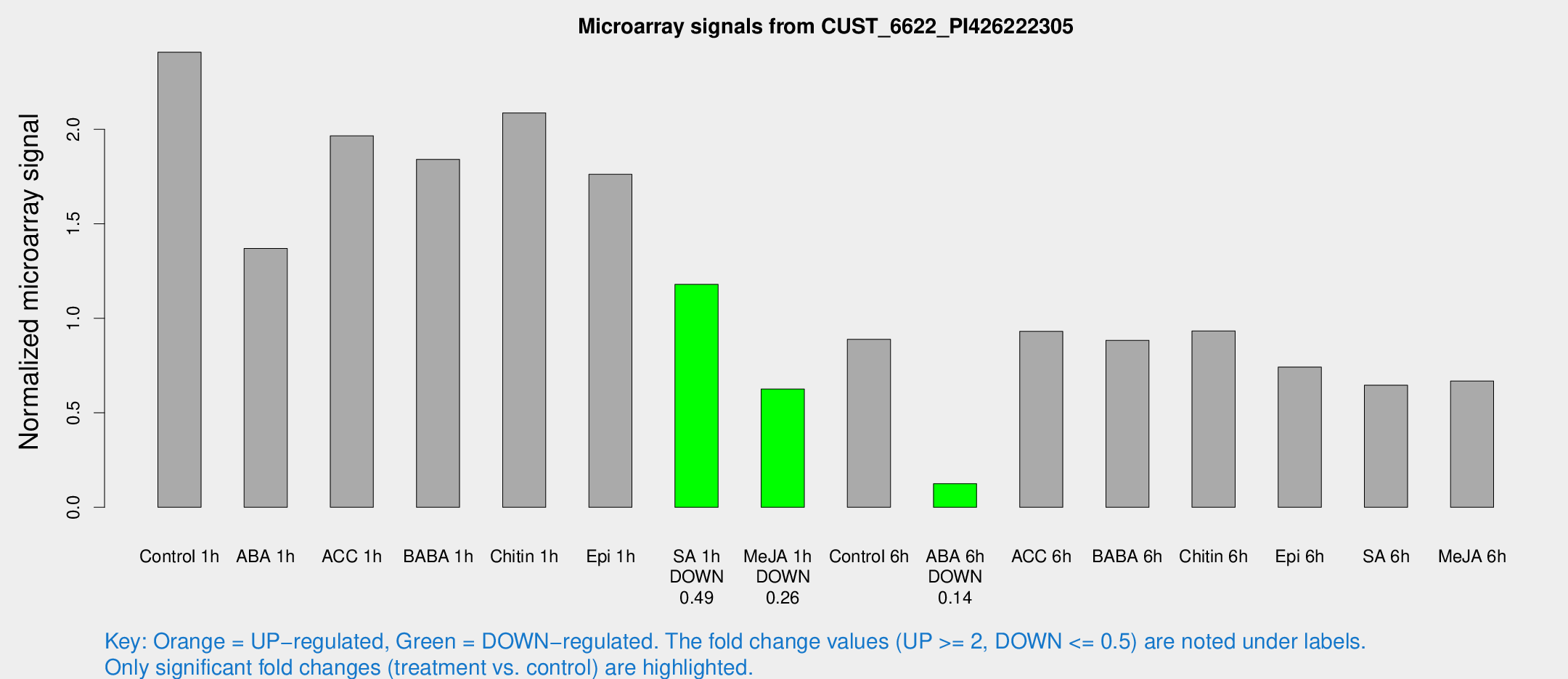
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5851.45 | 795.514 | 2.40822 | 0.250104 |
| ABA 1h | 3026.92 | 695.245 | 1.36982 | 0.20924 |
| ACC 1h | 4987.29 | 1007.17 | 1.96555 | 0.317855 |
| BABA 1h | 4411.99 | 846.677 | 1.8415 | 0.207344 |
| Chitin 1h | 4577.71 | 673.319 | 2.08697 | 0.202995 |
| Epi 1h | 3638.52 | 210.603 | 1.76201 | 0.120674 |
| SA 1h | 2977.65 | 525.598 | 1.17959 | 0.139627 |
| Me-JA 1h | 1225.85 | 102.479 | 0.626272 | 0.065084 |
| Control 6h | 2311.33 | 635.299 | 0.888979 | 0.207464 |
| ABA 6h | 321.178 | 39.123 | 0.125068 | 0.0199985 |
| ACC 6h | 2627.4 | 480.762 | 0.931062 | 0.0571178 |
| BABA 6h | 2464.7 | 545.095 | 0.883698 | 0.177804 |
| Chitin 6h | 2366.08 | 136.713 | 0.933577 | 0.0539267 |
| Epi 6h | 2058.88 | 394.314 | 0.74175 | 0.145954 |
| SA 6h | 1595.41 | 317.609 | 0.645919 | 0.0707798 |
| Me-JA 6h | 1789.54 | 563.413 | 0.668377 | 0.200406 |
Source Transcript PGSC0003DMT400014362 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G25440.1 | +3 | 7e-89 | 281 | 181/411 (44%) | B-box type zinc finger protein with CCT domain | chr1:8933939-8935284 REVERSE LENGTH=417 |
